魚類肌肉結構性蛋白:T 型肌鈣蛋白(Troponin T)的分子結
構、胚胎表現及基因調控﹝2/2﹞
計畫類別: 個別型計畫
計畫編號: NSC91-2313-B-002-289-
執行期間: 91 年 08 月 01 日至 92 年 07 月 31 日
執行單位: 國立臺灣大學漁業科學研究所
計畫主持人: 蔡懷楨
計畫參與人員: 王玉雪, 蕭崇德, 蔡維原
報告類型: 完整報告
處理方式: 本計畫可公開查詢
中 華 民 國 92 年 10 月 28 日
ARTICLE
Molecular Structure and Developmental Expression
of Three Muscle-Type Troponin T Genes in
Zebrafish
Chung-Der Hsiao, Wei-Yuan Tsai, Long-Shyan Horng, and Huai-Jen Tsai*
Troponin T (Tnnt), a troponin component, interacts with tropomyosin and is crucial to the regulation of striated muscle contraction. To gain insight into the molecular evolution and developmental regulation of Tnnt gene (Tnnt) in lower vertebrates, zebrafish Tnnt1 (slow Tnnt), Tnnt2 (cardiac Tnnt), and Tnnt3b (fast Tnnt isoform b) were characterized. The polypeptides of zebrafish Tnnt1, Tnnt2, and Tnnt3b were conserved in the central tropomyosin- and C-terminal troponin I-binding domains. However, the N-terminal hypervariable regions were highly extended and rich in glutamic acid in polypeptides of Tnnt1 and Tnnt2, but not Tnnt3b. The Tnnt2 and Tnnt3b contain introns, whereas Tnnt1 is intron-free. During development, large to small, alternatively spliced variants were detected in Tnnt2, but not in Tnnt1 or Tnnt3. Whole-mount in situ hybridization showed zebrafish Tnnt1 and Tnnt2 are activated during early somitogenesis (10 hr postfertilization, hpf) and cardiogenesis (14 hpf), respectively, but Tnnt3b is not activated until middle somitogenesis (18 hpf). Tnnt2 and Tnnt3b expression was cardiac- and fast-muscle specific, but Tnnt1 was expressed in both slow and fast muscles. We propose that three, distinct, muscle-type Tnnt evolved after the divergence of fish and deuterostome invertebrates. In zebrafish, the developmental regulation of Tnnt during somitogenesis and cardiogenesis is more restricted and simpler than in tetrapods. These new findings may provide insight into the developmental regulation and molecular evolution of vertebrate Tnnt. Developmental Dynamics 227:
266 –279, 2003.©2003 Wiley-Liss, Inc.
Key words: zebrafish; troponin T; expression pattern; alternative splicing; gene structure; molecular evolution
Received 6 August 2002; Accepted 21 February 2003
INTRODUCTION
In vertebrates, striated muscle is composed of three distinct fiber types: cardiac, slow-twitch, and fast-twitch (Oota and Saitou, 1999). Transgenic techniques have been used to study how trans-acting tran-scription factors and cis-acting ele-ments control cardiac and skeletal muscle-fiber specificity in mammals (Nakayama et al., 1996; Calvo et al., 1999; Wang et al., 2000; Koban et al., 2001; Yan et al., 2001). However, the
regulatory elements that restrict the transcription of genes encoding contractile proteins specific to fish cardiac or slow- or fast-twitch skele-tal muscles have been the subject of only a few studies (Kusakabe et al., 1999; Ju et al., 1999; Xu et al., 1999; Tan and Du, 2002). Fish are excellent models for studying the develop-mental and fiber-specific regulation of muscle diversity. Compared with mammals, the fish genome is more compact (Venkatesh et al., 2000), so
it is easier to dissect the enhancers and silencers that control tissue-spe-cific expression (Aparicio et al., 1995; Muller et al., 2002). In addition, a fish embryo is transparent (Briggs, 2002), so the processes of cardiac and skeletal muscle commitment, differ-entiation, and maturation can be observed directly in vivo. Finally, the organization of slow-twitch and fast-twitch muscles in fish is more homog-enous than in birds and mammals. In fish, slow- and fast-twitch muscles
Institute of Fisheries Science, National Taiwan University, Taipei, Taiwan
Grant sponsor: National Science Council, Republic of China; Grant number: NSC 90-2313-B-002-319.
Dr. Hsiao’s present address is Institute of Molecular Biology, Academia Sinica, 128, Sec. 2, Academia Road, Taipei 115, Taiwan. *Correspondence to: Huai-Jen Tsai, Institute of Fisheries Science, National Taiwan University, Taipei, Taiwan.
E-mail: hjtsai@ccms.ntu.edu.tw DOI 10.1002/dvdy.10305
DEVELOPMENTAL DYNAMICS 227:266 –279, 2003
are spatially separate: slow muscles form a superficial layer, and fast muscles a deep layer (Waterman, 1969; Koumans and Akster, 1995). These traits make it easy to assay developmental regulation of mus-cle-fiber specificity in fish.
Troponin T (Tnnt) is the tropomyo-sin-binding subunit of the troponin complex and is crucial to the regu-lation of striated muscle contraction (reviewed by Perry, 1998; Filatov et al., 1999). The occurrence of the Tnnt gene (Tnnt) is associated with the evolution of muscle tissues. For example, a single Tnnt has been identified in the smooth muscle of
Caenorhabditis elegans (Myers et
al., 1996), in the striated muscle of
Drosophila melanogaster (Fyrberg et
al., 1990, Benoist et al., 1998), and in the adductor muscle of scallops (In-oue et al., 1996, 1998). In deuteros-tome invertebrates, two distinct Tnnt were identified in the smooth and striated muscles of ascidians (Endo et al., 1996, 1997). In tetrapods, three fiber-specific Tnnt have evolved to regulate muscle contraction in car-diac (Cooper and Ordahl, 1984), slow-twitch (Gahlmann et al., 1987), and fast-twitch (Wilkinson et al., 1984; Bucher et al., 1999) muscles. In addition, each fiber-specific Tnnt can yield a variety of isoforms by alternative splicing (reviewed by Schiaffino and Reggiani, 1996; Perry, 1998). Transitional expression of alter-natively spliced Tnnt during different developmental stages and in differ-ent muscle types can modulate muscle calcium sensitivity and mus-cle performance in tetrapods (Jin et al., 1996; Schiaffino and Reggiani, 1996; Perry, 1998; Marden et al., 1999). In fish, several stage- (Ya-mano et al., 1991; Johnston et al., 1997) or tissue- (Thys et al., 1998, 2001) specific Tnnt have been iden-tified at the protein level. Unfortu-nately, little is known about the genomic structure and develop-mental regulation of Tnnt at the gene level (Waddleton et al., 1999; Xu et al., 2000). To better understand the mechanism controlling Tnnt mus-cle fiber diversification in lower ver-tebrates, we isolated three distinct fiber types of zebrafish Tnnt and compared their patterns of
spatio-temporal expression to those of their tetrapod counterparts.
RESULTS
Identification and
Characterization of Zebrafish
Tnnt
A BLAST search of the zebrafish ex-pression sequence tag (EST) data-base identified three zebrafish EST clones: AI883430, AW455177, and AI353765, that exhibited high se-quence homology with the trout
Tnnt1 (Waddleton et al., 1999),
ze-brafish Tnnt (Xu et al., 2000), and hu-man Tnnt2 (Mesnard et al., 1993), re-spectively. We also identified two potential Xenopus EST clones, BG514147 and BJ055552, that exhib-ited high sequence homology with chicken Tnnt2 (Cooper and Ordahl, 1984) and mouse Tnnt3 (Wang and Jin, 1997), respectively. EST clone-specific primers were designed to amplify potential Tnnt cDNAs from zebrafish and Xenopus by rapid am-plification of cDNA ends (RACE). Full-length cDNA of zebrafish and
Xenopus Tnnt were assembled with
sequences derived from 3⬘- and 5⬘-RACE. Three zebrafish and two
Xe-nopus Tnnt2 clones were identified.
Based on sequence analysis, the three zebrafish and two Xenopus
Tnnt2 clones were alternatively spliced variants derived from a sin-gle gene. In zebrafish, the longest, isolated Tnnt2 clone was identical to that reported by Sehnert et al. (2002). Two other alternatively spliced variants lacked 18 and 10 amino acid residues, respectively, in the N-terminal hypervariable region (Fig. 1A). In contrast, only one clone each of Tnnt1 and Tnnt3 was iso-lated from zebrafish. The Tnnt3 clone reported here and the Tnnt clone reported by Xu et al. (2000) ex-hibited many amino acid residue substitutions throughout the open reading frame and had distinct nu-cleotide sequences in both the 5 ⬘-and 3⬘-untranslated regions. Thus, these two Tnnt3 clones seemed to be encoded by a distinct gene. Be-cause they are co-orthologous to tetrapod Tnnt3 (Fig. 2), we named them Tnnt3a (⫽Tnnt, Xu et al., 2000)
and Tnnt3b (Table 1 summarizes their molecular features).
The number of deduced amino acid residues in Tnnt1 (290) and
Tnnt2 (282) was much greater than
in Tnnt3a (230) and Tnnt3b (232). Moreover, there were many more negatively charged residues (pri-marily glutamic acid) in Tnnt1 and
Tnnt2 than in Tnnt3b. Thus, Tnnt1 and Tnnt2 have acidic pI and are highly
negatively charged at pH 7, whereas Tnnt3b has a basic pI and is positively charged at pH 7. To better understand the relationship among different Tnnt, the zebrafish and
Xe-nopus Tnnt sequences were aligned
based on deduced amino acid se-quences (Fig.1A). The three types of zebrafish Tnnt and two types of
Xe-nopus Tnnt were highly conserved in
the central tropomyosin-binding do-main and C-terminal troponin I-bind-ing domain segments, but they were highly divergent in the N-terminal hy-pervariable region. The deduced amino acid sequences of zebrafish
Tnnt2 and Tnnt3b exhibited high
se-quence identity with tetrapod Tnnt2 (57.4 – 64.5%) and Tnnt3 (67.7– 69.0%), respectively, whereas Tnnt1 exhib-ited moderate identity with tetrapod
Tnnt1 (52.5–54.2%). Unexpectedly,
the amino acid identity of zebrafish and trout Tnnt1 (48.9%) was lower than that between zebrafish and tetrapod Tnnt1. Thus, Tnnt1 were more divergent than Tnnt2 and
Tnnt3 in teleost branch. Of interest, in Tnnt1, the number of amino acid
res-idues in the N-terminal hypervariable region decreased from 85 in ze-brafish to 53 in humans, but in Tnnt3 they increased from 25 in zebrafish to 46 in humans (Fig. 1B).
Phylogenetic Analysis of Tnnt
To examine the evolutionary rela-tionship between teleost and tetra-pod Tnnt, we conducted a phyloge-netic tree based on unspliced primary sequences of Tnnt. Ascidian
Tnnt from embryonic striated muscle
and the adult body wall (Endo et al., 1996, 1997) were included for com-parison. The occurrence of three dis-tinct types of Tnnt evolved after the divergence of deuterostome inver-tebrate and teleost lineages (Fig. 2). Zebrafish Tnnt2 and Tnnt3 clustered
with their tetrapod counterparts into two distinct monophyletic groups. Orthologous relationship between zebrafish and tetrapods in the Tnnt2 and Tnnt3 branches were strongly supported. In fish, the amino acid substitution rate in the Tnnt1 branch was faster than in the Tnnt2 and
Tnnt3 branches. There was not an
unambiguous orthologous relation-ship between zebrafish and tetra-pod Tnnt1 (Fig. 2). To determine whether this phenomenon was caused by sequence variation in the N-terminal hypervariable region, we constructed a phylogenetic tree based on partial Tnnt sequences
that omitted the N-terminal hyper-variable region. However, there was no significant difference in the tree topology (data not shown). That is, fish Tnnt1 was more closely related to the Tnnt2 clade than to the tetra-pod Tnnt1 clade.
Zebrafish Tnnt Gene Structure
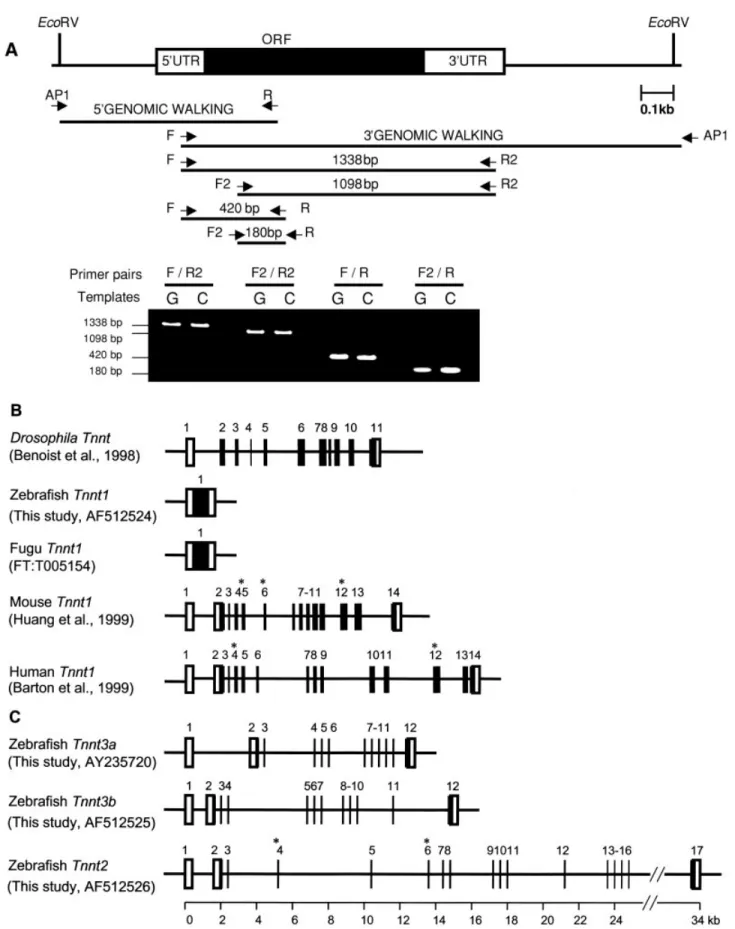
During the initial cloning of partial sequences (N-terminal hypervari-able region) of zebrafish Tnnt1, the polymerase chain reaction (PCR) products amplified from cDNA and genomic DNA by primers Tnnt1-F and Tnnt1-R were identical in size.
Furthermore, several primers that span the entire Tnnt1 cDNA were used for genomic PCR and reverse transcriptase-PCR (RT-PCR). This strategy consistently produced PCR products with the same size and se-quences (Fig. 3A). In addition, using PCR-based genomic walking, we obtained a genomic clone that covered the entire Tnnt1 (including 0.3-kb-upstream and 0.7-kb-down-stream regions; Fig. 3A). These results confirm that zebrafish Tnnt1 lacks in-trons. By using bioinformatics, we searched for Tnnt1 homologs in fugu draft genome sequences. We iden-tified one genomic clone (FT:
Fig. 1. Sequence alignment and homology analysis of Tnnt isolated in this study. A: The aligned, deduced amino acid sequences of zebrafish Tnnt1, Tnnt2, Tnnt3a, Tnnt3b, and Xenopus XTnnt2, XTnnt3. Gaps introduced during the alignment are shown by hyphens. Amino acid residues conserved among the Tnnt family members are highlighted in black; residues conserved in at least 50% are highlighted in gray. Putative tropomyosin (Tm)- and troponin I (TnI)-binding domains (Pearlstone et al., 1977) are indicated with solid underlines. Alternatively spliced exons detected in zebrafish and Xenopus Tnnt2 are indicated with dotted underlines. B: Schematic representation of the evolution among vertebrates of Tnnt N-terminal hypervariable regions. The gray boxes indicate the N-terminal hypervariable regions (residue numbers are on the right side). The empty boxes indicate the full-length Tnnt excluding the N-terminal hypervariable region. The black box indicates the 3⬘ alternatively spliced exon of human Tnnt1 (Gahlmann et al., 1987). The hatched box indicates the metal-binding cluster of turkey Tnnt3 (Jin and Samanez, 2001). The right panels list the amino acid identity between zebrafish and other species for each type of muscle Tnnt. F, full-length amino acid sequences; -N, partial sequences excluding N-terminal hypervariable region. 268 HSIAO ET AL.
T005154) that corresponds to fugu
Tnnt1. Gene prediction analysis showed it was also intronless. In con-trast, numerous introns interrupt
Tnnt1 orthologs in chickens (12
in-trons; Hirao et al., 2001), mice (13 introns; Huang et al., 1999; Huang and Jin, 1999), and humans (13 in-trons; Barton et al., 1999; Fig. 3B). This unexpected finding prompted us to
examine the gene structure of ze-brafish Tnnt2 and Tnnt3. An incom-plete (24 kb) genomic clone of ze-brafish Tnnt2, corresponding to exons 1 to 13, was reported by Seh-nert et al. (2002). By using gene mining and PCR-based genomic walking, we obtained complete genomic clones of zebrafish Tnnt2 (34 kb), Tnnt3a (12 kb), and Tnnt3b (15 kb; Fig. 3C). Like
chicken (9 kb and 18 exons; Cooper and Ordahl, 1985), rat (19 kb and 16 exons; Jin et al., 1992), and human
Tnnt2 (17 kb and 17 exons; Farza et
al., 1998), zebrafish Tnnt2 has a com-plex intron– exon organization (17 exons), but at 34 kb, it is two- to nearly fourfold larger than its tetra-pod counterparts. In contrast, the gene structures of zebrafish Tnnt3a
Fig. 2. An unrooted radial gene tree for Tnnt among vertebrates. The gene tree was constructed with the neighbor-joining method (Pearson et al., 1999), using 1,000 bootstrap values. The marker length of 0.1 corresponds to 10% sequence difference. The Tnnt1, Tnnt2, and Tnnt3 clades are marked in green, red, and blue, respectively. See the Experimental Procedure section for details on the sources of
Tnnt. [Color figure can be viewed in the online issue, which is available at www.interscience.wiley.com.]
TABLE 1. Summary of the Troponin T cDNAs Used in This Study
Gene names Gene abbreviations cDNA length (bp) Coding region (aa) Mr (kDa) pI GenBank accession no. References Zebrafish
Slow troponin T Tnnt1 1461 290 34.7 5.0 AF425255 This study
Cardiac troponin T Tnnt2 1047 282 34.0 5.1 AF434187 Sehnert et al. (2002); this study
Fast troponin T isoform a Tnnt3a 1098 230 27.8 9.9 AF180889 Xu et al. (2000) Fast troponin T isoform b Tnnt3b 1099 232 26.5 9.9 AF425741 This study
Xenopus
Cardiac troponin T XTnnt2 1047 272 32.6 5.5 AF467919 This study Fast troponin T XTnnt3 1521 276 32.9 5.1 AY114144 This study
Fig. 3. Comparison of Tnnt gene structure in different vertebrates. A: A schematic, scale diagram of Tnnt1 gene structure. Black box, coding region. Empty boxes, untranslated regions. Solid lines indicate 5⬘- and 3⬘-flanking regions. The primers used to clone the flanking regions of genomic DNA, and for RT-PCR on cDNA, are indicated below. PCR fragments amplified from genomic DNA (G) and cDNA (C) using different combinations of Tnnt1-specific primer pairs were the same size. B: The complete gene structures of zebrafish and fugu Tnnt1 are compared with that of Drosophila Tnnt (Benoist et al., 1998), mouse Tnnt1 (Huang et al., 1999), and human Tnnt1 (Barton et al., 1999).
C: The complete gene structures of zebrafish Tnnt3a, Tnnt3b, and Tnnt2. Exons are shown as boxes: black boxes are open reading frames,
empty boxes are 5⬘- or 3⬘-untranslated regions. Asterisks denote the alternatively spliced exons. 270 HSIAO ET AL.
(12 kb and 11 exons) and Tnnt3b (15 kb and 11 exons) are smaller and simpler than their tetrapod counter-parts in quail (33 kb and 25 exons; Bucher et al., 1999) and rat (16 kb and 19 exons; Breitbart and Nadal-Ginard, 1986). This comparison high-lights the unique, relatively compact genomic organization of Tnnt1
within the zebrafish Tnnt family.
Temporal Expression of Tnnt in
Zebrafish
We compared the spatiotemporal expression of Tnnt in zebrafish, a lower vertebrate, with that in tetra-pods. Total RNA was extracted from zebrafish embryos at eight different stages, ranging from 9 hpf (90% epi-boly) to 72 hpf (hatched fry), and from adult fish. RT-PCR was per-formed to compare the relative abundance of Tnnt transcripts. Tnnt1 was the first of the three types of Tnnt to be detected (Fig. 4). Tnnt1 was activated approximately 12 hpf and was expressed vigorously 24 hpf. However, Tnnt1 expression was very low in adults. Tnnt2 transcripts first appeared 15 hpf and remained
constant through adulthood. Tnnt3a (24 hpf) and Tnnt3b (21 hpf) tran-scripts appeared much later than
Tnnt1 (12 hpf) and Tnnt2 (15 hpf).
Each exhibited a distinct expression profile during development: Tnnt3a was expressed transiently in em-bryos, whereas Tnnt3b expression was constant from the embryonic stage through adulthood. Com-pared with sMyHC, which is ex-pressed specifically in slow muscle, and-actin, which is expressed con-stantly in the entire body, only low levels of Tnnt1 and Tnnt3a were ex-pressed in adults.
To detect alternatively spliced
Tnnt variants in zebrafish, RT-PCR was
carried out by using primers de-signed to amplify various fragments of the N-terminal hypervariable re-gion or the entire Tnnt open reading frame. There was no variation in the size of the RT-PCR products pro-duced by the primers for Tnnt1,
Tnnt3a, and Tnnt3b (data not shown). In contrast, one Tnnt2 primer pair (Tnnt2-F and Tnnt2-R3) pro-duced RT-PCR products of three sizes (Fig. 5A,B). The expected,
318-bp fragment (exon 1 to exon 8) was detected in embryos and adults (Fig. 5B). Two additional bands (282-bp and 264-bp), which first ap-peared 72 hpf, were expressed pre-dominately in adults. The two bands were cloned and sequenced, yield-ing 282-bp and 264-bp amplified products that corresponded to a splicing variant lacking exon 6 ignated Tnnt2-6E) and exon 4 (des-ignated Tnnt2-4E), respectively (Fig. 5C).
Spatial Expression of Tnnt in
Zebrafish
To examine the spatial expression of
Tnnt, whole-mount in situ
hybridiza-tion (WISH) was performed on ze-brafish embryos collected 9 to 72 hpf. WISH detected Tnnt transcripts earlier than RT-PCR. We found the onset of Tnnt2 expression in the bilat-eral precardiac mesoderm (Fig. 6A,F) occurred 2.5 hr earlier than re-ported in a previous study of the de-velopmental expression of Tnnt2 in zebrafish (Sehnert et al., 2002). Tnnt2-expressing cells rapidly migrated to-ward the notochord 17 hpf (Fig.
Fig. 4. Tnnt temporal expression patterns in developing zebrafish embryos as detected by reverse transcriptase-polymerase chain
reaction (RT-PCR). Total RNA was isolated from different developmental stages, and primers specific for sMyHC, Tnnt1, Tnnt2, Tnnt3a, and
Tnnt3b were used for RT-PCR.-actin was used as an internal control. Numbers in the right panel indicate the molecular mass of RT-PCR
6B,G) and fused to become a single heart cone 18 hpf. From 24 hpf on-ward, Tnnt2 transcripts were up-reg-ulated and expressed robustly in the looping heart tube (Fig. 6C,D,H,I). By 48 hpf, Tnnt2 transcripts were specif-ically and uniformly detected in both the atrium and ventricle (Fig. 6E,J). No Tnnt2 transcripts were de-tected in skeletal muscles. Thus, un-like its tetrapod counterparts, ex-pression of zebrafish Tnnt2 was highly cardiac-specific. To better under-stand the ontogeny of Tnnt2, the ex-pression of several cardiac-specific contractile proteins during early car-diogenesis was determined on tran-scription level. We found cmlc2 (Fig. 6L), cmlc1 (Fig. 6M), vmhc (Fig. 6N), and cTnC (Fig. 6O) are synchro-nously activated 17–18 hpf, whereas
Tnnt2 (Fig. 6K) is activated 14 –15 hpf.
Compared with sMyHC, a slow-muscle-specific marker activated 9 hpf (Fig. 7A), Tnnt1 was activated in adaxial cells (slow muscle precur-sors) 10 hpf, during early somitogen-esis (Fig. 7F). By 15 hpf, sMyHC (Fig. 7B) and Tnnt1 (Fig. 7G) were vigor-ously expressed in all developing
somites and adaxial cells. Tnnt3a and Tnnt3b were not detected until 16.5 and 18 hpf, respectively. By 18 hpf, all developing somites stained positive for sMyHC (Fig. 7C) and
Tnnt1 (Fig. 7H), but only 10 somites
stained positive for Tnnt3a (Fig. 7M) and only one was positive for Tnnt3b (Fig. 7R). Although Tnnt3a and
Tnnt3b were activated much later
than sMyHC and Tnnt1, they were sharply up-regulated from 18 hpf on-ward. By 24 hpf, only 3 and 5 newly forming somites were unstained for
Tnnt3a (Fig. 7N) and Tnnt3b (Fig. 7S),
respectively. To verify the muscle fi-ber-specificity of Tnnt1 and Tnnt3 in zebrafish, trunk-level tissue sections were examined. Compared with the specific staining of sMyHC in slow muscles (Fig. 7E), Tnnt1 transcripts (Fig. 7J) were detected in superficial slow and inner fast muscles, whereas
Tnnt3a (Fig. 7O) and Tnnt3b (Fig. 7T)
transcripts were fast-muscle–spe-cific. From 36 hpf onward, there was a wave of Tnnt1 down-regulation (Fig. 8E) that moved in a rostral– cau-dal direction, beginning in the oldest somites. sMyHC (Fig. 8A), Tnnt3a (Fig.
8I), and Tnnt3b (Fig. 8M) were not down-regulated. Transverse section (data not shown) and
high-magnifi-Fig. 6. Whole-mount in situ hybridization of
Tnnt2 and other cardiac-specific genes
dur-ing zebrafish cardiogenesis. Embryos were hybridized with Tnnt2 (A–K), cmlc2 (L),
cmlc1 (M), vmhc (N), and cTnC (O)
ribo-probes, respectively. A–E: Lateral views, an-terior of embryo to the left. F–I,K–O: Dorsal views, anterior of embryo to the top. J: Fron-tal view of embryo. Embryonic stages are indicated in each panel. h, hour; a, atrium; ht, heart tube; mp, myocardium precursor; v, ventricle. Scale bar⫽ 100 m in all pic-tures.
Fig. 7. Whole-mount in situ hybridization of
Tnnt1, Tnnt3a, Tnnt3b, and sMyHC during
zebrafish early somitogenesis. Embryos were hybridized with sMyHC (A–E), Tnnt1 (F–
J), Tnnt3a (K–O), and Tnnt3b (P–T)
ribo-probes, respectively. A,B,F,G,K,L,P,Q: Dorsal views. C,D,H,I,M,N,R,S: Lateral views. E,J,Q,T: Transverse sections at trunk level. The ante-rior is to the left in all whole-mount stained embryos. Dotted lines indicate the section level. Embryonic stages are indicated in each panel. h, hour. Scale bar⫽ 200 m in D,I,N,S, 140 m in C,H,M,R, 100 m in A,B,F,G,K,L,P,Q, 25m in E,J,O,T.
Fig. 5. Reverse transcriptase-polymerase chain reaction (RT-PCR) analysis of Tnnt2 alternatively spliced variants during zebrafish devel-opment. A: Schematic representation of the positions of the primers used to perform RT-PCR. Empty boxes indicate untranslated regions; solid boxes show open reading frames. B: Age-dependent expression of alternatively spliced, Tnnt2 variants. Alternatively spliced variants were detected by RT-PCR using the primer pair combinations indicated. Two, alternatively spliced variants were detected by primer pair
Tnnt2-F and Tnnt2-R3 in adult fish. C: Schematic illustration of the alternative splicing involved in the generation of Tnnt2 (unspliced form), Tnnt2-6E (excluding exon 6), and Tnnt2-4E (excluding exon 4). The alternatively spliced exons are underlined.
cation view (Fig. 8F) 36 hpf revealed that Tnnt1 was down-regulated in fast muscles, but it was still strongly expressed in pioneer slow muscles. By 48 hpf, Tnnt1 staining were unde-tectable in trunk muscles, but they had been up-regulated in fin buds and jaw and head muscles (Fig. 8G). At the same time, Tnnt3b (Fig. 8O) but not Tnnt3a (Fig. 8K) was weakly activated in fin buds and jaw and head muscles. The staining intensity of sMyHC (Fig. 8D), Tnnt1 (Fig. 8H),
Tnnt3a (Fig. 8L), and Tnnt3b (Fig. 8P)
in craniofacial muscles 72 hpf varied in different muscle fibers. For exam-ple, the sternohyoideus was heavily stained by Tnnt3a and Tnnt3b but weakly stained by sMyHC and Tnnt1. Jaw muscles were heavily stained by
sMyHC, Tnnt1, and Tnnt3b but weakly stained by Tnnt3a.
DISCUSSION
Molecular Characterization of
Tnnt in Zebrafish
In tetrapods, muscle-specific and development-regulated Tnnt have been characterized thoroughly at the protein and gene levels (Jin and Lin, 1988; Sabry and Dhoot, 1991; Wang and Jin, 1997; Ogut et al., 1999; Yonemura et al., 2000). How-ever, few tetrapod orthologs have been characterized at the gene level in teleosts (Waddleton et al., 1999; Xu et al., 2000; Sehnert et al., 2002). In this study, we isolated three
zebrafish Tnnt and two Xenopus Tnnt cDNAs with an EST-based approach. Like their tetrapod counterparts, the deduced amino acids of zebrafish and Xenopus Tnnt are highly con-served in the tropomyosin- and tro-ponin I-binding domains, but they diverse greatly in the N-terminal hy-pervariable region. Among verte-brates, fish Tnnt1 has the longest N-terminal extension, whereas Tnnt3 has the shortest N-terminal exten-sion. Extension of the N-terminal hy-pervariable region in vertebrates, and the C-terminal region in inverte-brates, plays a role in modulating the conformation of Tnnt isoforms (Benoist et al., 1998; Jin and Sa-manez, 2001). As negatively charged residues in the N-terminal
hypervari-Fig. 8. Whole-mount in situ hybridization of Tnnt1, Tnnt3a, Tnnt3b, and sMyHC genes during zebrafish late somitogenesis. Embryos were hybridized with sMyHC (A–D), Tnnt1 (E–H), Tnnt3a (I–L), and Tnnt3b (M–P) riboprobes, respectively. C,G,K,O: Dorsal views. A,B,E,F,I,J,M,N: Lateral views. D,H,L,P: Ventral views. The anterior is to the left in all pictures. Embryonic stages are indicated in each panel. am, adductor mandibulae; fb, fin bud; hh, hyohyoideus; hm, horizontal myospetum; sh, sternohyoideus; sm, somites. Muscle nomenclature is based on Schilling and Kimmel (1997). Scale bar⫽ 50 m in B,F,J,N, 100 m in A,C–E,G–I,K–M,O,P.
able region of Tnnt increase, sensitiv-ity of the myofibrillar contractile ap-paratus to Ca2⫹ (Ogut and Saitou,
1999) and tolerance of acidosis also increase (Ogut and Jin, 1998). Stud-ies that generate pCa/tension curves are needed to determine whether variation in the N-terminal hypervariable region of zebrafish
Tnnt1 and Tnnt3 is involved in
regu-lating contraction in red and white muscles.
During somitogenesis and cardio-genesis in tetrapods, changes in Tnnt isoforms expression resulting from al-ternative splicing (i.e., from large to small, and acidic to basic) are com-mon (Jin and Lin, 1988, Wang and Jin, 1997; Yonemura et al., 2002). In birds and mammals, alternative splicing is most frequent in Tnnt3 (Wang and Jin, 1997; Bucher et al., 1999) and Tnnt2 (Jin et al., 1996), and less frequent in Tnnt1 (Huang et al., 1999; Yonemura et al., 2000). As development proceeded, large to small alternatively spliced variants of
Tnnt2 isoforms were detected in
both zebrafish and Xenopus. Thus, our data provide strong evidence that alternative splicing is an evolu-tionarily conserved mechanism for generating Tnnt2 isoforms in both te-leosts and tetrapods. However, in zebrafish, unlike tetrapods, no alter-natively spliced variants were de-tected in Tnnt1, Tnnt3a, and Tnnt3b. Because zebrafish Tnnt1 lacks
in-trons, the absence of alternatively spliced variants is easily explained. However, it is difficult to explain the lack of alternatively spliced variants of Tnnt3a and Tnnt3b. In chickens, RNA-binding proteins and intronic el-ements related to muscle-specific splicing in skeletal and cardiac mus-cles have been identified in Tnnt (Ryan and Cooper, 1996; Charlet et al., 2002). Further studies are needed on the RNA-binding proteins and in-tronic elements of Tnnt3 in zebrafish.
Evolutionary Significance of
Zebrafish Tnnt1
Homology, phylogenetic, and ex-pression analyses indicate the three fiber types of zebrafish Tnnt evolved before teleost and tetrapod lin-eages diverged. However, the three distinct Tnnt probably evolved after deuterostome invertebrates and fish diverged. Before our study, there was no information on the gene structure of Tnnt in lower vertebrates, so the mechanism by which the three fiber types of Tnnt evolved was unknown. The position of one consti-tutively spliced intron of Drosophila
Tnnt coincides precisely with rat Tnnt3, leading Fyrberg et al. (1990) to
hypothesize that Tnnt predates the divergence of invertebrates and vertebrates. However, we found that both zebrafish and fugu Tnnt1 lack introns. We propose two
work-ing hypotheses to address this issue. First, Tnnt1 introns were lost in teleost lineage but gradually increased and expanded gene size in tetrapod lin-eage. In this case, fish Tnnt1 may represent the vertebrate Tnnt1 pro-totype. Similar evolutionary trends have also been proposed for bony fish rhodopsin (Fitzgibbon et al., 1995) and fugu SART1 (Bolland and Hewitt, 2001). Second, due to ge-nome duplication in fish (Postle-thwait et al., 1998; Taylor et al., 2001), zebrafish may have evolved two or more copies of Tnnt1, as occurred with Tnnt3. Searching for more Tnnt1 genes, and studying their genomic structure in zebrafish and other lower vertebrates will provide more insight into the molecular evolution of
Tnnt1.
Early Activation of Tnnt1 and
Tnnt2 During Somitogenesis
and Cardiogenesis
Slow muscles develop much earlier than fast muscles (van Swearingen and Lance-Jones, 1995, Stockdale, 1997) in the trunk, limb, and cranio-facial regions of vertebrates (Wacht-ler and Christ, 1992, Shu(Wacht-ler and Dal-rymple, 2001). The differential development of slow and fast mus-cles results in the asynchronous acti-vation of the slow and fast isoforms of muscle contractile proteins during somitogenesis. In tetrapods, the slow isoforms of muscle-specific proteins, such as the myosin heavy chain (Dhoot, 1986), troponin I (Kyprianou et al., 1997), and troponin T (Krishan et al., 2000, Wang et al., 2001), pre-dominate during early develop-ment. The fast isoforms of these pro-teins predominate later in development. Zebrafish Tnnt exhibits a similar pattern. The onset of ze-brafish Tnnt1 expression is 6 to 8 hr earlier than that of Tnnt3a and
Tnnt3b and also is much earlier than
that of several fast isoforms of mus-cle contractile proteins (Xu et al., 2000). The precocious expression of
Tnnt1 may help assemble the
tropo-myosin–troponin complex during early somitogenesis in both slow and fast muscles.
In zebrafish, the ontogenic expres-sion of muscle contractile proteins during cardiogenesis has not been
TABLE 2. Comparison of the Muscle Specificity of Three Troponin T Gene Transcripts in Early Developmental Stage Among Vertebrates
Tnnt1 Tnnt2 Tnnt3
Slow muscle Zebrafisha Mousec Chickenb,f
Chickenb
Mousec,d
Cardiac muscle Mousec,d Zebrafisha,e
Chickenb
Mousec
Fast muscle Zebrafisha Chickenb Zebrafisha,g
Chickenb Mousec Chickenb
Mousec,d Mousec aThis study. bYonemura et al., 2002. cWang et al., 2001. dKrishan et al., 2000. eSehnert et al., 2002. fNakada et al., 2000. gXu et al., 2000.
studied as much as expression of skeletal muscle proteins. Most of the cardiac-specific contractile proteins we examined were synchronously activated 17–18 hpf, whereas Tnnt2 was activated about 14 hpf. The precocious expression of Tnnt2 indi-cates it is crucial to assemble the tropomyosin–troponin complex dur-ing early cardiogenesis. This hypoth-esis is supported by zebrafish Tnnt2 mutants (Sehnert et al., 2002) and knock-down expression of Tnnt2 by morpholino oligos, which com-pletely stop the heart beat (Sehnert et al., 2002; Hsiao et al., unpublished observations).
Tnnt Expression in Fish Is More
Restricted Than in Tetrapods
In fetal to adult tetrapods, transi-tional expression of Tnnt transcripts in many different fiber-types is com-mon (summarized in Table 2). In chickens, Tnnt1 transcripts have been detected in slow, fast, and mixed muscles (Yonemura et al., 2002). In mice, Tnnt1 transcripts are expressed in all striated muscles dur-ing early fetal stages, but are re-stricted to slow muscle fibers during postnatal development (Krishan et al., 2000; Wang et al., 2001). Mouse
Tnnt2 transcripts occur in developing
skeletal muscles and the heart dur-ing the embryonic and fetal stages, but expression in skeletal muscles is selectively repressed after the late fetal stages (Wang et al., 2001). In zebrafish, only Tnnt1 is expressed in both slow and fast muscles. In con-trast, Tnnt2 and Tnnt3 exhibit high cardiac- and fast-muscle specificity during cardiogenesis and somito-genesis, respectively. This simplified muscle contractile protein expres-sion pattern also was reported in
Xe-nopus slow and fast troponin I
(Wark-man and Atkinson, 2002). Based on this evidence, we propose that the
cis-acting elements controlling
tis-sue- or stage-specific expression of
Tnnt in fish are simpler than their
tet-rapod counterparts. Transgenic ze-brafish lines controlled by Tnnt pro-moters should yield insights into the developmental regulation of muscle fiber specificity of Tnnt.
EXPERIMENTAL PROCEDURES
Extraction of Nucleic Acids
To obtain genomic DNA, an adult AB strain zebrafish (Danio rerio) was anesthetized by immersion in a tank of water containing 300g/ml
ethyl-m-aminobenzoate tricane methane
sulfonate (MS-222; Sigma). Its anal fin was removed, and genomic DNA was isolated from the fresh fin by us-ing the procedures described by Hsiao et al. (2001). To obtain total RNA, zebrafish embryos aged 24 to 72 hpf and Xenopus (Xenopus
lae-vis) embryos aged 1 to 7 dpf were
collected and immediately stored in liquid nitrogen. The whole, frozen embryos were homogenized with TRIzol reagent (Gibco BRL), and their total RNA was extracted according to the manufacturer’s instructions.
Molecular Cloning of Tnnt
cDNAs
The EST database maintained at NCBI (http://ncbi.nlm.nih.org) was searched for sequence annotations indicative of possible homology to zebrafish or Xenopus Tnnt. To clone full-length cDNA clones of Tnnt, 5 ⬘-and 3⬘-RACE were performed with EST clone-specific primers. First-strand cDNA was synthesized from 3 g of total RNA using a MATCH-MAKER Library Construction & Screening Kit (Clonetech). This method involved synthesis of full-length cDNA using CDS III oligo(dT) primer (5 ⬘-ATTCTAGAGGCCGA-GGCGGCCGACATG-d(T)30VN-3⬘),
coupled with (dC) tailing by Power-Script reverse transcriptase, followed by template switching and exten-sion with SMART III oligonucleotide (5 ⬘-AAGCAGTGGTATCAACGCAG-AGTGGCCATTATGGCCGGG-3⬘). The RNA/cDNA hybrid was then preampli-fied by long-distance PCR with the primer set 5⬘-PCR (5⬘-TTCCACC- CAAGCAGTGGTATCAACGCAGAGT-GG-3⬘) / 3⬘-PCR (5⬘-GTATCGATG- CCCACCCTCTAGAGGCCGAGG-CGGCCGACA-3⬘). For 3⬘-RACE, each preamplified PCR product was per-formed with primer sets Tnnt1-F (5 ⬘- AGAAGTAGCACCATGTGCGACAC-3⬘), Tnnt3b-F (5⬘-CTGTAGGTCTTGA GAGGGCCG-3⬘), Tnnt2-F (5⬘-TTC-CTCTGCATCTCTGCTGCGTGTC-3⬘), XTnnt2-F(5 ⬘-CCCAAGATTGTGGTGAA-AACATGTCTGATAC-3⬘), XTnnt3-F (5⬘- GAAAAAGTCTCACAACTGCTGC-GAC-3⬘) and 3⬘-PCR, respectively. For 5⬘-RACE, each preamplified PCR product was performed with primer sets of Tnnt1-R (5 ⬘-TTCAATTCGGT-TCTTCAACGCTAC-3⬘), Tnnt3b-R (5⬘-AATGTTCAGAGGCTTGCGTC-3⬘), Tnnt2-R (5 ⬘-AGGTAAAATCTATATT-GTTCAGTGAAATCTAACCG-3⬘), XT-nnt2-R (5 ⬘-ACTCTGTTTAAATGTAT-GTTTTCAACTCTAAC-3⬘), XTnnt3-R (5 ⬘-ATGAATTACAGGGCACAAC-GAACGC-3⬘) and 5⬘-PCR, respec-tively. Thirty-five cycles of PCR am-plification were performed by EXTaq DNA polymerase (Takara). Each cycle consisted of denatur-ation for 40 sec at 94°C, 1 min of annealing at 55°C, and 1 min of extension at 72°C. The last exten-sion step was extended for 10 min at 72°C. The PCR products were cloned into pGEM-T Easy vector (Promega) and both strands were sequenced on an ABI 373A se-quencer.
Bioinformatic Analysis of Tnnt
Sequences
Nucleotide sequences were trans-lated by using the sequence utilities available through the BCM Search Launcher interface (http://search-launcher.bcm.tmc.edu). Multiple se-quence alignment of the deduced amino acid sequences of unspliced
Tnnt were performed by using
Clus-talW (Thompson et al., 1994), and phylogenetic trees were con-structed by the neighbor-joining method (Pearson et al., 1999) through the DDBJ interface (http:// www.ddbj.nig.ac.jp/E-mail/clustalw-e. html). Confidence in the branch nodes was assessed with 1,000 boot-strap replicates. To identify potential
Tnnt1 genomic sequences in other
fish species, we used zebrafish Tnnt1 to BLAST search fugu draft genome sequences at the HGMP interface (http://fugu.hgmp.mrc.ac.uk). The intron– exon organization of poten-tial fugu Tnnt1 was predicted by GENSCAN (Burge and Karlin, 1997). The accession numbers of se-quences used in Figures 1 and 2 are
Tnnt1 from humans (Homo sapiens,
P13805), mouse (Mus musculus,
NP_035748), chicken (Gallus gallus, JC4970), brown trout (Salmo trutta, AAB58912), fugu (Fugu rubripes, FT: T005154), and zebrafish (AF425255);
Tnnt2 from humans (P45379), mouse
(BAB19881), chicken (TPCHTC),
Xe-nopus (AF467919), and zebrafish
(AF434187); and Tnnt3 from humans (NM_006757), mouse (NM_011620), turkey (Meleagris gallopavo,
AF210256), Xenopus (AY114144), ze-brafish (Tnnt3a, AAF78472 and
Tnnt3b, AF425741), Atlantic salmon
(S. salar, AAC24595); embryonic stri-ated muscle Tnnt (D85077) and adult body wall Tnnt (D50867) from an ascidian (Halocynthia rorezi).
Genomic Structure
Determination
A PCR-based strategy was used to elucidate the genomic organization of zebrafish Tnnt1 by using a Univer-sal Genomic Walker Kit (Clonetech). In brief, 3g of genomic DNA was digested with DraI, EcoRV, PvuI,
ScaI, or StuI and then ligated to
Ge-nome Walker Adapters. For 3 ⬘-genomic walking, the adapter-li-gated genomic DNA fragments were amplified by using the primer set Tnnt1-F/AP1 (5 ⬘-GTAATACGACT-CACTATAGGGC-3⬘). For 5⬘-genomic walking, the adapter-ligated genomic DNA fragments were first amplified by using the AP1 and
Tnnt1-R primer set, and then amplified
with the AP2 (5 ⬘-ACTATAGGGC-ACGCGTGGT-3⬘) and Tnnt1-R2 (5⬘-TGATGTGTCTCTTCAGGTTCGTC-3⬘) nested primer set (Fig. 3A). For Tnnt2,
Tnnt3a, and Tnnt3b, the primer pairs Tnnt2-F/Tnnt2-R4 (5
⬘-ATGTCCAGAG-GTTTGCGTCGATCAC-3⬘), Tnnt3a-F (5⬘-GGACATTGAGCAGCATTTCG-3⬘)/
Tnnt3a-R (5
⬘-CATGATGTCACACTATT-GTTTAGCAAACCT-3⬘), and Tnnt3b-F/
Tnnt3b-R were used to amplify
genomic DNA by long-distance PCR amplification with LATaq DNA poly-merase (Takara). Each of the thirty-five cycles of PCR amplification was comprised of denaturation for 30 sec at 94°C, 10 min of annealing and elongation at 68°C. The PCR products were cloned into pGEM-T Easy vector (Promega), and both strands were se-quenced on an ABI 373A sequencer.
RT-PCR Analysis
To detect the temporal expression of zebrafish Tnnt, RT-PCR was performed, as described in the cDNA cloning sec-tion, on the total RNA isolated from 9, 12, 15, 18, 24, 48, 72 hpf, and adult zebrafish (60 dpf). The primer pairs
Tnnt1-F/Tnnt1-R, Tnnt3a-F/Tnnt3a-R, Tnnt3b-F/Tnnt3b-R, Tnnt2-F3 (5
⬘-GTCT-GCACTTCGGCGGTTACA-3⬘)/Tnnt2-R, and sMyHC-F (5 ⬘-CAGGCTCCC-AGAATGAGATTGAAG-3⬘)/sMyHC-R (5⬘-AGCTTCATTCCTGCTGCGAG-3⬘) were used to amplify cDNA frag-ments of 420, 921, 273, 430, and 578 bp, respectively. To exclude false-positive bands, which could result from genomic DNA contamination, the total RNA was digested with RNase-free DNase I before RT-PCR. To confirm that the absence of a
Tnnt cDNA product was not due to a
deficiency in cDNA preparation, the 514-bp fragment of the zebrafish -actin (Kelly and Reversade, 1997) PCR product served as an RNA qual-ity control in each tissue sample.
Detection of Alternative RT-PCR
Products of Tnnt
To examine the potential alternative splicing products of zebrafish Tnnt2, total RNA isolated from 24, 48, 72 hpf, and adult zebrafish (60 dpf) was sub-jected to RT-PCR as described in the cDNA cloning section. The primer pairs Tnnt2-F/Tnnt2-R3 (5 ⬘-GGTGGAT-GTCATCAAAATCGACTC-3⬘), Tnnt2-F2 (5 ⬘-AAGATTCCAGATGGAGAAAGAG-TCG-3⬘)/Tnnt2-R2 (5⬘-ATGTCCAGAG-GTTTGCGTCGATCAC-3⬘), and Tnnt2-F3/Tnnt2-R were used to amplify cDNA fragments with predicted sizes of 318, 419 and 430 bp, respectively.
WISH
Zebrafish embryos were obtained by natural mating and were reared at 28.5°C in water containing 0.2 mM phenylthiocarbamide (Sigma) to suppress melanin formation. Em-bryos younger than 24 hpf were staged on the basis of somite num-ber, whereas older embryos were staged on the basis of hpf (Kimmel et al., 1995). Embryos were dechori-onated by pronase (10 g/ml) di-gestion and fixed overnight at 4°C in
PBS containing 4% paraformalde-hyde. Digoxigenin (DIG) -labeled RNA probes for Tnnt1, Tnnt2, Tnnt3a (Xu et al., 2000), Tnnt3b, sMyHC (AF425742), cmlc2 (Yelon et al., 1999), cmlc1 (Reiter et al., 2001),
vmhc (Yelon et al., 1999), and cTnC
(AF434188), were used to perform WISH according to procedures de-scribed by Thisse et al. (1993). Color reactions were carried out in alka-line phosphatase buffer using nitro-blue tetrazolium chloride and 5-bromo-4-chloro-3-indolyl phosphate (Roche) as substrates. Stained embryos were placed in 100% glycerol, and evalu-ated with a differential interference contrast microscope (DMR, Leica) with a color digital camera (COOL-PIX 996, Nikon) attached. For histo-logic examination, some stained embryos were embedded in paraf-fin and sectioned at 10-m intervals.
NOTE ADDED IN PROOF
During the revision of this manu-script, one publication has ap-peared that is consistent with our data. Antin et al. (2002) reported the precocious expression of Tnnt2 mRNA in early chick embryos. They found most gene transcripts of car-diac muscle contratile protein are detected after HH stage 11, while Tnnt2 mRNA is detected as early as HH stage 5.
ACKNOWLEDGMENTS
The authors thank Dr. Z. Gong, Na-tional University of Singapore, for providing us the Tnnt3a (⫽Tnnt) cDNA clone and Dr. W.M. Fu, Na-tional Taiwan University, for providing us the Xenopus embryos. The com-plete cDNA sequences of zebrafish
Tnnt1, Tnnt2, and Tnnt3b were
sub-mitted to GenBank under accession nos. AF425255, AF434187, and AF425741, respectively. The com-plete genomic sequences of ze-brafish Tnnt1, Tnnt2, Tnnt3a, and
Tnnt3b were submitted to the
Gen-Bank under accession nos. AF512524, AF512525, AY235720, and AF512526, respectively.
REFERENCES
Antin PB, Zhang W, Bales MA, Garriock RJ, Yatskievych TA. 2002. Precocious
ex-pression of cardiac troponin T in early chick embryos is independent of bone morphogenetic protein signaling. Dev Dyn 225:135–141.
Aparicio S, Morrison A, Gould A, Gilthorpe J, Chaudhuri C, Rigby P, Krumlauf R, Brenner S. 1995. Detecting conserved regulatory elements with the model genome of the Japanese puffer fish, Fugu rubripes. Proc Natl Acad Sci U S A 92:1684 –1688.
Barton PJ, Cullen ME, Townsend PJ, Brand NJ, Mullen AJ, Norman DA, Bhavsar PK, Yacoub MH. 1999. Close physical link-age of human troponin genes: organi-zation, sequence, and expression of the locus encoding cardiac troponin I and slow skeletal troponin T. Genomics 57:102–109.
Benoist P, Mas JA, Marco R, Cervera M. 1998. Differential muscle-type expres-sion of the Drosophila troponin T gene. A 3-base pair microexon is involved in visceral and adult hypodermic muscle specification. J Biol Chem 273:7538 – 7546.
Bolland DJ, Hewitt JE. 2001. Intron loss in the SART1 genes of Fugu rubripes and
Tetraodon nigroviridis. Gene 271:43–49.
Breitbart RE, Nadal-Ginard B. 1986. Com-plete nucleotide sequence of the fast skeletal troponin T gene. Alternatively spliced exons exhibit unusual interspe-cies divergence. J Mol Biol 188:313– 324.
Briggs JP. 2002. The zebrafish: a new model organism for integrative physiol-ogy. Am J Physiol Regul Integr Comp Physiol 282:R3–R9.
Bucher EA, Dhoot GK, Emerson MM, Ober M, Emerson CP. 1999. Structure and evolution of the alternatively spliced fast troponin T isoform gene. J Biol Chem 274:17661–17670.
Burge C, Karlin S. 1997. Prediction of com-plete gene structures in human genomic DNA. J Mol Biol 268:78 –94. Calvo S, Venepally P, Cheng J,
Buon-anno A. 1999. Fiber-type-specific tran-scription of the troponin I slow gene is regulated by multiple elements. Mol Cell Biol 19:515–525.
Charlet BN, Logan P, Singh G, Cooper TA. 2002. Dynamic antagonism between ETR-3 and PTB regulates cell type-spe-cific alternative splicing. Mol Cell 9:649 – 658.
Cooper TA, Ordahl CP. 1984. A single car-diac troponin T gene regulated by dif-ferent programs in cardiac and skele-tal muscle development. Science 226: 979 –982.
Cooper TA, Ordahl CP. 1985. A single car-diac troponin T gene generates embry-onic and adult isoforms via develop-mentally regulated alternate splicing. J Biol Chem 260:11140 –11148.
Dhoot GK. 1986. Selective synthesis and degradation of slow skeletal myosin heavy chains in developing muscle fi-bers. Muscle Nerve 9:155–164.
Endo T, Matsumoto K, Hama T, Ohtsuka Y, Katsura G, Obinata T. 1996. Distinct tro-ponin T genes are expressed in
embry-onic/larval tail striated muscle and adult body wall smooth muscle of as-cidian. J Biol Chem 271:27855–27862. Endo T, Matsumoto K, Hama T, Ohtsuka Y,
Katsura G, Obinata T. 1997. Temporal and spatial expression of distinct tropo-nin T genes in embryonic/larval tail stri-ated muscles and adult body wall smooth muscle of ascidian. Cell Struct Funct 22:197–203.
Farza H, Townsend PJ, Carrier L, Barton PJ, Mesnard L, Bahrend E, Forissier JF, Fiszman M, Yacoub MH, Schwartz K. 1998. Genomic organisation, alterna-tive splicing and polymorphisms of the human cardiac troponin T gene. J Mol Cell Cardiol 30:1247–1253.
Filatov VL, Katrukha AG, Bulargina TV, Gusev NB. 1999. Troponin: structure, properties, and mechanism of func-tioning. Biochemistry 64:969 –985. Fitzgibbon JA, Hope SJ, Slobodyanyuk SJ,
Bellingham J, Bowmaker JK, Hunt DM. 1995. The rhodopsin-encoding gene of bony fish lacks introns. Gene 164:273– 277.
Fyrberg E, Fyrberg CC, Beall C, Saville DL. 1990. Drosophila melanogaster. Tropo-nin-T mutations engender three distinct syndromes of myofibrillar abnormali-ties. J Mol Biol 216:657–675.
Gahlmann R, Troutt AB, Wade RP, Gun-ning P, Kedes L. 1987. Alternative splic-ing generates variants in important functional domains of human slow skel-etal troponin T. J Biol Chem 262:16122– 16126.
Hirao C, Hosoda K, Yonemura I, Miyazaki JI. 2001. Muscle subdivision: 1. Regula-tion of muscle genes. Dev Growth Dif-fer 43:S71.
Hsiao CD, Hsieh FJ, Tsai HJ. 2001. En-hanced expression and stable trans-mission of transgenes flanked by in-verted terminal repeats from adeno-associated virus in zebrafish. Dev Dyn 220:323–336.
Huang QQ, Jin JP. 1999. Preserved close linkage between the genes encoding troponin and troponin T, reflecting an evolution of adapter proteins coupling the Ca(2⫹) signaling of contractility. J Mol Evol 49:780 –788.
Huang QQ, Chen A, Jin JP. 1999. Genomic sequence and structural or-ganization of mouse slow skeletal mus-cle troponin T gene. Gene 229:1–10. Inoue A, Ojima T, Nishita K. 1996. Cloning
and sequencing of a cDNA for Aka-zara scallop troponin T. J Biochem 120: 834 –837.
Inoue A, Ojima T, Nishita K. 1998. Cloning and sequencing of a cDNA for tropo-nin T of Ezo-giant scallop striated mus-cle. Fish Sci 64:459 –463.
Jin JP, Lin JJC. 1988. Rapid purification of mammalian cardiac troponin T and its isoform switching: in rat hearts during development. J Biol Chem 263:7305– 7315.
Jin JP, Samanez RA. 2001. Evolution of a metal-binding cluster in the NH(2)-ter-minal variable region of avian fast skel-etal muscle troponin T: functional
diver-gence on the basis of tolerance to structural drifting. J Mol Evol 52:103– 116.
Jin JP, Huang QQ, Yeh HI, Lin JJ. 1992. Complete nucleotide sequence and structural organization of rat cardiac troponin T gene. A single gene gener-ates embryonic and adult isoforms via developmentally regulated alternative splicing. J Mol Biol 227:1269 –1276. Jin JP, Wang J, Zhang J. 1996. Expression
of cDNAs encoding mouse cardiac troponin T isoforms: characterization of a large sample of independent clones. Gene 168:217–221.
Johnston I, Cole N, Viera V, Davidson I. 1997. Temperature and developmen-tal plasticity of muscle phenotype in herring larvae. J Exp Biol 200:849 –868. Ju B, Xu Y, He J, Liao J, Yan T, Hew CL,
Lam TJ, Gong Z. 1999. Faithful expres-sion of green fluorescent protein (GFP) in transgenic zebrafish embryos under control of zebrafish gene promoters. Dev Genet 25:158 –167.
Kelly GM, Reversade B. 1997. Character-ization of a cDNA encoding a novel band 4.1-like protein in zebrafish. Bio-chem Cell Biol 75:623–632.
Kimmel CB, Ballard WW, Kimmel SR, Ull-mann B, Schilling TF. 1995. Stages of embryonic development of the ze-brafish. Dev Dyn 203:253–310.
Koban MU, Brugh SA, Riordon DR, Dellow KA, Yang HT, Tweedie D, Boheler KRA. 2001. Distant upstream region of the rat multipartite Na(⫹)-Ca(2⫹) exchanger NCX1 gene promoter is sufficient to confer cardiac-specific expression. Mech Dev 109:267–279.
Koumans JTM, Akster HA. 1995. Myogenic cells in development and growth of fish. Comp Biochem Physiol 110A:3–20. Krishan K, Morgan MJ, Zhao W, Dhoot GK. 2000. Slow troponin T mRNA in striated muscles is expressed in both cell type and developmental stage specific manner. J Muscle Res Cell Motil 21:527– 536.
Kusakabe R, Kusakabe T, Suzuki N. 1999. In vivo analysis of two striated muscle actin promoters reveals combinations of multiple regulatory modules re-quired for skeletal and cardiac muscle-specific gene expression. Int J Dev Biol 43:541–554.
Kyprianou P, Madgwick A, Morgan M, Krishan K, Dhoot GK. 1997. Expression pattern of troponin I and distinct alter-natively spliced developmental iso-forms of troponin T in vitro and in neo-natally denervated rat skeletal muscles. Basic Appl Myol 7:287–293. Marden JH, Fitzhugh GH, Wolf MR, Arnold
KD, Bowan B. 1999. Alternative splicing, muscle calcium sensitivity, and the modulation of dragonfly flight perfor-mance. Proc Natl Acad Sci U S A 96: 15304 –15309.
Mesnard L, Samson F, Espinasse I, Durand J, Neveux JY, Mercadier JJ. 1993. Mo-lecular cloning and developmental ex-pression of human cardiac troponin T. FEBS Lett 328:139 –144.
Muller F, Blader P, Strahle U. 2002. Search for enhancers: teleost models in com-parative genomic and transgenic analysis of cis regulatory elements. Bioessays 24:564 –572.
Myers CD, Goh PY, Allen TS, Bucher EA, Bogaert T. 1996. Developmental ge-netic analysis of troponin T mutations in striated and nonstriated muscle cells of
Caenorhabditis elegans. J Cell Biol 132:
1061–1077.
Nakada K, Miyazaki JI, Saba R, Hiraba-yashi T. 1997. Natural occurrence of fast- and fast/slow-muscle chimeric fi-bers in the expression of troponin T iso-forms. Exp Cell Res 235:93–99.
Nakayama M, Stauffer J, Cheng J, Ban-erjee-Basu S, Wawrousek E, Buonanno A. 1996. Common core sequences are found in skeletal muscle slow- and fast-fiber-type-specific regulatory ele-ments. Mol Cell Biol 16:2408 –2417. Ogut O, Jin JP. 1998. Developmentally
regulated, alternative RNA splicing-generated pectoral muscle-specific troponin T isoforms and role of the NH2
-terminal hypervariable region in the tolerance to acidosis. J Biol Chem 273: 27858 –27866.
Ogut O, Granzier H, Jin JP. 1999. Acidic and basic troponin T isoforms in mature fast-twitch skeletal muscle and effect on contractility. Am J Physiol 276:C1162– C1170.
Oota S, Saitou N. 1999. Phylogenetic re-lationship of muscle tissues deduced from superimposition of gene trees. Mol Biol Evol 16:856 –867.
Pearlstone JR, Johnson P, Carpenter MR, Smillie LB. 1977. Primary structure of rab-bit skeletal muscle troponin-T. Se-quence determination of the NH2
-ter-minal fragment CB3 and the complete sequence of troponin-T. J Biol Chem 252:983–989.
Pearson WR, Robins G, Zhang T. 1999. Generalized neighbor-joining: more re-liable phylogenetic tree reconstruc-tion. Mol Biol Evol 16:806 –816.
Perry SV. 1998. Troponin T: genetics, prop-erties and function. J Muscle Res Cell Motil 19:575–602.
Postlethwait JH, Yan YL, Gates MA, Horne S, Amores A, Brownlie A, Donovan A, Egan ES, Force A, Gong Z, Goutel C, Fritz A, Kelsh R, Knapik E, Liao E, Paw B, Ransom D, Singer A, Thomson M, Ab-duljabbar TS, Yelick P, Beier D, Joly JS, Larhammar D, Rosa F. 1998. Vertebrate genome evolution and the zebrafish gene map. Nat Genet 18:345–349. Reiter JF, Verkade H, Stainier DY. 2001.
Bmp2b and Oep promote early myo-cardial differentiation through their regulation of gata5. Dev Biol 234:330 – 338.
Ryan KJ, Cooper TA. 1996. Muscle-spe-cific splicing enhancers regulate inclu-sion of the cardiac troponin T alterna-tive exon in embryonic skeletal muscle. Mol Cell Biol 16:4014 –4023.
Sabry MA, Dhoot GK. 1991. Identification of and pattern of transitions of cardiac, adult slow and slow skeletal muscle-like embryonic isoforms of troponin T in de-veloping rat and human skeletal mus-cles. J Muscle Res Cell Motil 12:262–270. Schiaffino S, Reggiani C. 1996. Molecular diversity of myofibrillar proteins: gene regulation and functional significance. Phys Rev 76:371–423.
Schilling TF, Kimmel CB. 1997. Muscoskel-etal patterning in the pharyngeal seg-ments of the zebrafish embryo. Devel-opment 124:2945–2960.
Sehnert AJ, Huq A, Weinstein BM, Walker C, Fishman M, Stainier DY. 2002. Car-diac troponin T is essential in sarcomere assembly and cardiac contractility. Nat Genet 31:106 –110.
Shuler CF, Dalrymple KR. 2001. Molecular regulation of tongue and craniofacial muscle differentiation. Crit Rev Oral Biol Med 12:3–17.
Stockdale FE. 1997. Mechanisms of for-mation of muscle fiber types. Cell Struct Funct 22:37–43.
Tan X, Du J. 2002. Differential expression of two MyoD genes in fast and slow muscles of gilthead seabream (Sparus
aurata). Dev Genes Evol 212:207–217.
Taylor JS, Van de Peer Y, Braasch I, Meyer A. 2001. Comparative genomics pro-vides evidence for an ancient genome duplication event in fish. Philos Trans R Soc Lond B Biol Sci 356:1661–1679. Thisse C, Thisse B, Schilling TF, Postlethwait
JH. 1993. Structure of the zebrafish snail1 gene and its expression in wild-type, spadetail and no tail mutant em-bryos. Development 119:1203–1215. Thompson JD, Higgins DG, Gibson TJ.
1994. CLUSTAL W: improving the sensi-tivity of progressive multiple sequence alignment through sequence weight-ing, positions-specific gap penalties and weight matrix choice. Nucleic Ac-ids Res 22:4673–4680.
Thys TM, Blank JM, Schachat FH. 1998. Rostral-caudal variation in troponin T and parvalbumin correlates with differ-ences in relaxation rates of cod axial muscle. J Exp Biol 201:2993–3001. Thys TM, Blank JM, Coughlin DJ, Schachat
F. 2001. Longitudinal variation in muscle protein expression and contraction ki-netics of largemouth bass axial muscle. J Exp Biol 204:4249 –4257.
van Swearingen J, Lance-Jones C. 1995. Slow and fast muscle fibers are prefer-entially derived from myoblasts migrat-ing into the chick limb bud at different developmental times. Dev Biol 170:321– 337.
Venkatesh B, Gilligan P, Brenner S. 2000. Fugu: a compact vertebrate reference genome. FEBS Lett 476:3–7.
Wachtler F, Christ B. 1992. The basic em-bryology of skeletal muscle formation in vertebrates: an avian model. Semin Dev Biol 3:217–227.
Waddleton DM, Jackman DM, Bieger T, Heeley DH. 1999. Charactersation of Troponin-T from salmonid fish. J Muscle Res Cell Motil 20:315–324.
Wang J, Jin JP. 1997. Primary structure and developmental acidic to basic transition of 13 alternatively spliced mouse fast skeletal muscle troponin T isoforms. Gene 193:105–114.
Wang Q, Sigmund CD, Lin JJ. 2000. Iden-tification of cis elements in the cardiac troponin T gene conferring specific ex-pression in cardiac muscle of trans-genic mice. Circ Res 86:478 –484. Wang Q, Reiter RS, Huang QQ, Jin JP, Lin
JJ. 2001. Comparative studies on the expression patterns of three troponin T genes during mouse development. Anat Rec 263:72–84.
Warkman AS, Atkinson BG. 2002. The slow isoform of Xenopus troponin I is ex-pressed in developing skeletal muscle but not in the heart. Mech Dev 115:143– 146.
Waterman RE. 1969. Development of the lateral musculature in the teleost,
Brachydanio rerio: a fine structural
study. Am J Anat 125:457–493. Wilkinson JM, Moir AJG, Waterfield MD.
1984. The expression of multiple forms of troponin T in chicken fast muscle may result from differential splicing of a single gene. Eur J Biochem 143:47–56. Xu Y, He J, Tian HL, Chan CH, Liao J, Yan
T, Lam TJ, Gong Z. 1999. Fast skeletal muscle-specific expression of a ze-brafish myosin light chain 2 gene and characterization of its promoter by di-rect injection into skeletal muscle. DNA Cell Biol 18:85–95.
Xu Y, He J, Wang X, Lim TM, Gong Z. 2000. Asynchronous activation of 10 muscle-specific protein (MSP) genes during ze-brafish somitogenesis. Dev Dyn 219:201– 215.
Yamano K, Miwa S, Obinata T, Inui Y. 1991. Thyroid hormone regulates de-velopmental changes in muscle during flounder metamorphosis. Gen Comp Endocrinol 81:464 –472.
Yan Z, Serrano AL, Schiaffino S, Bassel-Duby R, Williams RS. 2001. Regulatory elements governing transcription in specialized myofiber subtypes. J Biol Chem 276:17361–17366.
Yelon D, Horne SA, Stainier DY. 1999. Re-stricted expression of cardiac myosin genes reveals regulated aspects of heart tube assembly in zebrafish. Dev Biol 214:23–37.
Yonemura I, Hirabayashi T, Miyazaki J. 2000. Heterogeneity of chicken slow skeletal muscle troponin T mRNA. J Exp Zool 286:149 –156.
Yonemura I, Mitani Y, Nakada K, Akutsu S, Miyazaki J. 2002. Developmental changes of cardiac and slow skeletal muscle troponin T expression in chicken cardiac and skeletal muscles. Zool Sci 19:215–223.