Biochemical characterization and cloning of transglutaminases
responsible for hemolymph clotting in Penaeus monodon
and Marsupenaeus japonicus
☆
Maw-Sheng Yeh
a, Ling-Rong Kao
b, Chang-Jen Huang
b,c, Inn-Ho Tsai
b,c,⁎
a
Department of Food and Nutrition, Hung Kuang University, Sha Lu, Taiwan
b
Institute of Biochemical Sciences, National Taiwan University, Taipei, Taiwan
c
Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan
Received 29 October 2005; received in revised form 22 March 2006; accepted 7 April 2006 Available online 21 April 2006
Abstract
To investigate the shrimp blood clotting enzyme, a transglutaminase in the hemocytes of Penaeus monodon (abbreviated as TGH) was purified.
TGH is an abundant homodimeric cytosolic protein with 84.2 kDa subunits. It clotted shrimp plasma and incorporated fluorescent dansylcadaverine
into succinyl casein upon activation by CaCl
2in vitro. IC
50for the activation was 3 mM, which is below the shrimp plasma Ca
2+level. Showing
similar properties as other type II transglutaminase, TGH was particularly unstable after activation. MALDI-TOF/TOF mass-analyses of tryptic
peptides of P. monodon TGH confirmed its identity to STG I (AY074924) previously cloned. A possible allele of the other isozyme STG II
(AY771615) has also been cloned from the P. monodon cDNA and designated as PmTG. The predicted PmTG protein sequence is 58% similar to that
of STG I and 99.2% to that of STG II. Likewise, a novel enzyme Mj-TGH was purified and cloned from Marsupenaeus japonicus hemocytes. Results
of sequence alignment and phylogenetic analyses of these transglutaminases suggest that STG I and Mj-TGH are 83% identical and orthologous to
each other, while PmTG/STG II and a previously cloned M. japonicus transglutaminase (AB162767) are their paralogs. Protein of the latter two could
not be isolated, their regulated expression was discussed.
© 2006 Elsevier B.V. All rights reserved.
Keywords: Transglutaminase; Hemolymph coagulation; Regulation by Ca2+; cDNA sequence; Shrimp (Penaeus monodon and Marsupenaeus japonicus)
1. Introduction
Transglutaminases (EC 2.3.2.13) are known to play critical
roles in blood coagulation and other biochemical process
in-volving protein cross-linking or related super-structures.
En-zymes in this family catalyze calcium-dependent acyl-transfer
reactions between glutamine residues and lysine residues in
protein substrates. The role of Ca
2+is to induce enzyme
con-formational changes essential for substrate-binding and catalysis
[1
–4]
. The formation of intra-molecular or intermolecular 1-(
γ-glutamyl) lysine bonds lead to polymerized or insoluble
pro-teins. In addition, the enzymes mediate other post-translational
modification of proteins, e.g., deamidation and amine
incorpo-ration
[1]
.
The enzymes are widely distributed in tissues and body fluids
of animals. For example, blood coagulation factor XIIIa
cata-lyzes the cross-linking of fibrin monomer during blood
coag-ulation
[5]
, another transglutaminase toughens skin epidermis
and hair follicle epithelia
[6]
in vertebrates. The enzyme is also
involved in forming the fertilization plug in female rodents
[7]
.
Up to ten transglutaminases in various invertebrate species have
so far been reported
[8–11]
, including those responsible for
blood coagulation in Crustacean
[9,10]
. The enzymes were also
found to undergo acetylation
[1]
, phosphorylation, or
fatty-acylation
[12]
that probably regulates their subcellular
localiza-tion and catalytic activities. Some of them are involved in signal
transduction and apoptosis
[1,13]
.
Abbreviations: Mj, Marsupenaeus japonicus; Pm, Penaeus monodon; TGH, hemocyte transglutaminase; RFE, relative fluorescence enhancement
☆ The cDNA sequences of PmTG and Mj-TGH have been deposited in
GenBank with accession No. AF469484 and DQ436474, respectively. ⁎ Corresponding author. Fax: +886 2 23635038.
E-mail address:bc201@gate.sinica.edu.tw(I.-H. Tsai).
1570-9639/$ - see front matter © 2006 Elsevier B.V. All rights reserved. doi:10.1016/j.bbapap.2006.04.005
Tiger shrimp (Penaeus monodon) is an economically
impor-tant species cultured in Taiwan and southeastern Asia.
Coagula-tion of the hemolymph is part of the innate immune response of
Crustaceans, it prevents leakage of hemolymph from sites of
injury and dissemination of invaders such as bacteria throughout
the body. Much earlier, we have purified, characterized and cloned
the clottable protein in the hemolymph of P. monodon
[14]
.
Previously, two transglutaminases of P. monodon, STG I and STG
II were cloned and expressed as the shrimp clotting enzymes
[8,10]
. In the present study, we first reported purification and
biochemical characterization of major transglutaminases in
shrimp hemocytes (abbreviated TGH) and their stability and
regulation. Another transglutaminase from P. monodon
(nated as PmTG), and a novel hemocyte transglutaminase
desig-nated as Mj-TGH from Marsupenaeus japonicus (kuruma prawn)
were herein cloned and sequenced. Results of peptide sequencing
and peptide mass fingerprinting of the purified TGHs from both
shrimps enabled us to identify them with previously reported STG
I of P. monodon
[8]
and Mj-TGH, respectively. Moreover,
trans-glutaminases from invertebrates were subjected to phylogenetic
analysis, and the regulations of their expression were discussed.
2. Materials and methods
2.1. Shrimp hemocytes and the reagents
A total of 90 live specimens of P. monodon and 50 live specimens of M. japonicus were obtained from local markets. The hemolymph was withdrawn by a syringe of 2.5 ml with a needle of 23 G and 1.25 inch long containing 1/8 volume of anticoagulant (0.1 M sodium citrate, 0.4 M NaCl. It was centrifuged at 1600×g for 5 min at 4 °C, the supernatant was harvested for clotting assay. The pellet was washed twice with a cold buffer (1.5 mM EDTA, 0.4 M NaCl, 0.25 M sucrose in 50 mM Tris–HCl at pH 7.5). Casein and monodansylcadaverin were obtained from (Sigma Chem. Co., USA). Succinyl-casein was synthesized from casein and succinyl anhydride (Pierce, USA) as previously described [15].
2.2. Transglutaminase assay
Succinyl-casein (500 μg/ml) were mixed with 50 μM monodansylcada-verine in 50 mM Tris–HCl (pH 7.6) containing 5–10 mM dithioerythreitol and 6 mM CaCl2. This substrate solution was thermostated at 23 °C in a
cuvette before adding hemocyte lysate or purified P. monodon TGH. Incorporation of dansylcadaverin to succinyl-casein by TGH was followed by a fluorospectrometer (model MPF-4, Hitachi, Japan) at the excitation/ emission wavelengths of 360/500 nm, respectively [15]. Upon adding the TGH, the reaction started immediately in the presence of CaCl2 and no lag
phase was observed. Initial enzymatic rate within 1 min was measured to minimize errors due to enzyme instability, and relative fluorescence enhancement (RFE) was calculated based on the percent fluorescence increase relative to the blank. The shrimp plasma was diluted with equal volume of 50 mM Tris–HCl (pH 7.5) with 10 mM CaCl2 and then added
purified TGH at a final concentration of 14–35 μg/ml. The clotting time was monitored at 23 °C since the shrimps live in natural marine environment at 15–35 °C.
2.3. Purification and characterization of shrimp TGH
After washing, the hemocytes in pellet were lysed by 10 mM Tris–HCl buffer containing 1.5 mM EDTA and 0.1 mM ATP[16]and centrifuged to remove cell debris. The supernatant (i.e., hemocyte lysate) was loaded on a Fractogel TSK DEAE-650(M) column (1.4 × 7.5 cm) at 4 °C, which had been equilibrated in 50 mM Tris–HCl buffer (pH 7.5) with 1.5 mM EDTA and 0.1 mM ATP. The sample was eluted with a linear gradient of 0–0.3 M NaCl, fractions of 1 ml were collected and assayed for transglutaminase activity. Active fractions were pooled and further purified by a Sephacryl S-300 column (0.9 × 60 cm) at 4 °C, which was equilibrated and eluted with the 50 mM Tris–HCl buffer containing 1.5 mM EDTA and 0.1 mM ATP.
Molecular weight and purity of P. monodon TGH were studied by SDS-PAGE[17]. Native molecular weight of TGH was also estimated by elution volume on a gel filtration column, by interpolation from a calibration plot of “ratios of the elution volume to the void volume” against log molecular weight of the markers. In addition, matrix-assisted laser desorption/ionization-time-of-flight (MALDI-TOF) mass spectrometer (Model G2025, Hewlett-Packard, USA) was used to determine its accurate mass, in which 1μl of purified TGH (0.4 mg/ml) after dialysis against distilled water was added with 1μl sinapinic acid before crystallized under vacuum.
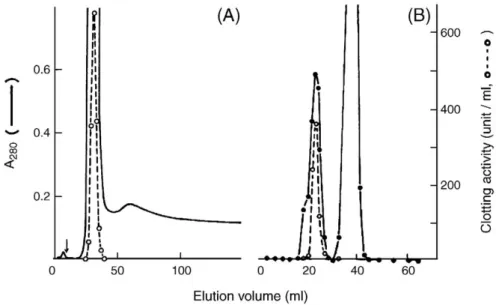
Fig. 1. Purification of TGH from P. monodon hemocyte lysate. (A) About 12 ml hemocyte lysate was separated by a Fractogel TSK DEAE-650 (M) column at 4 °C. The column was eluted with a gradient of 0–0.3 M NaCl in 50 mM Tris–HCl (pH 7.5), containing 1.5 mM EDTA and 0.1 mM ATP. Arrow indicated start of the gradient. (B) partially purified TG was subjected to gel filtration on a Sephacryl S-300 column. Fractions of 1 ml were collected at 4 °C, and transglutaminase activity (○–○) was assayed as detailed in Materials and methods.
2.4. Protein fragmentation and amino acid sequencing
Likewise, the enzyme Mj-TGH was purified from the hemocytes of another aqua-cultured species M. japonicus. Both TGHs (1 mg) of P. monodon and M. japonicus were denatured in 100μl of 6 M guanidine hydrochloride in 0.2 M Tris–HCl (pH 8.5) and reduced by dithiothreitol at 50 °C for 1 h. After adding excess iodoacetic acid and incubated at room temperature for 30 min, the S-carboxymethyl protein was desalted and lyophilized. Amino acid sequences were analyzed by an automatic sequencer (Model 477A, Applied Biosystem, USA).
For fragmentation, M. japonicus TGH was dissolved in 100μl of 70% formic acid with 0.2 mg of CNBr at room temperature for 24 h. In addition, it was digested with Lys-C endopeptidase (Promega, Madison, WI, USA) in 50 mM Tris–HCl (pH 8.0) at 37 °C for 20 h at an enzyme to substrate ratio of 1:50 (w/w). The reaction was stopped by adding dithiothreitol and heated at 95 °C for 5 min. Then the oligopeptides were purified by reversed-phase HPLC on a C18column,
then N-terminal sequences of three of the fragments.
2.5. Enzyme assay and effect of calcium and other reagents
For the fluorescence assay, CaCl2at a final concentration of 10 mM was
added to P. monodon TGH (77μg/ml) in 50 mM Tris–HCl, 1.5 mM EDTA, 0.1 mM ATP (pH 7.5) to start the reaction and initial rate (RFL/min) was
measured[15]. For clotting assay, clotting time was measured after adding the enzyme to a buffer containing 60μl supernatant of the centrifuged hemolymph and equal volume of 50 mM Tris–HCl (pH 7.5) containing 20 mM CaCl2.
To investigate its thermostability, purified TGH in the buffer (32μg/ml) were incubated at various temperature up to 50 °C. After a period of time, 30μl aliquots were withdrawn for the fluorescence assay. In addition, we have studied effects of pre-incubation of TGH with the chlorides of IIA group metal ion or iodoacetamide which modified active site CysSH of TGH. The remaining enzyme activity was assayed by the fluorescence assay.
2.6. cDNA synthesis and cloning
Standard procedures in molecular biology were used for preparation of plasmid DNA, restriction enzyme digestion, DNA agarose gel electrophoresis, DNA ligation, and the transformation of bacteria[18]. Total RNA was purified from tiger shrimp and kuruma prawn using the RNAzol B kit (Biotecx, Friendswood, TX, USA)[19]. The mRNA was purified using QuickPrepR Micro mRNA purification kit (Amersham Pharmacia, Freiburg, Germany). The first strand cDNA was synthesis using oligo(dT)-primer and random primers, and it was used as templates in subsequent polymerase chain reaction (PCR)[20].
To amplify the transglutaminase cDNAs of both species, we have used an antisense oligo(dT) and another degenerate primer designed from the conserved protein sequence (VPAHTIK) of M. japonicus TGH, i.e., sense 5′-GTNCCN
Fig. 3. (A) Correlation between clotting and amine-incorporating activities of purified TGH. After adding Tris–HCl buffer containing CaCl2 to the same
volume of shrimp plasma, the clotting was started from the addition of TGH (final TGH concentration ranged from 14 to 35μg/ml). In separate experiments, initial rates of dansylcadaverin incorporation were determined by fluorescence assay. (B) Apparent Ca2+dependency of P. monodon TGH activity. Purified TGH
(18μg/ml) was used in the fluorescence assay at various CaCl2concentrations in
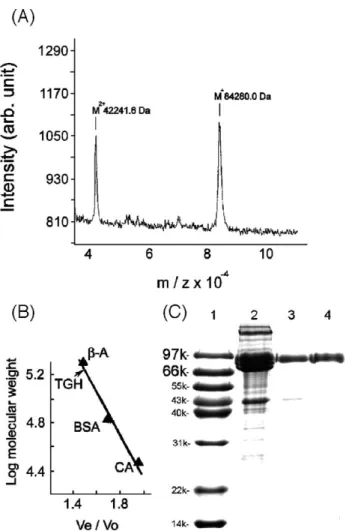
Tris buffer at 23 °C. Fig. 2. (A) Molecular mass of purified P. monodon TGH as determined by
MALDI-TOF mass spectrometry. (B) Estimation of the TGH molecular weight by gel filtration. Protein markers used were:β-amylase (β-A, 200 kDa), bovine serum albumin (BSA, 67 kDa) and carbonic anhydrase (CA, 29 kDa). Veand Vo
are elution-volumes for the protein marker and blue-dextran, respectively. (C) SDS-PAGE on 12% acrylamide gel. Lane 1, molecular weight markers; lane 2, hemocyte lysate; lanes 3 and 4 purified TGH with non-reducing and reducing sample buffer, respectively. Molecular weights of the markers were shown at left.
CANCA(TC)ACNAT(TCA) AA-3′. Except for the first and last cycles, each of the 40 cycles was: 94 °C for 30 s, 42 °C for 30 s, and 72 °C for 1 min. The first cycle included an extended denaturation time (2 min) during which polymerase was added while the last cycle had an extended (10 min) elongation period. The cDNAs of 578 bp and 1071 bp were thus produced from tiger shrimp and kuruma prawn, respectively. These cDNAwere purified and ligated into pGEM-T (Promega). Each clone was sequenced by using AmpliTaq DNA polymerase and fluorescent dideoxynucleotides on a DNA sequencer (Applied Biosystems model 310)[21].
2.7. Rapid amplification of cDNA ends (RACE)
Total RNA was isolated from both shrimps and poly (A)+ RNA was prepared. The first-strand cDNA was synthesized using a modified poly-T primer and 1μg of poly (A)+RNA. The second-strand cDNA was obtained
using an enzyme cocktail containing RNase H, DNA polymerase, and DNA ligase. Asymmetric adaptor primers (AP primers) were then ligated to both ends of the double-stranded cDNA. An aliquot of this cDNA was diluted 1:100 and subjected to PCR. The 5′ ends of mRNAs of PmTG and Mj-TGH were obtained by RACE methods using the Marathon cDNA amplification kit (Clontech, Palo Alto, CA, USA). Adaptor primers AP1 (external) and AP2 (internal) were supplied with the kit. RACE was performed with a 27mer sense primer (AP1) and antisense primers, 5′-GTAGTAGTCACCGTACGTCACTTCC-3′ for PmTG/ TGH and 5′-TGATAAGGTCATGGTCCACCAGCTTG-3′ for Mj-TGH, respectively. The second round of PCR was carried out with a nested 25mer sense primer (AP2) and nested antisense primers, 5′-CTTCGAGAAC GATTTCTCCCTCACG-3′ for TGH and 5′-GTAGTAGTCAGAGTAGTA
CACCTCC-3′ for Mj-TGH, respectively. Except for the first and last cycles, each cycle was: 94 °C for 30 s, 63 °C for 30 s, and 72 °C for 1.5 min. The first cycle included an extended denaturation time (2 min) during which polymerase was added while the last cycle had an extended (10 min) elongation period. After 40 PCR cycles, the products were cloned into pGEM-T vector (Promega) and sequenced[20,21].
2.8. RNA isolation from P. monodon tissues and RT-PCR
Total RNA was isolated from gill, heart, hemocyte, hepatopancreas, intestine, lymphoid organ, and muscle of P. monodon using the RNAzol reagent (Biotecx, Friendswood, TX, USA). RNAase-free DNase (Promega) was added to eradicate the genomic DNA contamination. First-strand cDNA was amplified from 2μg total RNA using 10 pmole oligo-dT primer in a 25-μl reaction containing 30 U RNasin (Promega), 1 mM dNTP, 10 mM dithiothreitol, and 300 U Superscript II (Life Technologies). After being incubated at 42 °C for 1 h in 1.5 mM MgCl2, 0.25 mM dNTP and 0.5 U ExTaq (Takara Shuzo Co.,
Japan), 2μl of the solution were added to 48 μl solution of the primers (designed from PmTG; forward, 5′-TGGAAGTGACGTACGGTGAC TACTAC G-3′ and reverse, 5′-GTTGTACAGCTGGCTGGAGTTGAAGG-3′). After amplified by PCR at the annealing temperature of 60 °C for 30 s (25–40 cycles), the 390 bp PCR products were examined on a 1.2% agarose gel.
2.9. Molecular phylogenetic analysis
Sequences of invertebrate transglutaminases were retrieved by BlastP[22]. Their amino acid sequences were aligned with the program CUSTAL W[23]. Phylogenetic tree was build by the program PHYLIP (http://www. evolution. genetics.washington.edu./phylip.html)[24]. Degree of confidence of the lineage at each node was determined by the bootstrap analyses of 1,000 replicates[25].
2.10. In-gel digestion of P. monodon TGH and peptide mass
finger-printing
After manually excised from the polyacrylamide gel, the electrophoresis band of P. monodon TGH was cut into pieces, which were dehydrated with acetonitrile for 10 min, vacuum dried, re-hydrated with 100 mM dithioerythritol in 25 mM NH4HCO3 (pH 8.5) at 37 °C for 1 h. They were subsequently
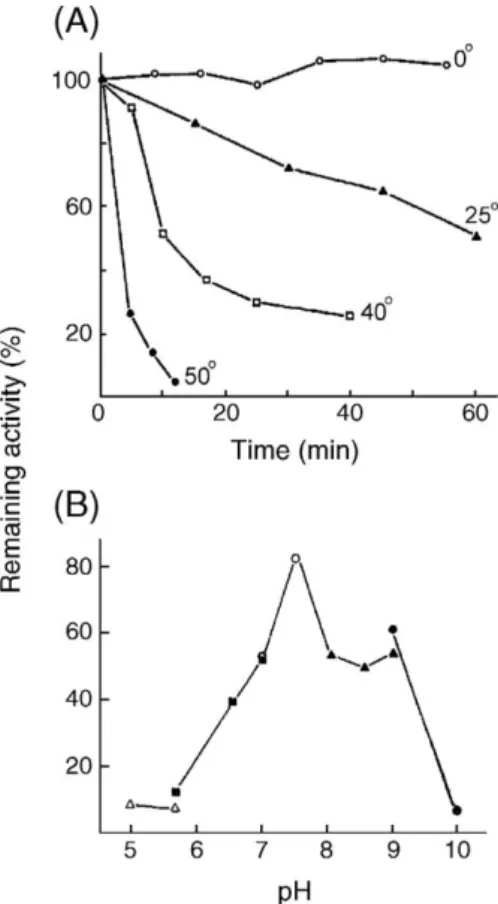
Fig. 4. (A) Thermostability of P. monodon TGH. The enzyme (0.3 mg protein/ml) in 50 mM Tris–HCl, 1.5 mM EDTA (pH 7.5) was pre-incubated at different temperature (0–50 °C) for 10 min. At indicated time, aliquots were withdrawn for amine-incorporation assay. (B) Stability of P. monodon TGH at various pH. The hemocyte lysate was diluted with 0.1 M of the following buffers with 0.4 mM EDTA at 4 °C and incubated 16 h before the amine-incorporation assay at 23 °C; sodium acetate (pH 5.0–5.8) (▵–▵), imidazole–HCl (pH 5.8–7.0) (
▪
–▪
), HEPES (pH 7.0–7.8) (○–○), Tris–HCl (pH 7.9–8.9) (▴–▴), sodium borate (pH 9.0–10) (U–U).Fig. 5. Effects of metal chlorides on the enzymatic activity of P. monodon TGH. The hemocyte lysate (0.4 mg protein/ml in 50 mM Tris–HCl, 0.1 mM ATP, 0.1 mM EDTA, pH 7.5) was preincubated with 5.3 mM metal chlorides at 4 °C (▵–▵, MgCl2;
▪
–▪
, BaCl2;□–□, SrCl2;▴–▴, CaCl2). Aliquots of samplewere withdrawn at the time indicated and amine-incorporation activity of TGH was measured at 23 °C.
alkylated with 65 mM iodoacetamide in NH4HCO3at 27 °C for 1 h in the dark,
washed twice with 50% acetonitrile in NH4HCO, dehydrated with acetonitrile
for 10 min and dried, then hydrolyzed with 25 ng modified trypsin (Promega, Madison, USA) in 25 mM NH4HCO3(pH 8.5) at 37 °C for 16 h. The
oligo-peptides were extracted twice with 50% acetonitrile containing 5% formic acid for 15 min and dried under vacuum. Their masses were analyzed with MALDI-TOF/TOF and matched by Mascot Search programs.
3. Results
3.1. Purification and characterization of P. monodon TGH
Enhancement of the substrate fluorescence upon
incorporat-ing dansylcadaverine into succinyl casein was measured to assay
Fig. 6. The cDNA and deduced amino acid sequence of PmTG. Sequences homologous to those of Mj-TGH peptides purified by HPLC are underlined. Potential N-glycosylation sites and the RGD motif are doubly underlined.
TGH during purification (
Fig. 1
). The yield was high (0.6 mg
TGH from 7.6 mg of the total hemocyte lysate proteins).
Increase of the specific activities of TGH after the two
purification steps was about 4-folds. The activity was hardly
detected in the shrimp plasma. The molecular mass of the
purified TGH was determined as 84280 ± 168 Da by
MALDI-TOF mass spectrometry (
Fig. 2
A). However, the molecular
weight measured by a gel filtration column was about 170–
180 kDa (
Fig. 2
B), and it was estimated to be about 85–90 kDa
by SDS-PAGE under both reducing and non-reducing condition
(
Fig. 2
C). Thus, the TGH most likely existed as non-covalent
homodimers, which could be dissociated in the presence of SDS
during electrophoresis. We found that TGHs of both shrimps
were N-terminal blocked, thus no results could be obtained by
automatic sequencing. Presumably like many other
transgluta-minases, the N-terminus was acetylated after removal of the
initiator-methionine residue. This acetylation possibly serves as
a signal for protein secretion
[1]
.
3.2. Enzymatic properties of P. monodon TGH and the effect of
Ca
2+We noticed that the amine-incorporating activity of purified
TGH and its hemolymph-clotting activity were well correlated
(
Fig. 3
A). Like other member of group II transglutaminase
family, Ca
2+is an essential cofactor or specific activator of
shrimp TGH. The half-maximum concentration of Ca
2+required
for the incorporation of dansylcadaverine of P. monodon TGH
Fig. 7. The cDNA and deduced protein sequence of Mj-TGH. Sequences confirmed by protein sequencing are underlined. The polyadenylation-signal, potential N-glycosylation sites and the RGD motif are doubly underlined.
was about 3 mM (
Fig. 3
B). Similar level of Ca
2+was required for
clotting of hemolymph by TGH. This is consistent with the
normal Ca
2+requirement of other transglutaminases
[1]
. Other
metal chlorides, e.g., 10–100 mM of SrCl
2, BaCl
2or MgCl
2,
could not substitute CaCl
2in both TGH assays (data not shown).
The strict dependency on Ca
2+is different from that for Factor
XIII, which can be activated by other divalent metal ions
although less effectively
[26]
.
Fig. 8. Amino acid sequence alignment of transglutaminase variants of tiger shrimp and kuruma prawn. Residues identical in all the four sequences are shown in black, and those identical in any three are marked in gray. RGD motifs are doubly underlined and the catalytic triads are shown in bold italics. Dots indicate putative calcium binding sites, and sequences matching those of the purified Mj-TGH oligopeptides are underlined.
P. monodon TGH appeared to be labile at temperature
N25 °C
and pH outside the range 7–9 (
Figs. 4
A and B). Lyophilization
or frozen–thaw could inactivate the enzyme by more than 50%.
However, more than 85% of the activity could retain for several
days if purified TGH or the hemocyte lysate was stored in the
presence of metal chelator (e.g., EDTA) at pH 7–9 at 0–4 °C, or
kept in buffer in the presence of EDTA and 35% (v/v) glycerol
below
−14 °C.
Like other type II transglutaminases, P. monodon TGH was
inhibited or inactivated by certain divalent metal ion (e.g., 10
–
20
μM ZnCl
2) which competed Ca
2+binding
[3,26,27]
,
thiol-reactive agents (e.g., organomercurials and iodoacetamide), or
2–10 mM histamine and other primary amines (data not shown).
Like other transglutaminases
[1,28]
, addition of 5 mM Ca
2+to
the medium greatly facilitated the inhibition of TGH by 3–
10
μM mercurial or 0.1 mM iodoacetamide (data not shown),
suggesting exposure of the TGH active site by a Ca
2+induced
conformational change. Notably, TGH activity was decayed in
minutes at mM concentrations of Ca
2+. Addition of EDTA could
stop the Ca
2+-induced decay immediately but could not reverse
it, and the presence of dithiothreitol in the enzyme buffer only
weakly protected this decay (data not shown). Moreover,
pre-incubation with other metal chlorides also inactivated the
enzyme although less effectively than CaCl
2(
Fig. 5
).
3.3. Cloning, sequencing and phylogenic analysis
Automatic sequencing of two oligopeptides obtained by
HPLC-purification of Lys-C digest of Mj-TGH showed their
amino acid sequences as MRQEYVLSDVGTVFVGSYP and
EXLTWEYKPPEGTRAXRVSILN, respectively. The sequence
of another purified CNBr cleavage product was YYTGVPAH
TIKRHDGELHLEPSQT. Degenerate PCR primers were
de-signed based on these Mj-TGH partial sequences. In addition to
the cDNA encoding STG I (AY074924), a 578-bp cDNA
en-coding the C-terminal part of a novel P. monodon
transglutami-nase (designated as PmTG) was produced by PCR. On the other
hand, a 1071 bp cDNA encoding the C-terminal part of Mj-TGH
was also amplified. The full-length cDNA of PmTG and Mj-TGH
were then completed by 5′ RACE. PmTG had a total length of
2,387 bp, including 36 bp of the 5′-untranslated region, an open
reading frame of 2,271 bp, and 81 bp of the 3′-untranslated
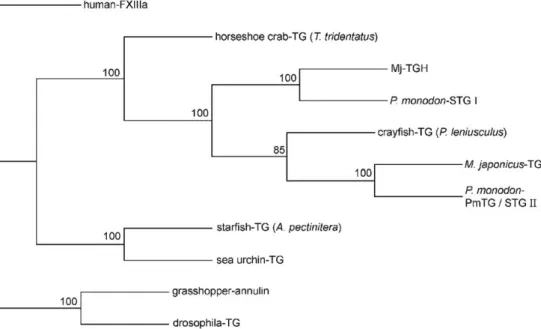
Fig. 9. Phylogenetic tree for invertebrate transglutaminases based on the amino acid sequences. Human factor XIIIa (NP-000120) was used as the out-group. Besides PmTG and Mj-TGH, the species and GenBank accession Nos. are: P. monodon STG I (AY074924) and STG II (AY771615), M. japonica-TG (AB162767), crayfish TG (AAK69205), horseshoe crab TG (A45321), starfish TG (BAB20439), sea urchin (CAD28789), drosophila TG (AAF52590) and grasshopper annulin (P52183). Bootstrap values are shown at the nodes.
Fig. 10. RT-PCR analysis of tissue expression of PmTG. Total RNA was extracted from various tissues of P. monodon. Specific primers were designed to amplify a 390-bp fragment of PmTG cDNA. PCR products at various cycles were analyzed by electrophoresis on 1.2% agarose gel containing ethidium bromide.
region. The putative initiating ATG codon, according to Kozak's
rule
[29]
, is at nucleotide 33 (
Fig. 6
). The open reading frame of
PmTG is predicted to encode a protein of 757 residues (pI 5.47)
and similar to STG II
[10]
. Its ORF differs from that of STG II by
15 nucleotides and resulted in the substitution of six amino acid
residues, i.e., L158
→F, T182→N, Q212→P, V495→A,
S499
→G, and E541→G. None of them are in the catalytic core
domain of the enzymes.
We also cloned Mj-TGH cDNA which contained 2996 bp
(included 54 bp of the 5′-untranslated region, an open reading
frame of 2274 bp, and 669 bp of the 3′-untranslated region). Its
authenticity was confirmed by sequence analyses of the
fragments obtained from CNBr cleavage and protease digestion
of the purified M. japonicus TGH. The putative initiating ATG
[29]
is at nucleotide 55, and its open reading frame encodes an
enzyme of 758 residues (pI 5.76) with seven potential
N-glycosylation sites and one RGD motif (
Fig. 7
). The predicted
Mj-TGH sequence is 83% and 59% identical to those of STG I
and STG II/PmTG, respectively (
Fig. 8
). PmTG and Mj-TGH
probably are both cytosolic, since they do not contain a typical
signal-peptide or trans-membrane domains.
Amino acid sequences of all the invertebrate
transglutami-nases so far solved were aligned and a phylogenetic tree has been
constructed to study their structural relationships (
Fig. 9
). All the
Crustacean enzymes are linked together in the tree and closer to
those of other marine arthropods than to those of insects.
Apparently, STG I and Mj-TGH are orthologous to each other.
3.4. RT-PCR analyses and tissue expression profile
The expression patterns of PmTG mRNA in various tissues of
P. monodon were analyzed by RT-PCR. As shown in
Fig. 10
, a
390-bp DNA fragment could be amplified in contrast to the
negative control. The expression levels were highest in
hemo-cyte and muscle, moderate in gill, intestine and lymphoid organ,
and lowest in heart and hepatopancreas.
3.5. In-gel digestion and mass analysis of TGH
Purified P. monodon TGH band in gel was digested with
modified trypsin before analyzed by
peptide-mass-fingerprint-ing. The masses matched nicely with those of the peptides
pre-dicted from P. monodon STG I
[8]
. We found at least 34 TGH
peptides matching those of STG I (
Table 1
), representing a total
of 396 amino acids or 52% (396/757) of the entire sequence and
evenly distributed.
Table 1
Mass match between the tryptic peptides of TGH and STG I (MSeand MScare experimental mass of TGH peptides and calculated mass of STG I peptides,
respectively)
Tryptic peptides of STG I position MSe MSc
FTNDLELCR 14–22 1167.57 1167.52 EDREFEIATELNEIADALSASK 24–45 2450.13 2450.18 EFEIATELNEIADALSASK 27–45 2050.95 2050.01 EFEIATELNEIADALSASKDPK 27–48 2390.14 2390.19 IVSVDFQPMYNAVNHHCELYELV 49–73 3047.36 3048.41 RGGLLSMVVK 82–91 1058.62 1058.63 GGLLSMVVK 83–91 902.47 902.53 LNQNVALASAQELK 92–105 1497.73 1497.82 FYFSFGSRPNVEK 106–118 1576.77 1576.77 GTQAHLVVTGK 119–129 1109.60 1109.62 IEKPHDDWDIR 132–142 1422.68 1422.69 VAAEVTLR 168–175 857.49 857.50 FDTHVYILFNPWNENDGTYLPEEK 186–209 2940.31 2940.36 MVNSNDDSGLLVGK 275–288 1447.63 1447.70 WDGEYEDGR 289–297 1125.42 1125.44 WDGEYEDGRAPAK 289–301 1492.64 1492.66 WLGSIEILEMYLR 302–314 1621.82 1621.85 VVTNFASAHDTNISLSIDEYFDEDGDQIDAADR 344–376 3642.54 3642.62 YVNPAGIR 377–384 888.41 888.48 YDVPFVLAEVNADVVR 444–459 1804.91 1804.94 LATQTSSVGR 473–482 1018.53 1018.54 AGPVATGGFFK 489–499 1050.48 1050.55 SDREDITWEYKPPEGTR 500–516 2077.94 2077.97 VSILNAAR 520–527 841.45 842.50 EDVTFQIEDR 544–553 1250.56 1250.58 TINVVMTCASMYYTGAK 577–593 1908.83 1909.86 HDGELHLEPSQTK 600–612 1489.70 1489.72 LVDHDLIK 627–634 950.47 951.54 VSFDNPIPLLLK 679–690 1354.76 1354.79 LDGPGLMRPK 696–705 1081.62 1082.59 MNFEMNLYPK 718–727 1285.55 1285.58 KAVEGTLIVQFNSR 728–741 1560.84 1560.86 AVEGTLIVQFNSR 729–741 1432.74 1432.77 ELCNLTGAK 742–750 1005.52 1005.48
4. Discussion
In shrimps, coagulation is initiated by activation of hyaline
cells which release its content including the clotting enzymes
[30
–32]
. P. monodon TGH effectively polymerizes shrimp
clottable proteins to form stabilized gel (
Fig. 3
). Notably, the
purified P. monodon TGH exists as weakly associated dimmers,
like the A subunit of mammalian FXIII which is release from
megakaryocytes as homodimers and also present as dimmers
when crystallized. However, the shrimp TGH is possibly active
as monomers like FXIII
[1]
. Dependent on seawater salinity, the
Ca
2+concentrations in tiger shrimp hemolymph range from 6.4
to 10 mM
[28]
. Thus, when TGH is released from hemocytes it
would be fully activated to initiate the coagulation (
Fig. 3
).
However, metal ions and high chloride ion
[2]
in the
hemo-lymph possibly reduce its enzymatic activity within minutes, as
a feedback regulation to avoid excess clotting.
The X-ray crystallographic structure
[33,34]
showed that the
catalytic domain (i.e., region 200
–550) of FXIIIa contains a
hydrogen-bonded Cys
–His–Asp triad similar to that found in
enzymes of the papain family
[35]
. The catalytic triad appears to
be also conserved in PmTG (and Mj-TGH), i.e., Cys330 (325),
His397 (392), Asp420 (415). Their Ca
2+-binding sites
presum-ably include the conserved Asn460, Asp462, Glu512, and
Glu517 (
Fig. 8
). With highly conserved region 350–353, His
358 (353) and Glu451 (466) in shrimp TGH may be associated
as in FXIIIa
[33]
. Many conserved Trp residues in the core
domain of TGH (
Fig. 8
) possibly are involved in stabilization of
the transition state during the enzymatic reaction
[36]
. It is
known that Ca
2+unmasks the active site Cys and widens the
protein substrate binding site
[1,37]
and the active site possibly
undergoes irreversible conformational changes. Being 35%
similar to FXIIIA, TGH is relatively unstable after activation at
ambient temperature and in the presence of bivalent metal ions
(
Fig. 5
). Instability of other transglutaminases has been reported
before
[2,38]
, but its exact mechanism remains to be explained
by 3D structural analyses of both the active and the inactive
enzymes or by mutagenesis study
[39]
.
The RGD (or Arg–Gly–Asp) motif, which is also present in
fibrinogen, is known to bind to integrins or specific membrane
receptors
[40]
. Positions of RGD motifs are remarkably
conserved in all invertebrate transglutaminases (
Fig. 8
) and
human keratinocyte
[8]
. This motif probably is essential for the
shrimp enzyme to bind integrins and form complex with adhesion
molecules
[1,41]
, and possibly trigger certain signal pathways,
e.g., RhoA activation and cell spreading
[42]
. Interestingly, the
RGD are all flanked by Pro and/or Gly residues (i.e.,
“proline
brackets”) and structurally constrained on the protein surface
[43]
, consistent with their role for protein
–protein interaction.
Both STG I and PmTG were transcribed in P. monodon
hemocytes. However, results of the peptide mass fingerprinting
of P. monodon TGH confirmed its identity with STG I (
Table 1
)
but not with PmTG or STG II. The molecular mass of P.
monodon TGH was determined as 84280 ± 168 Da. The
pre-dicted P. monodon TGH mass would be 84713 Da (based on the
STG I sequence,
Fig. 8
) assuming no disulfide bond was formed
and N-terminal acetylation took place after removal of Met 1, as
in the case of FXIIIa
[44]
. Alternatively, the mass would be
84285 Da if four residues (Met–Pro–Thr–Val) were removed.
Thus, the determined mass suggests that TGH possibly is
processed by deleting N-terminal residues before acetylation but
not glycosylated although they contain seven potential
N-glycosylation sites (
Figs. 6 and 7
).
Differed by merely six amino acid substitutions, PmTG and
STG II probably are alleles of the same locus with natural
polymorphism. The phylogenetic tree (
Fig. 9
) shows that the
shrimps have at least two paralogous transglutaminases, i.e.,
PmTG/STG II and STG I in P. monodon, and
Mj-transglutami-nase (AB162767) and Mj-TGH in M. japonica. Paralogs are
homologous proteins resulted from gene-duplication and they
usually co-expressed in the same organism, while orthologs are
those from the same gene lineage but expressed in different
organisms
[45]
. Previous immunodiffusion experiments showed
that antibodies prepared against of P. monodon clottable protein
reacted with M. japonicus clottable protein
[32]
. The coagulation
systems for this two genetically related species
[46]
presumably
are similar. Results in
Fig. 8
confirmed a high structural
similarity between the hemocyte enzymes (STG I and Mj-TGH)
of both species, but many distinct substitutions between STG I
and PmTG at their core regions (residues 200–500) suggesting
possible different substrate specificities. Protein sequence of a
previously cloned crayfish transglutaminase
[9]
appears to be
more close to that of PmTG or STG II (62%) than that of STG I
(55%), whether it is the only isozyme or the real functional TGH
of this fresh-water Crustacean remains to be studied.
The recombinant STG I from insect-baculovirus expression
system did not show clotting activities
[8]
, and recombinant
STG II was postulated to be the clotting enzyme of P. monodon
[10]
. However, STG I was expressed in Sf21 cells cultured at
26 °C and the cell lysate were frozen before assay, the
re-combinant STG I probably had lost most of its activity under the
culture conditions or freeze-and-thaw procedure. The instability
of STG I probably lead to the wrong conclusion that STG 1 is
not responsible for the hemolymph coagulation
[8]
. Moreover,
in vitro assays of recombinant STG II may also be misleading,
since the clot generated by recombinant STG II was less
branched and instable
[10]
, not like the clot formed naturally in
crayfish
[47]
and horseshoe crab
[2]
.
Based on microscopic observation, shrimp hemocytes
com-prised of hyaline cells, semi-granular cells, and granular cells
[48–50]
. Coagulation in shrimp is mediated by the circulating
hyaline cells
[31]
. We have cloned more cDNAs encoding
PmTG than those encoding STG I although STG I was the only
transglutaminase purified from P. monodon hemocytes. By
RT-PCR, we also found PmTG gene highly expressed in hemocytes
and muscle, but less expressed in other tissues (
Fig. 10
),
consistent with the expression patterns of STG II gene
[10]
.
Thus, PmTG mRNA is relatively abundant in the hemocytes,
while STG I gene is constitutive expressed at low-level in
various tissues
[8]
. The mRNA of a crayfish transglutaminase
(possibly orthologous to those of PmTG/STG II) was also
reported to be expressed in semigranular and granular cells
[9]
.
Since shrimp have an open circulatory system and hemocytes
are likely to rest onto different tissue surfaces, the Northern blot
results of PmTG (
Fig. 10
) need to be confirmed by in situ
hybridization. Previous in situ hybridization pointed out a low
mRNA level of STG I in circulating hemocytes but a high level in
mitotic cells of hematopoietic tissue
[8]
, and the RT-PCR
experiment also showed that STG I mRNA is decreased when
hemocytes matured and released into the hemolymph. Taken
together, these results suggest that transcription and translation of
STG I in the hemocytes possibly take place in early
develop-mental stages but down-regulated in later developdevelop-mental stages.
In conclusion, this is the first biochemical study on shrimp
hemocyte transglutaminases (TGH) responsible for hemolymph
coagulation. The purified shrimp TGHs are shown to be
relatively unstable after Ca
2+-dependent activation. We also
cloned two novel transglutaminases, PmTG from P. monodon
and Mj-TGH from M. japonicus, and confirmed the presence of
two transglutaminase genes in both species. By peptide mass
fingerprinting (
Table 1
) or sequencing of purified TGH peptide
fragments (
Fig. 8
), both STG I and Mj-TGH have been identified
to be the functional TGH in the two shrimp species. We failed to
isolate their paralogous enzymes, PmTG/STG II and another
transglutaminase previously cloned from M. japonicus, although
their mRNAs were expressed in shrimp hemocytes. The specific
role and regulated expression of these transglutaminases in
different Crustacean remain to be investigated.
Acknowledgements
We thank Yuh-Ling Chen for supplying peptide sequence
data of M. japonica transglutaminase and Chien-Hong Lu for
preparing the shrimp cDNA. MALDI-TOF/TOF analyses were
carried out by the Core Facilities for Proteomics and Structural
Biology Research, Institute Biological Chemistry, Academia
Sinica, Taiwan. This research was supported by grants from
Academia Sinica and Hung Kuang University, Taiwan, ROC.
References
[1] L. Lorand, R.M. Graham, Transglutaminases, cross-linking enzymes with pleiotropic functions, Nat. Rev., Mol. Cell Biol. 4 (2003) 140–156. [2] F. Tokunaga, M. Yamada, T. Miyata, Y.L. Ding, M. Hiranaga-Kawabata,
T. Muta, S. Iwanaga, A. Ichinose, E.W. Davie, Limulus hemocyte trans-glutaminase cDNA cloning, amino acid sequence, and tissue localization, J. Biol. Chem. 268 (1993) 252–261.
[3] C.S. Greenberg, P.J. Brickbichler, R.H. Rice, Transglutaminases: multifunction cross-linking enzymes that stabilize tissue, FASEB J. 5 (1991) 3071–3077. [4] D. Aeschlimann, M. Paulsson, Transglutaminase: protein crosslinking
enzymes in tissues and body fluid, Thromb. Haemost. 71 (1994) 402–415. [5] V.C. Yee, I. Le Trong, P.D. Bishop, L.C. Pedersen, R.E. Stenkamp, D.C. Teller, Structure and function studies of factor XIIIa by X-ray crystallography, Semin. Thromb. Hemost. 22 (1996) 377–384.
[6] E. Candi, G. Melino, G. Mei, E. Tarcsa, S.I. Chung, L.N. Marekov, P.M. Steinert, Biochemical, structural, and transglutaminase substrate properties of human lorcrin, the major epidermal cornified cell envelope protein, J. Biol. Chem. 270 (1995) 26382–26390.
[7] H.J. Lin, C.W. Lo, Y.H. Chen, Localization of the transglutaminase cross-linking site in SVS III, a novel glycoprotein secreted from mouse seminal vesicle, J. Biol. Chem. 277 (2002) 3632–3639.
[8] C.C. Huang, K. Sritunyalucksana, K. Soderhall, Y.L. Song, Molecular cloning and characterization of tiger shrimp (Penaeus monodon) transglutaminase, Dev. Comp. Immunol. 28 (2004) 279–294.
[9] R. Wang, Z. Liang, M. Hall, K. Soderhall, A transglutaminase involved in the coagulation system of the freshwater crayfish. Pacifastacus leniuscu-lus. Tissue localization and cDNA cloning, Fish Shellfish Immunol. 11 (2001) 623–637.
[10] M.Y. Chen, K.Y. Hu, C.C. Huang, Y.L. Song, More than one type of transglutaminase in invertebrates? A second type of transglutaminase is involved in shrimp coagulation, Dev. Comp. Immunol. 29 (2005) 1003–1016.
[11] F. Tokunaga, T. Muta, S. Iwanaga, A. Ichinose, E.W. Davie, K. Kuma, T. Miyata, Limulus hemocyte transglutaminase cDNA cloning, amino acid sequence, and tissue localization, J. Biol. Chem. 268 (1993) 262–268. [12] P.M. Steinert, S.Y. Kim, S.I. Chung, L.N. Marekov, The transglutaminase
enzyme is variably acylated by myristate and palmitate during differentiation in epidermal keratinocytes, J. Biol. Chem. 271 (1996) 26242–26250.
[13] G. Melino, M. Piacentini, Tissue transglutaminase in cell death: a downstream or a multifunctional upstream effector, FEBS Lett. 430 (1998) 59–63. [14] M.S. Yeh, C.J. Huang, J.H. Leu, Y.C. Lee, I.H. Tsai, Molecular cloning and
characterization of a hemolymph clottable protein from tiger shrimp (Penaeus monodon), Eur. J. Biochem. 266 (1999) 624–633.
[15] L. Lorand, O.M. Lockridge, L.K. Campbell, R. Myhrman, J. Bruner-Lorand, Transamidating enzymes II. A continuous fluorescent method suited for automating measurements of factor XIII in plasma, Anal. Biochem. 4 (1971) 221–231.
[16] S.C. Brenner, F. Wold, Human erythrocyte transglutaminase purification and properties, Biochim. Biophys. Acta 522 (1978) 74–84.
[17] U.K. Laemmli, Cleavage of structural proteins during the assembly of the head of bacteriophage T4, Nature 227 (1970) 680–685.
[18] J. Sambrook, E.F. Fritsch, T. Maniatis, Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1989.
[19] P. Chomczynski, N. Sacchi, Single-step method of RNA isolation by acid guanidinium thiocyanate–phenol–chloroform extraction, Anal. Biochem. 162 (1987) 156–159.
[20] K.B. Mullis, F.A. Faloona, Specific synthesis of DNA in vitro via a poly-merase-catalyzed chain reaction, Methods Enzymol. 155 (1987) 335–350. [21] F. Sanger, S. Nicklen, A.R. Coulson, DNA sequencing with
chain-terminating inhibitors, Proc. Natl. Acad. Sci. U. S. A. 74 (1977) 5463–5467. [22] S. McGinnis, T.L. Madden, BLAST: at the core of a powerful and diverse set of sequence analysis tools, Nucleic Acids Res. 32 (2004) W20–W25. [23] J.D. Thompson, D.G. Higgins, T.J. Gibson, CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice, Nucleic Acids Res. 22 (1994) 4673–4680.
[24] J. Felsenstein, PHYLIP: the PHYLogeny Inference Package, version 3.573. Computer program distributed by the U. of Washington, Dept. of Genetics, Seattle, 1992.
[25] J. Felsenstein, Confidence limits on phylogenies: an approach using the bootstrap, Evolution 39 (1985) 783–791.
[26] C.G. Curtis, K.L. Brown, R.B. Credo, R.A. Domanik, A. Gray, P. Stenberg, L. Lorand, Calcium-dependent unmasking of active center cysteine during activation of fibrin stabilizing factor, Biochemistry 13 (1974) 3774–3780.
[27] R.B. Credo, C.G. Curtis, L. Lorand, Alpha-chain domain of fibrinogen controls generation of fibrinoligase (coagulation factor XIIIa). Calcium ion regulatory aspects, Biochemistry 20 (1981) 3770–3778.
[28] R.P. Ferraris, Fe D. Parado-Estepa, J.M. Ladja, Effect of salinity on the osmotic, chloride, total protein and calcium concentrations in the hemolymph of the prawn Penaeus monodon (Fabricius), Comp. Biochem. Physiol. 83A (1986) 701–708.
[29] M. Kozak, An analysis of 50-noncoding sequences from 699 vertebrate messenger RNAs, Nucleic Acids Res. 15 (1987) 8125–8148.
[30] P. Kopacek, M. Hall, K. Sorderhall, Characterization of a clotting protein isolated from the plasma of the freshwater crayfish Pacifastcaus leniusculus, Eur. J. Biochem. 213 (1993) 591–597.
[31] S.A. Omori, G.G. Martin, J.E. Hose, Morphology of hemocyte lysis and clotting in the ridge back prawn, Sicyonia ingentis, Cell Tissue Res. 255 (1989) 117–123.
[32] M.S. Yeh, Y.L. Chen, I.H. Tsai, The hemolymph clottable proteins of tiger shrimp, Penaeus monodon, and related species, Comp. Biochem. Physiol. 121B (1998) 169–176.
[33] V.C. Yee, L.C. Pedersen, I. Le Trong, P.D. Bishop, R.E. Stenkamp, D.C. Teller, Three-dimensional structure of a transglutaminase: human blood coagulation factor XIIIa, Proc. Natl. Acad. Sci. U. S. A. 91 (1994) 7296–7300.
[34] B.A. Fox, V.C. Yee, L.C. Pedersen, I. Le Trong, P.D. Bishop, R.E. Stenkamp, D.C. Teller, Identification of the calcium binding site and a novel ytterbium site in blood coagulation factor XIII by X-ray crystallography, J. Biol. Chem. 274 (1999) 4917–4923.
[35] L.C. Pedersen, V.C. Yee, P.D. Bishop, I. Le Trong, D.C. Teller, R.E. Stenkamp, Transglutaminase factor XIII uses protease-like catalytic triad to crosslink macromolecules, Protein Sci. 3 (1994) 1131–1135. [36] S.E. Iismaa, S. Holman, M.A. Wouters, L. Lorand, R.M. Graham, A.
Husain, Evolutionary specialization of a tryptophane indole group for transition-state stabilization by eukaryotic transglutaminases, Proc. Natl. Acad. Sci. U. S. A. 100 (2003) 12636–12641.
[37] R. Casadio, E. Polverini, P. Mariani, F. Spinozzi, F. Carsughi, A. Fontana, P. Polverino de Laureto, G. Matteucci, C.M. Bergamini, The structural basis for the regulation of tissue transglutaminase by calcium ions, Eur. J. Biochem. 262 (1999) 672–679.
[38] J.E. Folk, P.W. Cole, Transglutaminase: mechanistic features of the active site as determined by kinetic and inhibitor studies, Biochim. Biophys. Acta 122 (1966) 244–264.
[39] V. Schroeder, E. Meili, T. Cung, P. Schmutz, H.P. Kohler, Characterisation of six novel A-subunit mutations leading to congenital factor XIII deficiency and molecular analysis of the first diagnosed patient with this rare bleeding disorder, Thromb. Haemost. 95 (2006) 77–84.
[40] E. Ruoslahti, RGD and other recognition sequences for integrins, Annu. Rev. Cell Dev. Biol. 12 (1996) 697–715.
[41] S.S. Akimov, A.M. Belkin, Cell surface tissue transglutaminase is involved in adhesion and migration of monocytic cells on fibronectin, Blood 98 (2001) 1567–1576.
[42] A. Salsmann, E. Schaffner-Reckinger, F. Kabile, S. Plancon, N. Kieffer, A new functional role of the fibrinogen RGD motif as the molecular switch that selectively triggers integrin alphaIIbbeta3-dependent RhoA activation during cell spreading, J. Biol. Chem. 280 (2005) 33610–33619. [43] R.M. Kini, Proline brackets and identification of potential functional sites
in proteins: toxins to therapeutics, Toxicon 36 (1998) 1659–1670. [44] K. Ikura, H. Yokota, R. Sasaki, H. Chiba, Determination of amino- and
carboxyl-terminal sequences of guinea pig liver transglutaminase: evidence for amino terminal processing, Biochemistry 28 (1989) 2344–2348.
[45] M. Kanehisa, S. Goto, S. Kawashima, A. Nakaya, The KEGG databases at GenomeNet, Nucleic Acids Res. 30 (2002) 42–46.
[46] S. Lavery, T.Y. Chan, Y.K. Tam, K.H. Chu, Phylogenetic relationships and evolutionary history of the shrimp genus Penaeus s.l. derived from mitochondrial DNA, Mol. Phylogenet. Evol. 31 (2004) 39–49. [47] M. Hall, R. Wang, R. van Antwerpen, L. Sottrup-Jensen, K. Soderhall, The
crayfish plasma clotting protein: a vitellogenin-related protein responsible for clot formation in crustacean blood, Proc. Natl. Acad. Sci. U. S. A. 96 (1999) 1965–1970.
[48] A. Tsing, J.M. Arcier, M. Brehelin, Hemocytes of Penaeid and Palaemonid shrimps: morphology, cytochemistry, and hemograms, J. Invertebr. Pathol. 53 (1989) 64–77.
[49] J. Rodriguez, V. Boulov, E. Mialhe, E. Bachere, Characterization of shrimp haemocytes and plasma components by monoclonal antibodies, J. Cell Sci. 108 (1995) 1043–1050.
[50] C.B.T. Van de Braak, R. Faber, J.H. Boon, Cellular and humoral char-acteristics of Penaeus monodon (Fabricius, 1798) haemolyph, Comp. Haematol. Int. 6 (1996) 194–203.