Venom phospholipases of Russell's vipers from Myanmar and eastern
India
—Cloning, characterization and phylogeographic analysis
☆
Inn-Ho Tsai
a,⁎
, Hsin-Yu Tsai
a, Ying-Ming Wang
a, Tun-Pe
b, David A. Warrell
ca
Institute of Biological Chemistry, Academia Sinica, and College of Life Sciences, National Taiwan University, Taipei 106, Taiwan
b
Department of Medical Research, Yangon, Myanmar
c
Nuffield Department of Clinical Medicine, John Radcliffe Hospital, Headington, Oxford OX3 9DU, UK Received 19 February 2007; received in revised form 22 April 2007; accepted 24 April 2007
Available online 5 May 2007
Abstract
Venoms of Russell's vipers (genus Daboia) are known for their deadly coagulopathic and other effects. We herein studied various isoforms of
venom phospholipases A
2(PLAs) from two Daboia species at their geographic boundary. From Myanmar Daboia siamensis venom (designated
as DsM), four PLAs (designated DsM-aI, aI', aII' and bI') were purified, and the cDNAs encoding two acidic (DsM-aI and aII) and two basic
PLAs (DsM-bI and S1) were also cloned from its venom-glands. DsM-S1 is identical to the major venom PLA of southern India Daboia russelii,
but the protein is absent from the venom. Additionally, four PLAs (designated DrK-aI, aII, bI and bII) were cloned from cDNA obtained from
venom glands of a Kolkata D. russelii, and the PLAs were purified from the pooled venom (designated as DrK). The acidic DrK-aI is the most
neurotoxic and lethal among these PLAs; DsM-aI which differs from DrK-aI by only the Phe2 substitution shows greatly reduced enzymatic
activity and lethality. Both acidic PLAs do not form dimeric complex with basic PLAs in the same venoms. DsM-bI' is neurotoxic and lethal but
its orthologous DrK-bI (97% identical to DsM-bI') is a much weaker toxin. Given the fact that most of the orthologous PLAs of DrK and DsM
share 97–100% sequence identity, Daboia vipers of Myanmar and Kolkata must be closely related. Molecular phylogenetic analyses on 30 venom
PLAs of Eurasian vipers' revealed co-evolution of five subtypes of venom PLAs in both Daboia and Vipera genera. Our results shed light on the
intra- and inter-species variations and structure
–function relationships of viperid venom PLAs.
© 2007 Elsevier B.V. All rights reserved.
Keywords: Venom phospholipase A2; Cloning and sequencing; Phylogenetic analysis; Geographic variation; Russell's viper (Daboia siamensis, Daboia russelii)
1. Introduction
Russell's vipers (Viperinae of the genus Daboia) are
important causes of morbidity and mortality in South and
Southeast Asia. Two species of Daboia, Daboia russelii
[1]
and Daboia siamensis
[2]
, have been identified. D. russelii is
mainly distributed in the Indian sub-continent while D.
siamensis is discontinuously spread over a wide area from
Myanmar to Taiwan and south to Indonesia. Clinical effects and
antigenicities of Daboia venoms vary throughout these
different geographic ranges
[2–8]
. Envenoming by D.
siamen-sis poses a remarkably severe problem especially in Myanmar
where it has been the country's 5th leading single cause of death
[3–5]
. Clinical observations also indicated higher case fatalities
and more severe hemorrhagic manifestations in Daboia
envenoming in Myanmar and eastern India than in other
regions
[2,6
–9]
.
Snake venom is the richest source of secreted
phospholi-pases A
2(PLA; EC 3.1.1.4), the Ca
+ 2-dependent enzymes that
hydrolyzes the 2-acyl ester of phosphoacylglyceride
[9,10]
.
Paralogous PLA variants in viperid venoms have been derived
from gene duplication and rapidly evolved to acquire
functional diversity
[11–13]
. Up to 65% of the venom proteins
of Daboia from Taiwan, Thailand, southern India and Pakistan
Abbreviations: DsM, Daboia siamensis (Myanmar); DrK, Daboia russelii (Kolkata); dPPC,L-dipalmitoyl phosphatidylcholine; PLA, Phospholipase A2;
HPLC, high performance liquid chromatography
☆GenBank accession numbers for the novel nucleotide sequences of venom
PLAs are DQ090654-7 for those from D. siamensis (Myanmar), and DQ090658-61 for those from D. russelii (Kolkata).
⁎ Corresponding author. POB 23-106, Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan. Tel.: +886 22 3620264; fax: +886 22 3635038.
E-mail address:bc201@gate.sinca.edu.tw(I.-H. Tsai).
1570-9639/$ - see front matter © 2007 Elsevier B.V. All rights reserved. doi:10.1016/j.bbapap.2007.04.012
are PLAs, many of them have been cloned or fully sequenced
[14–16]
. Notably, most of the lethal or neurotoxic PLAs found
in snake venom are basic proteins, but two acidic PLAs of high
lethal potency have recently been purified from Daboia
venoms of eastern India
[9]
and Myanmar
[17,18]
. Since
their complete sequences were not solved, the structure
–
activity relationships of both PLA-toxins were difficult to
understand.
The aim of the present study is to resolve structures and
functions of novel venom PLAs from Daboia subspecies and
better understand their geographic variations. We thus purified,
characterized and cloned venom PLA isoforms of Daboia
vipers from Myanmar (i.e. Burma) and Kolkata (eastern India),
at the boundary between D. russelii and D. siamensis ranges.
The results were compared with the venom PLA data of Daboia
from adjacent areas, including Pakistan, southern India, Sri
Lanka, Thailand, and Taiwan
[16–19]
. By phylogenetic
ana-lyses and sequence alignments of 30 Viperinae venom PLAs,
the classification and structure
–function relationships of these
PLAs are discussed.
2. Materials and methods
2.1. Venoms and other materials
D. siamensis venom samples from southern Myanmar were kindly given by Prof. R.D.G. Theakston, Liverpool School of Tropical Medicine (UK) while another pooled sample from northern Myanmar was obtained from Prof. Y. Y. Shu of Kuangxi Medical University, China. Meanwhile, live specimens of Daboia were also caught near Yangon (Myanmar) and Kolkata (eastern India), respectively. Fresh venom glands were dissected immediately from each snake after they were euthanized 48–60 h after venom collection. The glands were preserved for several weeks in the RNAlater solution (Ambion, USA) prior to RNA extraction.
Modifying enzymes, restriction enzymes and the pGEM-T vector were purchased from Promega Corp. (Madison, USA). SyntheticL-dipalmitoyl phos-phatidylcholine (dPPC) was purchased from Avanti Polar Lipids (Alabaster, AL, USA). Triton X-100, sodium deoxycholate, acetylcholine, and other chemicals were of reagent grade from either Merck (Darmstadt, Germany) or Sigma (St. Louis, USA).
2.2. Purification of venom PLA
Crude Daboia venom was dissolved in 0.1 M ammonium acetate buffer (pH 6.5) and then fractionated on a FPCL equipped with a Superdex G75 column (HR10/30, Pharmacia) pre-equilibrated with the same buffer, at room temperature. The PLA2-containing factions were pooled and freeze-dried. The
venom proteins were further purified by reversed-phase HPLC using a C18
column (4.5 × 250 mm, 10 μ, Vydac) equilibrated with 0.07% aqueous trifluoroacetic acid (TFA, solvent A), and eluted with a linear gradient of 20– 45% CH3CN containing 0.07% trifluoroacetic acid (solvent B). Each peak was
dried in a vacuum-centrifuge device (Labconco, USA). Concentrations of crude venom and PLA in solutions were determined spectrophotometrically at 280 nm, assuming an extinction coefficient of 1.5 at 1.0 mg/ml[14]and method of Bradford[20].
2.3. Protein sequences and masses
Purity of venom protein was assessed by SDS-PAGE and N-terminal sequencing. The N-terminal sequence of each purified protein was determined by an Applied Biosystems amino acid sequencer (Model Procise 492)[21]. Its molecular weight was analyzed by QSTAR XL nano-ESI mass spectrometer System (Applied Biosystems, Foster City, USA).
2.4. Cloning and sequencing
One Daboia specimen from each region was euthanized 2 days after venom extraction, at which point the venom glands were removed immediately for RNA extraction. The mRNA was prepared and the cDNA synthesis kits (Stratagene, USA) was used, as previously described[14]. Using venom gland cDNA as a template, the PCR[22]was conducted using SuperTaq DNA Polymerase with a pair of pairs of mixed-base oligonucleotide primers (sense primer 1: TCTGGATT-SAGGAGGATGA GG; antisense primer 2: GCCTGCAGRACTTAGCA), that were specifically designed based on conserved 3′ and 5′ untranslated regions of several homologous cDNA encoding other viperid venom PLAs [12,14]. In addition, a sense primer 3 (GCGGAGATGATCGTNAARATG) based on the amino acid sequence AEMIVK was used in conjunction with primer 2 in an attempt to specifically amplify Drk-aI' and DsM-aI'.
After treatment with polynucleotide kinase, the amplified DNA fragment was inserted into the pGEM-T easy vector (Promega Biotech, Wisconsin, USA). It was then transformed into Escherichia coli strain JM 109. Only the white transformants were picked-up and the cDNA clones were selected by restriction enzyme gel pattern. The DNA sequencing System model 373A and the Taq-Dye-Deoxy terminator cycle sequencing kit (PE Applied Biosystems, USA) were used to determine the cDNA sequences by dideoxynucleotide method[23]. Full amino acid sequences of venom PLAs were deduced from the nucleotide sequences of cDNA.
2.5. Enzymatic activities and pharmacological effects
PLA2activities were measured by stat titration method at 37 °C on a
pH-stat apparatus (Radiometer RTS 822, Copenhagen, Denmark). Released fatty acids, from 3 mML-dipalmitoyl phosphatidylcholine (dPPC) mixed either with an equal concentration of deoxycholate or two fold concentration of Triton X-100 in 0.10 M NaCl, were titrated at pH 7.4 with 6 mM NaOH. The initial rate was recorded for more than five min and corrected for non-enzymatic spontaneous rate. Specific activity was expressed asμmol of dPPC hydrolyzed per min per mg of the enzyme. In addition, 1.0% (w/w) fresh egg yolk in 0.15 M NaCl was used as substrate for comparing PLA activities by pH-stat titration methods.
Induction of edema by venom PLA on hind paw of anaesthetized Wistar rats (150–160 g body mass) was monitored up to 5 h by a plethysmometer, as described previously[12]. To test the lethal effects of crude venom and purified PLAs, ICR mice of 30–35 g body weight were used[9,24]. The venom proteins were dissolved in sterile PBS and centrifuged at 10,000×g to remove insoluble materials before injected to the mice. The medium lethal dose (LD50) was
estimated by intraperitoneal (i.p.) injection of graded doses of the protein to the mice. Effects of venom proteins on four mice in each group were examined; the lethality was estimated as the dosage-range which was able to kill about 50% of the test animals within 24 h.
2.6. Phylogenetic analyses
Amino acid sequences of venom PLAs from various species under Viperinae subfamily were retrieved by Blast search[25]and aligned along with those solved in the present study by AlignX in VectorNTI (Invitrogen, USA) program. Cladograms were constructed based on these sequences by neighbor-joining algorithm using program PHYLIP [26]; degree of confidence for the internal linage of the tree was determined by bootstrap methods[27].
3. Results
3.1. Purification and biochemical characterization
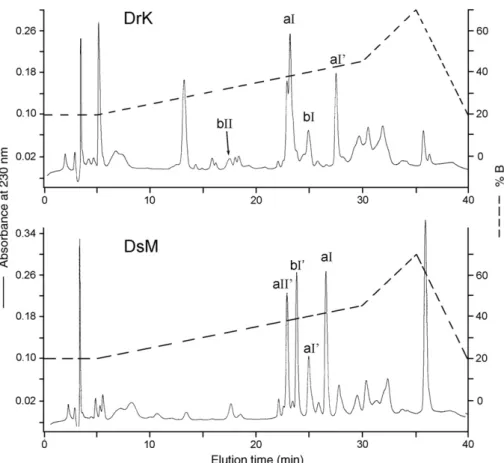
Patterns of gel filtration of the crude venoms by Superdex
G75 column on a FPLC system were shown in
Fig. 1
. The
PLA-containing fractions were lyophilized and further purified by
reversed-phase HPLC (not shown). Four venom PLAs could be
purified and identified from each of DrK and DsM (northern
Myanmar). Their protein masses, N-terminal sequences and
enzyme activities were also determined, as listed in
Table 1
. The
PLA isoforms were designated according to their pI values
deduced from the protein sequence (a, for acidic; b for basic), as
well as the N-terminal sequence homology. In addition, small
amount of the crude venom was fractionated by HPLC directly
(
Fig. 2
) to confirm the PLA-profiles obtained by the two-step
procedure.
The acidic DrK-aI is the most neurotoxic and lethal among
these PLAs, similar to a previous report
[9]
. Notably, DrK-aI
showed strong preference for the anionic substrate over the
zwitterionic substrate (
Table 1
). This is consistent with previous
findings for other venom presynaptic neurotoxins, including
β-bungarotoxin, crotoxin
[28]
and ammodytoxin
[29]
. Notably,
several PLA isoforms of DsM and DrK bear a Phe2 substitution
which was seldom observed in other venom PLAs, and they all
showed much lower catalytic activities than the PLAs
contain-ing Leu2 when both types of micellar substrates were used in
vitro (
Table 1
). When 1.0% (w/w) solution of egg yolk was used
as substrate, DrK-aI had a low specific activity of 23
μmol/min/
mg at 37 °C, while the activities of the Phe2-PLAs (DsM-aI and
Drk-aI') were too low to be measured.
Since cDNA had been obtained from venom glands of a D.
siamensis specimen collected near Yangon (southern
Myan-mar), PLAs were also purified from several venom batches of
D. siamensis specimens, which had been collected in the
same area (supplied by Prof. R.D.G. Theakston). HPLC
profiles, masses, and N-terminal sequences of the PLA isoforms
in these samples suggested that the PLAs are present in different
proportions and possibly show some individual variations. Each
of the venom sample contains four or five PLA-variants, mass
of the major PLAs have been determined to be 13607, 13810,
13830, and 14084 Da, respectively. Thus, DsM-PLA variant
with mass of 13830 appears to be present in Daboia venom of
southern Myanmar and DrK but not that of northern Myanmar
(
Table 2
).
3.2. cDNA cloning and protein sequence alignments
PCR has been carried out using specifically designed primers
for venom PLAs with the venom gland cDNAs as templates
[12,14]
. After the cDNAs were amplified and cloned, their
nucleotide sequences were determined to predict the PLA
protein sequences and pI values. Based on sequencing more
than 45 selected PLA clones for each species, sequences of four
distinct Asp49 PLAs, respectively, could be deduced from the
cDNA sequences of both DrK and DsM (
Table 2
). Their
complete amino acid sequences were then aligned with those of
most related venom PLAs obtained by BlastP search
[25]
. Three
putative categories of the Asp49-PLAs from Viperinae venoms
(i.e. acidic Asn1, basic Asn1, and Ser1) were aligned in
Fig. 3
A,
B and C, respectively.
Most of the cDNA deduced sequences (
Table 2
) could match
those of the purified PLA isoforms (
Table 1
), except that the
masses of purified DsM-aII' (13810) and DsM-bI' (14084) do
not match DsM-aII (13830) and DsM-bI (14055) predicted from
the cDNA sequences, respectively, though their N-terminal
sequences compromised. Further PCR experiments using
primers 3 and 2, followed by cloning and sequencing of
another 20 cDNAs, failed to produce any new clones encoding
DsM-aI'. It lays the possibility that the two PLA pairs, DsM-aII
and aII', DsM-bI and bI', are allelic proteins from different
individual snakes or geographic samples.
3.3. Lethal and edematous effects
DrK and DsM appear to evolved functionally different
PLAs, in spite of the fact that their PLA sequences are very
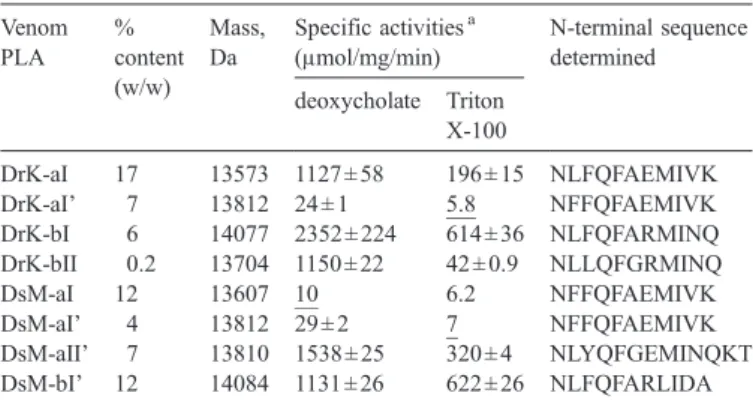
Table 1
Characterization of purified venom PLAs from DrK and DsM venoms Venom PLA % content (w/w) Mass, Da Specific activitiesa (μmol/mg/min) N-terminal sequence determined deoxycholate Triton X-100 DrK-aI 17 13573 1127 ± 58 196 ± 15 NLFQFAEMIVK DrK-aI' 7 13812 24 ± 1 5.8 NFFQFAEMIVK DrK-bI 6 14077 2352 ± 224 614 ± 36 NLFQFARMINQ DrK-bII 0.2 13704 1150 ± 22 42 ± 0.9 NLLQFGRMINQ DsM-aI 12 13607 10 6.2 NFFQFAEMIVK DsM-aI' 4 13812 29 ± 2 7 NFFQFAEMIVK DsM-aII' 7 13810 1538 ± 25 320 ± 4 NLYQFGEMINQKT DsM-bI' 12 14084 1131 ± 26 622 ± 26 NLFQFARLIDA Masses were determined by ESI-MS spectrometry.
a Enzymatic hydrolysis of dPPC were measured in the presence of
deoxycholate or Triton X-100 with 10 mM CaCl2 at 37 °C. Values shown
are median ± S.E. of results from three independent experiments, and underlined values are averages of two experimental results.
Fig. 1. Gel filtration of the crude venom of DrK and DsM. Crude venom was dissolved and injected into a Superdex G75 (HR10/30) column on a FPLC system. The elution was carried out with the equilibration buffer, 0.1 M ammonium acetate (pH 6.4), at a flow rate of 1.0 ml/min. Fractions of 1.0 ml were collected. The PLA-containing fractions (shown by bars) were pooled.
similar. Purified DrK-aI at doses between 0.06 and 0.10
μg/g
was lethal to about half of the mice in 24 h. In contrast, DsM-aI,
which differed from DrK-aI by only residue 2, had very low
enzymatic activity (
Table 1
) and was not so toxic to mice
(LD
50N3.0 μg/g). All the mice injected with DsM-aI survived
in our experiments but were hypo-locomotive the first few
hours. We also compared the edema-inducing effects of DsM-aI
and DrK-aI on rat hind paws. Both PLAs at the dose of 10
μg
effectively induced edema but DrK-aI had a stronger and faster
effect than DsM-aI. Relative swelling of paw was peaked in 4 or
5 h after the injection, and reached 38.5% and 30.3% for DrK-aI
and DsM-aI, respectively (
Fig. 4
).
While toxicity of DsM-aI was weak, the basic DsM-bI' in the
same venom was more neurotoxic and lethal and its toxicity was
not affected or enhanced by the presence of DsM-aI (
Table 3
).
Notably, DrK-bI has good enzyme activity but it is not as lethal
as DsM-bI' to mice although both PLAs share about 97%
sequence identity (
Fig. 3
B).
3.4. Phylogenetic analyses
Cladogram was established based on the amino acid
sequences of venom PLAs from selected Eurasia Viperinae
including most Vipera and Daboia. A basic venom
G6D49-PLA from Trimeresurus puniceus (an Indonesia pitviper) was
used as the out-group (
Fig. 5
). Remarkably, this robust tree of
Viperinae venom Asp49-PLAs shows that the PLAs with Asn at
the N-terminus (designated as N1) form a distinct cluster
separated from those with Ser at the N-terminus (designated as
S1). In the N1-PLA cluster, at least four subtypes of PLAs with
distinct N-terminal sequences are selectively present in various
Fig. 2. Purification of PLAs by reversed-phase HPLC. Solubilized crude venoms of DrK and DsM in solvent A were fractionated on a C18-Vydac HPLC column. The
PLA-containing peak was identified by ESI-MS and N-terminal sequencing and annotated (as shown inTable 1).
Table 2
cDNA data for venom PLAs of DrK and DsM PLA cloned Calculated mass (Da) pI No. of clones
(Signal peptide) and residues 1–10 DrK-aI 13573 4.6 3 (MRTLWIVAVCLIGVEG) NLFQFAEMIV DrK-aIIa 13830 4.9 4 (MRTLWIVAVCLIGVEG) NLYQFGEMIN DrK-bI 14076 8.7 5 (MRTLWIVAMCLIGVEG) NLFQFARMIN DrK-bII 13704 8.4 3 (MRTLWIVAVCLIGVEG) NLLQFGRMIN DsM-aI 13607 4.6 11 (MRTLWIMAVCLIGVEG) NFFQFAEMIV DsM-aIIa 13830 4.9 2 (MRTLWIVAVCLIGVEG) NLYQFGEMIN DsM-bI 14055 8.7 2 (MRTLWIVAMCLIGVEG) NLFQFARLID DsM-S1a 13625 8.4 2 (MRTLWIVAVCLIGVEG) SLLEFGKMIL
Isoelectric point (pI) and molecular mass were predicted from the deduced protein sequence.
a
Vipera and Daboia venom species (
Fig. 5
), including two acidic
subtypes with either F3 or Y3 (or T/S 3) substitution, and a
highly basic subtype containing A11K12 and a less basic
subtype containing K11M12.
4. Discussion
Contents of venom PLAs in DsM and DrK (28–38% of the
venom proteins by weight) are lower than those in Daboia
venoms from Thailand, China, Taiwan and Pakistan (55–65%
of the total proteins). Various effects elicited by the Daboia
PLAs have been reported, including neurotoxic
[14,17]
,
cytotoxic, myotoxic
[16]
, hypotensive, and/or anticoagulating
[30]
, and some PLAs may show more than one of the effects. In
the present study, four PLAs were purified from each of DrK
and DsM, and eight novel venom PLAs have been cloned and
fully sequenced (
Table 2
,
Fig. 3
). PLAs of all geographic venom
samples of Daboia are found to contain Asp49, and many of the
DrK and DsM PLAs contain rather unique Ala6 substitution.
We previously reported the N-terminal sequences of five
PLAs purified from D. siamensis (formerly Vipera r. siamensis)
venom purchased from Miami Serpentarium (Florida, USA)
[19]
. However, our later analyses of Daboia venom obtained
from Thai Red-Cross Society, Bangkok revealed that it
contained only the RV4/RV7 heterodimeric PLA (unpublished
results), just like Taiwan D. siamensis venom
[14]
. This was
supported by another cloning study using Thai D. siamensis
venom glands
[15]
. In fact, the N-terminal sequences of three
minor PLAs (S1-2, S1-1 and S3) in previous report
[19]
are
identical to those of DsM-aI, DsM-aII and DsM-bI,
respec-tively (
Table 1
). Thus, the D. siamensis venom from Miami
Serpentarium was heterogeneous and possibly contained 20%
of the venom from Myanmar.
Results in
Tables 1 and 2
are helpful for the identification and
matching of purified PLAs with their cDNA clones. However,
masses of purified DsM-aI' and of DsM-bI' cannot exactly
match those of DsM-aI and of DsM-bI (deduced from the
cDNA data). The discrepancies could be attributed to the fact
that we used venom glands of Daboia specimen from southern
Myanmar for cDNA cloning, but the pooled northern Myanmar
Daboia venom was used for protein analyses. We suspect that
the PLA pairs, DsM-aI and aI', DsM-bI and bI', are probably
allelic proteins from different geographic samples. However,
possibilities that we did not obtain all the transcripts of PLA
from DsM venom glands, or that some of purified PLAs have
undergone post-translational modification or processing, could
not be ruled out. In addition, Blast-search helped to detect
similar data recently reported by other researchers after our data
have been deposited. For examples, D. r. siamensis PLA-III
(AY303800) and PLA-II (AY286006) appear to be identical to
DsM-aI and DsM-bI, while D. r. russelii PLA-I (DQ365974)
and PLA-II (DQ365975) are identical to DrK-aI and DrK-bI,
respectively. Another clone (AY256974) in the databank was
possibly a hybrid of DsM-aII and aI, and the other (DQ365977)
appeared to be a hybrid of DrK-aI and bII, they possibly derived
from artifacts of PCR experiments.
Envenoming by both DsM and DrK was reported to be more
hemorrhagic than by Daboia specimens from other geographic
areas
[3–5,7,8]
, which is attributed to special expression of
haemorrhagic metalloproteinases in both DsM and DrK (results
to be published). Except for the absence of a DrK-bII homolog
in DsM (
Table 1
), all the deduced DsM-PLA sequences are
≥97% similar or identical to their orthologous PLAs in DrK
(
Fig. 3
), and a PLA-isoform identical to DsM-aI is also present
in low content in DrK. These venom similarities suggest that
Daboia vipers from Myanmar and eastern India are closely
related species and constitute a special lineage of the Daboia
populations. Results of previous studies on western India
Da-boia venom (supplied by Haffkine Institute, Mumbai) also
showed that an acidic PLA (FrIII-3, pI 4.2) had most potent
neuromuscular blocking action among the five PLAs purified
[31]
. However, venom differences between the eastern and
western India D. russelii have been reported
[7,8]
.
Morphological differences between the D. siamensis
popula-tions of Thailand and Myanmar
[32]
have been reported, and
heterologous antivenoms in four times greater dosages were found
to be necessary to neutralize toxic effects of each venom
[33]
. The
fact that DsM is more similar to DrK than to the D. siamensis
venoms from Thailand, China and Taiwan might be explained by
the notion that dispersal of Daboia
“out of India” probably was
Table 3
Lethal potencies of crude venom and purified PLAs on ICR mice
Venom or PLA Lethality, (i.p.)μg/g mouse body weight DrK-aI 0.06bLD50b0.13 DsM-aI LD50N3.0 DrK-bI LD50N3.0 DsM-bI' 0.13bLD50b0.20 DsM-aI + DsM-bI' (1:1) 0.40bLD50b0.50 DrK soluble venom 1.0bLD50b1.33 DsM soluble venom 0.3bLD50b0.4
Fig. 4. Oedema-inducing activities of DrK-aI and DsM-aI on rat paw. Time course of the swelling of rat hind-paw was followed after injection of 10μg of the PLA dissolved in 100μl PBS. Volume of the paw was measured by a plethysmometer, % paw swelling (relative to the volume measured 1 min after the injection) was the average from results of two independent experiments.
Fig. 3. Alignments of the amino acid sequences of Asp49 PLAs from Viperinae venom: (A) acidic N1-PLAs; (B) basic N1-PLAs; (C) S1-PLAs. Representative sequences were retrieved from BlastP search[25]. Single-letter codes of amino acids and the numbering system of Renetseder et al.[51]are used. Residues identical to those in the top line are denoted with dots, gaps are marked with hyphens. PLAs and the species (as the original taxonomic designation submitted) and their GenBank or SwissProt accession numbers are: C. cerastes Ccer, P21789; Vipera russelii (southern India) VRV-PL VIII, P59071; Vipera r. russelii (Pakistan) Vrr-R1, P81458; Vipera russelii formosensis RV-4 and RV-7, Q02471 and P31100; E. mafocmahonii Ema-PLA 2, P24294; Echis ocellatus Eo-A, P59171; Pseudocerastes persicus Cb-I, AAB36097, Cb-ICb-I, AAB36096; T. puniceus Tpu-G6D49, AAR14167; Vipera a. ammodytes Ammodytin I2, P34180, Ammodytoxin-A, B and C, P00626, P14424 and P11407; Vipera a. meridionalis Vipoxin-A, P04084, Vipoxin-B, P14420; Vipera aspis zinnikeri Vaspin-An, AF548351; Vipera b. berus V6, AY159811; Vipera palaestinae VP8 AAC78084.
hampered by the high mountain ranging from the eastern border of
Myanmar. This bio-geographical distribution has also been
proposed for frog and some other vertebrate groups
[34]
.
Based on the mitochondria DNA analyses
[35,36]
, the
genus Daboia is closely related to European and western Asian
Vipera and Macrovipera. Moreover, both Daboia and Vipera
are specifically equipped with venom procoagulating
compo-nents, i.e. Factor-X and Factor-V activators
[37,38]
. It is
interesting that their venom powder can be either white or
yellow color in appearance
[39]
, and the white venoms were
less necrotic or hemorrhagic than the yellow ones
[6]
. Five
subtypes of venom Asp49 PLA genes have been found for
geographic samples of Vipera a. aspis
[40,41]
and possibly
other Vipera
[42]
. The phylogenetic tree of PLAs in
Fig. 5
unravels not only the close relationship between the two genera
Daboia and Vipera but also parallel evolution of their venom
PLA paralogs. It has been suggested that gene duplications
followed by accelerated evolution of surface functional
domains have resulted in venom PLA diversity
[13,43]
.
Venom components may be differentially expressed according
to geological separation
[44]
, adaptation to prey-ecology
[45]
or other factors
[2]
.
A lethal acidic PLA designated as PFIIc' was previously
purified from eastern Indian Daboia venom
[9]
. DrK-aI
prob-ably is PFIIc' because they share identical N-terminal sequences
and high content in the venom (15% by weight), and are highly
lethal to mice. In contrast, the most lethal PLA of DsM is the
basic DsM-bI', which is present in equal abundance as DsM-aI
(
Table 3
). Notably, enzymatic activity and toxicity of DsM-bI'
were not affected by the addition of equal concentration of
DsM-aI (
Table 3
), thus DsM-aI does not play a chaperon role.
The venom content of DrK-bI is much lower than that of DrK-aI
(
Fig. 2
), they possibly do not associate into dimers either. It
is found that DrK-bI is much less lethal than DsM-bI or bI'
(
Table 3
) although they differed by only four residues (
Fig. 3
B).
Since Thr41 has been conserved in all the neurotoxic or myotoxic
PLAs listed in
Fig. 3
as well as crotoxin B and agistrodontoxin
[11]
while Ser41 is usually present in less toxic PLAs and
crotoxin A, it is speculated that the substitution of Thr41 might be
one of the reasons behind the lower toxicity of DrK-bI relative to
DsM-bI or bI'.
In both pancreatic and snake venom PLAs, Leu2 is highly
conserved
[46]
, and probably is important for binding of the
substrates fatty acyl chain. Mutagenesis of Leu2 to Trp2 in
pancreatic PLA resulted in 33-folds decrease of the enzymatic
rate
[47]
. DsM-aI differs from DrK-aI only at residue 2, the
40–45 folds reduction in the catalytic rates and the lethality
of DsM-aI relative to DrK-aI thus can be solely attributed
to its Phe2-substitution (
Table 1
). The hydrolytic products
of neurotoxic PLAs are suggested to be crucial for their
presynaptic toxicities
[48]
. Mutants of PLA-neurotoxins with
lower catalytic activities than the native form usually showed
decreased toxicities
[29,49]
. The N-terminal structures of
PLA have also been known to be critical for the neurotoxicities
[49,50]
. Thus, single amino acid substitution at strategic
position of a venom PLA toxin may greatly affect its biological
functions.
Having an identical N-terminal sequence to that of
daboiatoxin previously purified from Myanmar D. siamensis
venom
[17,18]
, DsM-aI is much less lethal than daboiatoxin
(
Table 3
). This discrepancy and the reason why the viper
expresses such inactivated DsM-aI remain puzzling. Although
DsM-aI showed significant edematous effect (
Fig. 4
), its other
functions remain to be investigated. Moreover, the sequence of
Fig. 5. Phylogenetic tree based on amino acid sequences of Viperinae venom Asp49 PLAs. The dataset includes full sequences of 30 PLAs from Viperinae venom, and a pitviper PLA (Tpu-G6D49) was used as out-group. Their pI and special N-terminal substitutions were shown in parentheses. Neurotoxic PLAs or PLA-subunits are marked with asterisks. Species names and accession numbers are: Pseudocerastes persicus Cb-II, AAB36096; Vipera a. aspis Ammodytins, Vaa-Am I1, AY159807 and Vaa-Am I2, AY158637; Vipera aspis zinnikeri Vaspin-Bz, AY158635, Vipera b. berus Vbb, P31854; Vipera palaestinae VP7, AAC78085. Values at the nodes indicate the percentage of 1000 bootstrap replicates.“D or V ” above branches denote origin of the PLA from Daboia or Vipera venom, respectively.
DrK-aI is 91% identical to that of RV7 from Taiwanese Daboia
siamensis venom (
Fig. 3
A). The substitutions of about ten
amino acid residues in RV7, including E7K, T70M and D89N,
Y120N, could transform this non-lethal chaperone-like subunit
[14]
to a highly lethal and neurotoxic DrK-aI, or vise versa.
These naturally occurring isoforms with minor structural
differences but great functional variations thus provide a new
platform for further investigation of structure-function
relation-ships of venom PLAs.
Acknowledgments
We are grateful to Prof. Antony Gomes (Univ. of Calcutta)
for his help in obtaining venom glands of Daboia from Kolkata.
We also thank Prof. Shu, Yu-Yen (Kuangxi Medical University,
China) and Prof. R.G. Theakston (Liverpool, UK) for the
precious gifts of Daboia venom from Myanmar.
References
[1] K. Adler, H.M. Smith, S.H. Prince, P. David, D. Chiszar, Russell's Viper: Daboia russelii not Daboia russellii, due to Classical Latin rules, Hamadryad 25 (2000) 83–85.
[2] D.A. Warrell, Geographic and intraspecies variation in the clinical manifestations of envenoming by snakes, in: R.S. Thorpe, W. Wuster, A. Malhotra (Eds.), Venomous Snakes, Clarendon Press, Oxford, U. K., 1997, pp. 189–204.
[3] Than-Tahn, R.A. Hutton, M.L. Khin-Ei-Han, Soe-Soe, Tin-Nu-Swe, R.E. Phillips, D.A. Warrell, Haemostatic disturbances in patients bitten by Russell's viper (Vipera russelli siamensis) in Burma, Br. J. Haematol. 69 (1988) 513–520.
[4] Than-Tahn, N. Francis, Tin-Nu-Swe, Myint-Lwin, Tun-Pe, Soe-Soe, Maung-Maung-Oo, R.E. Phillips, D.A. Warrell, Contribution of focal haemorrhage and microvascular fibrin deposition to fatal envenoming by Russell's viper (Vipera russelli siamensis) in Burma, Acta Trop. 46 (1989) 23–38.
[5] Tin-Nu-Swe, Tin-Tun, Myint-Lwin, Thein-Than, Tun-Pe, J.I. Robertson, B.J. Leckie, R.E. Phillips, D.A. Warrell, Renal ischemia, transient glomerular leak and acute renal tubular damage in patients envenomed by Russell's vipers (Daboia russelii siamensis) in Myanmar, Trans. R. Soc. Trop. Med. Hyg. 87 (1993) 678–681.
[6] A.V. Kumar, T.V. Gowda, Novel non-enzymatic toxic peptide of Daboia russelii (Eastern region) venom renders commercial polyvalent antivenom ineffective, Toxicon 47 (2006) 398–408.
[7] N.B. Prasad, B. Uma, K.G.S. Bhatt, T.V. Gowda, Comparative characterization of the variable properties of venoms of Russell's viper (Vipera russelli) venom from Indian peninsula, Biochim. Biophys. Acta 1428 (1999) 121–136.
[8] A.K. Mukherje, S.K. Ghosal, C.R. Maity, Some biochemical properties of Russell's viper (Daboia russelli) venom from Eastern India: correlation with clinico-pathological manifestation in Russell's viper bite, Toxicon 38 (2000) 163–175.
[9] A.K. Chakraborty, R.H. Hall, A.C. Ghose, Purification and characteriza-tion of a potent hemolytic toxin with phospholipase A2activity from the
venom of Indian Russell's viper, Mol. Cell. Biochem. 237 (2002) 95–102. [10] J.M. Danse, S. Gasparini, A. Menez, Molecular biology of snake venom phospholipases A2, in: R.M. Kini (Ed.), Venom Phospholipase A2
Enzyme: Structure, Function and Mechanism, J. Wiley and Sons, London, 1997, pp. 29–71.
[11] Y.H. Chen, Y.M. Wang, M.J. Hseu, I.H. Tsai, Molecular evolution and structure–function relationships of crotoxin-like and asparagine 6-containing phospholipases A2 in pit viper venoms, Biochem. J. 381
(2004) 25–34.
[12] I.H. Tsai, Y.M. Wang, Y.H. Chen, T.S. Tsai, M.C. Tu, Venom
phospholipases A2of bamboo viper (Trimeresurus stejnegeri): molecular
characterization, geographic variations and evidence of multiple ances-tries, Biochem. J. 377 (2004) 215–223.
[13] T. Ogawa, M. Kitajima, K.I. Nakashima, Y. Sakaki, M. Ohno, Molecular evolution of group II phospholipase A2, J. Mol. Evol. 41 (1995)
867–877.
[14] Y.M. Wang, P.J. Lu, C.L. Ho, I.H. Tsai, Characterization and molecular cloning of neurotoxic phospholipases A2 from Taiwan viper (Vipera
russelli formosensis), Eur. J. Biochem. 209 (1992) 635–641.
[15] I. Nuchprayoon, A. Sai-Ngam, S. Suntrarachun, J. Noiphrom, N. Pakmanee, L. Chanhome, S. Nuchprayoon, V. Sitprija, Molecular cloning of phospholipase A2 from a Thai Russell's viper venom gland cDNA
library, J. Med. Assos. Thail., Suppl. 84 (Suppl 1) (2001) S99–S105. [16] V.T. Gowda, J. Schmidt, J.L. Middlebrook, Primary sequence
determina-tion of the most basic myonecrotic phospholipase A2from the venom of
Vipera russelli, Toxicon 32 (1994) 665–673.
[17] Maung-Maung-Thwin, P. Gopalakrishnakone, R. Yuen, C.H. Tan, A major lethal factor of the venom of Burmese Russell's viper (Daboia russelii siamensis): isolation, N-terminal sequencing and biological activities of daboiatoxin, Toxicon 33 (1995) 63–76.
[18] Maung-Maung-Thwin, P. Gopalakrishnakone, R. Yuen, C.H. Tan, Synaptosomal binding of125I-labelled daboiatoxin, a new phospholipase A2neurotoxin from the venom of Daboia russelli siamensis, Toxicon 34
(1996) 183–199.
[19] I.H. Tsai, P.J. Lu, Y.C. Su, Two types of Russell's viper revealed by variation in phospholipases A2from venom of the sub-species, Toxicon 34
(1996) 99–109.
[20] M.M. Bradford, A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding, Anal. Biochem. 72 (1976) 248–254.
[21] M.W. Hunkapiller, L.E. Hood, Analysis of phenylthiohydantoins by ultrasensitive gradient high-performance liquid chromatography, Methods Enzymol. 91 (1983) 486–493.
[22] K.B. Mullis, F.A. Faloona, Specific synthesis of DNA in vitro via a polymerase-catalyzed chain reaction, Methods Enzymol. 155 (1987) 335–350. [23] T. Maniatis, E.F. Fritsch, J. Sambrook, Molecular Cloning, a Laboratory Manual, 2nd ed.Cold Spring Harbor Laboratory, Cold Spring Harbor, NY, 1989.
[24] G.P. Jayanthi, T.V. Gowda, Synergistic interaction of a protease and protease inhibitors from Russell's viper (Vipera russelli) venom, Toxicon 28 (1990) 65–74.
[25] S.F. Altschul, T.L. Madden, A.A. Schaffer, J. Zhang, Z. Zhang, W. Miller, D.J. Lipman, Gapped BLAST and PSI-BLAST: a new generation of protein database search programs, Nucleic Acids Res. 25 (1997) 3389–3402. [26] J. Felsenstein, PHYLIP: the PHYLogeny Inference Package, version
3.573. Computer Program Distributed by the U. of Washington, Dept. of Genetics, Seattle, U.S.A. , 1992.
[27] J. Felsenstein, Confidence limits on phylogenies: an approach using the bootstrap, Evolution 39 (1985) 783–791.
[28] F. Radvanyi, B. Saliou, C. Bon, P.N. Strong, The interaction between the presynaptic phospholipase neurotoxins beta-bungarotoxin and crotoxin and mixed detergent- phosphatidylcholine micelles. A comparison with non-neurotoxic snake venom phospholipases A2, J. Biol. Chem. 262
(1987) 8966–8974.
[29] T. Petan, I. Krizaj, M.H. Gelb, J. Pungercar, Ammodytoxins, potent presynaptic neurotoxins, are also highly efficient phospholipase A2
enzymes, Biochemistry 44 (2005) 12535–12545.
[30] E. Carredano, B. Westerlund, B. Persson, M. Saarinen, S. Ramaswamy, D. Eaker, H. Eklund, The three-dimensional structures of two toxins from snake venom throw light on the anticoagulant and neurotoxic sites of phospholipase A2, Toxicon 36 (1998) 75–92.
[31] H.C. Huang, C.Y. Lee, Isolation and pharmacological properties of phospholipases A2 from Vipem russelli (Russell's viper) snake venom,
Toxicon 22 (1984) 207–217.
[32] W. Wuster, S. Otsuka, R.S. Thorpe, A. Malhotra, Morphological variation in Russell's viper in Burma and Thailand, Herpetol. J. 2 (1992) 99–101. [33] Win-Aung, N. Pakmanee, O. Khow, C. Nambut, V. Sitprija, Cross neutralization of the lethal activities of Myanmar and Thai Russell's
viper venoms by Thai and Myanmar antivenoms, J. Nat. Toxins 10 (2001) 335–342.
[34] F. Bossuyt, M.C. Milinkovitch, Amphibians as indicators of early Tertiary “Out-of-India” dispersal of vertebrate, Science 292 (2001) 93–95. [35] P. Lenk, S. Kalyabina, M. Wink, U. Joger, Evolutionary relationships
among the true vipers (Reptilia: Viperidae) inferred from mitochondrial DNA sequences, Mol. Phylogenet. Evol. 19 (2001) 94–104.
[36] T. Garrigues, C. Dauga, E. Ferquel, V. Choumet, A.B. Failloux, Molecular phylogeny of Vipera Laurenti, 1768 and the related genera Macrovipera and Daboia (Gray, 1842), with comments about neurotoxic Vipera aspis aspis populations, Mol. Phylogenet. Evol. 35 (2005) 35–47.
[37] E. Siigur, A. Aaspollu, K. Trummal, K. Tonismagi, I. Tammiste, N. Kalkkinen, J. Siigur, Factor X activator from Vipera lebetina venom is synthesized from different genes, Biochim. Biophys. Acta 1702 (2004) 41–51.
[38] E. Siigur, A. Aaspollu, J. Siigur, Molecular cloning and sequence analysis of a cDNA for factor V activating enzyme, Biochem. Biophys. Res. Commun. 262 (1999) 328–332.
[39] R.W.P. Master, F. Kornalik, Biochemical differences in yellow and white venoms of Vipera ammodytes and Russell's viper, J. Biol. Chem. 240 (1965) 139–142.
[40] V. Jan, R.C. Maroun, A. Robbe-Vincent, L. De Haro, V. Choumet, Toxicity evolution of Vipera aspis aspis venom: identification and molecular modeling of a novel phospholipase A2heterodimer neurotoxin, FEBS Lett.
527 (2002) 263–268.
[41] I. Guillemin, C. Bouchier, T. Garrigues, A. Wisner, V. Choumet, Sequences and structural organization of phospholipase A2 genes from
Vipera aspis aspis, V. aspis zinnikeri and Vipera berus berus venom. Identification of the origin of a new viper population based on ammodytin I1 heterogeneity, Eur. J. Biochem. 270 (2003) 2697–2706.
[42] D. Kordis, A. Bdolah, F. Gubensek, Positive Darwinian selection in Vipera palaestinae phospholipase A2genes is unexpectedly limited to the third
exon, Biochem. Biophys. Res. Commun. 251 (1998) 613–619.
[43] R.M. Kini, Y.M. Chan, Accelerated evolution and molecular surface of venom phospholipase A2enzymes, J. Mol. Evol. 48 (1999) 125–132.
[44] W. Wuster, S. Otsuka, A. Malhotra, R.S. Thorpe, Population systematics of Russell's viper: a multivariate study, Biol. J. Linn. Soc. 47 (1992) 97–113.
[45] J.C. Daltry, W. Wuster, R.S. Thorpe, The role of ecology in determining venom variation in Malayan pit viper, C. rhodostoma, in: R.S. Thorpe, W. Wuster, A. Malhotra (Eds.), Venomous Snakes, Clarendon Press, Oxford, UK, 1997, pp. 155–172.
[46] D.L. Scott, S.P. White, Z. Otwinowski, W. Yuan, M.H. Gelb, P.B. Sigler, Interfacial catalysis: the mechanism of phospholipase A2, Science 250
(1990) 1541–1546.
[47] X. Liu, H. Zhu, B. Huang, J. Rogers, B.Z. Yu, A. Kumar, M.K. Jain, M. Sundaralingam, M.D. Tsai, Phospholipase A2 engineering. Probing the
structural and functional roles of N-terminal residues with site-directed mutagenesis, X-ray, and NMR, Biochemistry 34 (1995) 7322–7334. [48] M. Rigoni, P. Caccin, S. Gschmeissner, G. Koster, A.D. Postle, S.G.
Rossetto, C. Montecucco, Neuroscience: Equivalent effects of snake phospholipase A2neurotoxins and lysophospholipid-fatty acid mixtures,
Science 310 (5754) (2005) 1678–1680.
[49] M. Rouault, L.D. Rash, P. Escoubas, E. Boilard, J. Bollinger, B. Lomonte, T. Maurin, C. Guillaume, S. Canaan, C. Deregnaucourt, J. Schrevel, A. Doglio, J.M. Gutierrez, M. Lazdunski, M.H. Gelb, G. Lambeau, Neurotoxicity and other pharmacological activities of the snake venom phospholipase A2 OS2: the N-terminal region is more important than
enzymatic activity, Biochemistry 45 (2006) 5800–5816.
[50] I.H. Tsai, Y.M. Wang, Effect of site directed mutagenesis on the activity of recombinant trimucrotoxin, a neurotoxic phospholipase from Trimere-surus mucrosquamatus venom, Toxicon 36 (1998) 1591–1598. [51] R. Renetseder, S. Brunie, B.W. Dijkstra, J. Drenth, P.B. Sigler, A
com-parison of the crystal structures of phospholipase A2from bovine pancreas