行政院國家科學委員會專題研究計畫 成果報告
臺灣地區從醫院病患分離出 Chlorhexidine 抗藥性金黃色葡
萄球菌菌株之分子流行病學、臨床特質及 chlorhexidine 抗
藥性機轉的研究
計畫類別: 個別型計畫 計畫編號: NSC93-2314-B-002-080- 執行期間: 93 年 08 月 01 日至 94 年 07 月 31 日 執行單位: 國立臺灣大學醫學院內科 計畫主持人: 盛望徽 共同主持人: 張上淳 計畫參與人員: 盛望徽, 王振泰, 張上淳 報告類型: 精簡報告 處理方式: 本計畫可公開查詢中 華 民 國 94 年 10 月 20 日
Emergence of Reduced Chlorhexidine Susceptibility in
Methicillin-Resistant Staphylococcus aureus at a Teaching Hospital
Jann-Tay Wang,* Wang-Huei Sheng,* Ching-Tzu Dai,* Chia-Ling Wu,*
Hui-Ming Tai,* Mei-Ling Chen,† Shan-Chwen Chang*
*
Department of Internal Medicine and †Laboratory Medicine, National Taiwan
University Hospital, Taipei, Taiwan
Jann-Tay Wang and Wang-Huei Sheng contributed equally to this article
Running title: MRSA with chlorhexidine resistance
Correspondance to: Dr. Shan-Chwen Chang, Department of Internal Medicine, National Taiwan University Hospital, No. 7 Chung-Shan South Road, Taipei 100, Taiwan e-mail: sc4030@ha.mc.ntu.edu.tw Tel.: 886-2-23123456 ext. 5401 Fax.: 886-2-23971412 Research Text words: 2637 Abstract words: 148
Abstract
We studied the chlorhexidine susceptibility, molecular typing, and the distribution of the qacA/B gene of methicillin-resistant Staphylococcus aureus (MRSA) isolates obtained in 1990, 1994, 1995, and 2002 at the National Taiwan University Hospital (NTUH). The prevalence rate of MRSA with a high minimum
inhibitory concentration (MIC) of chlorhexidine (≧ 4 μg/mL) was found to increase
rapidly during this 13-year period, from 3.3% in 1990 to 66.7% in 2002. Nearly all of the MRSA isolates with high chlorhexidine MICs carried the qacA/B gene and
belonged to a single molecular type. The prevalence rate of MRSA isolates with high chlorhexidine MIC correlates well with that of MRSA isolates belonging to the molecular type. We conclude that the prevalence rate of MRSA with high
chlorhexidine MIC increased rapidly during past 13 years at the NTUH, which was associated with an endemic strain, carrying the qacA/B gene, spreading in the hospital.
Key words: methicillin-resistant Staphylococcus aureus, MRSA, chlorhexidine, pulsed-field gel electrophoresis, qacA/B
Introduction
The first clinical isolate of methicillin-resistant Staphylococcus aureus (MRSA) at the National Taiwan University Hospital (NTUH) was found in 1981 (1). Thereafter, the nosocomial MRSA infection rate at NTUH increased rapidly, especially during 1990s and the prevalence rate of methicillin resistance in S. aureus isolates causing nosocomial infection at the NTUH remained at 60 – 80% during the past 10 years (2-3). The possible reasons for this high prevalence rate of MRSA nosocomial infection included poor adherence of healthcare workers (HCWs) to isolation
precautions leading to transfer of MRSA between patients via their colonized hands or the inanimate environment (4, 5), use and/or abuse of broad-spectrum antibiotics (3,6,7), increase of therapeutic modalities that breach the integrity of the
mucocutaneous system, and introduction of an endemic MRSA strain into the institute (5, 8-10).
To prevent the transfer of MRSA between patients via HCWs’ hands, antiseptics such as chlorhexidine were used for hand washing (11). Based on the history of antibiotic resistance, some authors predicted confidently and then proved the
emergence and wide spread of resistance to antiseptics in S. aureus, especially MRSA (12-16).
Resistance of S. aureus to antiseptics, including chlorhexidine, is conferred by two gene families, qacA/B and smr (14). The qacA/B gene confers a high-level resistance to antiseptics, and the smr gene family confers a low-level resistance. With the existence of chlorhexidine resistance in S. aureus, hand disinfectants based solely on chlorhexidine (such as Hibiscrub) might become limitedly effective against S.
aureus, which probably would not stop the spread of MRSA in a medical institute
using this kind of hand disinfectant for hand washing of health care workers (HCWs) (17).
Hibiscrub (Zeneca Ltd. Co., Macclesfield Cheshire England, UK), which contains chlorhexidine gluconate (4% w/v) as its major antibacterial component, has been an important disinfectant for hand washing for more than 20 years at the NTUH. The present longitudinal study was carried out to understand the changes of
susceptibility to chlorhexidine and the prevalence of MRSA isolates carrying the
qacA/B gene at a teaching hospital with a high prevalence rate of MRSA nosocomial
Methods
Hospital settings
NTUH is a major teaching hospital with a 2,200-bed capacity located in northern Taiwan. It offers both primary and tertiary medical cares.
Bacterial strains
A total of 240 MRSA isolates, with 60 isolates in each year, were randomly selected from the MRSA isolates obtained from clinical specimens, including blood, sputum, pus, discharge, and other body fluids in 1990, 1994, 1995, and 2002 at the NTUH. No duplicate isolates from a single patient or an outbreak were included.
Bacterial identification
Identification of S. aureus was based on colony morphology on trypticase soy agar supplemented with 5% sheep blood (BBL, Microbiology Systems, Cockeysville, MD, USA), Gram stain, and a positive Staphylase test (Oxoid Ltd, Basingstoke, England). The S. aureus isolates obtained were screened for methicillin resistance using the standard disk diffusion method, with use of Mueller-Hinton agar (BBL, Microbiology Systems) and 1 μg oxacillin disk after incubation for 24 hours at 35℃ (18).
Susceptibility test
Chlorhexidine minimum inhibitory concentration (MIC) was determined by agar dilution method as described by the National Committee for Clinical Laboratory Standards (NCCLS) (19). The chlorhexidine MIC for MRSA isolates carrying the
qacA/B gene is typically equal to or greater than 4 μg/mL according to previous
reports (15, 20, 21). Therefore, an isolate with a high chlorhexidine MIC was defined in this study as having a MIC of chlorhexidine equal to or greater than 4 μg/mL.
Pulsed-field gel electrophoresis (PFGE)
Molecular typing of all 240 MRSA isolates was done using PFGE according to the methodology described in details previously (22). The banding pattern of the PFGE results was interpreted according to the criteria suggested by Bannerman et al and Jorgensen et al (23, 24). Various types and subtypes of the PFGE pattern were named as those we used in our previous studies (22).
Detection of the qacA/B gene
Bacterial DNA was extracted using the InstaGeneTM matrix (Bio-Rad
Laboratories, Hercules, CA) according to the manufacturer’s protocol. Briefly, an isolated bacterial colony was suspended in 1 mL of autoclaved water in a microfuge tube and then centrifuged for 1 minute at 6,000 g. The supernatant was removed. The
pellet was then added into 200 μL of InstaGeneTM
matrix (Bio-Rad) and vortexed, followed by heating at 56℃ for 15 minutes. The sample was vortexed again and heated at 100℃ for 8 minutes and then centrifuged to pellet the matrix. The
supernatant was used in subsequent polymerase chain reactions (PCR). The following primers were used for the amplification of the qacA/B genes: the forward primer 5’-CTATGGCAATAGGAGATATGGTGT, and the reverse primer
5’-CCACTACAGATTCTTCAGCTACATG (14). PCR was performed as described elsewhere (14). DNA sequences of the PCR products were determined by Applied Biosystem 377 DNA sequencer (PE-ABD, Foster City, CA).
Results
Chlorhexidine susceptibility
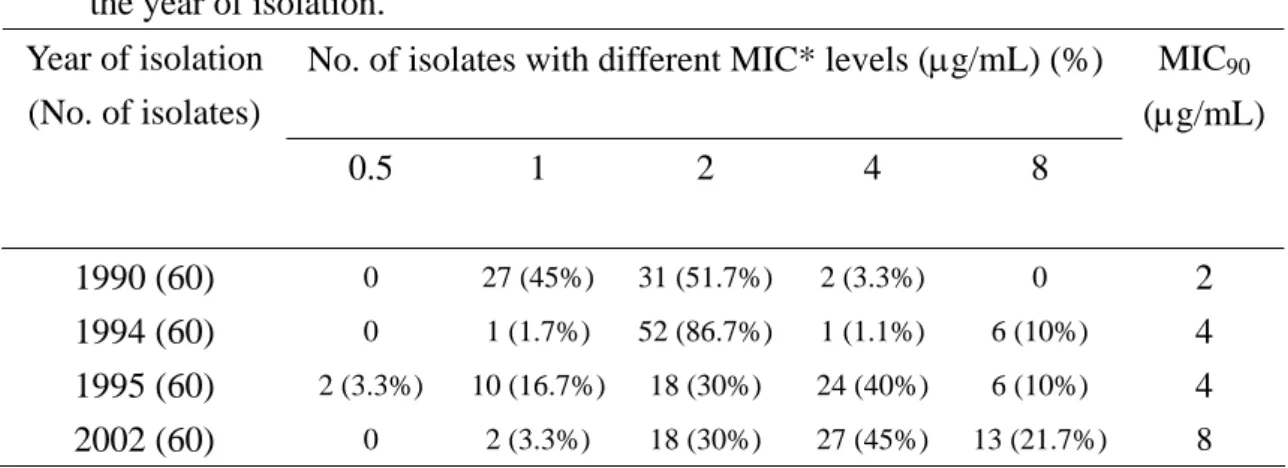
The chlorhexidine MICs of the 240 MRSA isolates collected from different
years are shown in Table 1. The MIC90 increased from 2 μg/mL in 1990 to 8 μg/mL in
2002. The prevalence rate of MRSA isolates with high chlorhexidine MIC (≧ 4 μg/mL) increased markedly during the study period, from 3.3% to 66.7%.
PFGE patterns
The PFGE banding patterns of the 60 MRSA isolates collected in 1990 revealed great diversity, with no predominant pattern evident. In contrast, analysis of the banding pattern of the 60 MRSA isolates isolated in 1994 revealed the predominance of the type B strain. This strain comprised 31 of the 60 isolates (51.7%), with type A and type C (consist of only one subtype, C1) comprising 17(28.3%) isolates and seven (11.7%) isolates, respectively. The six remaining isolates belonged to six other minor types (types D, E, F, G, H, and I).
Among the 60 isolates isolated in 1995, 32 isolates (53.3%) belonged to three subtypes (C1, C2, and C3) of type C, 15 isolates (25%) belonged to type B, and the other 13 isolates (21.7%) belonged to four other minor types (types A, D, F, and J).
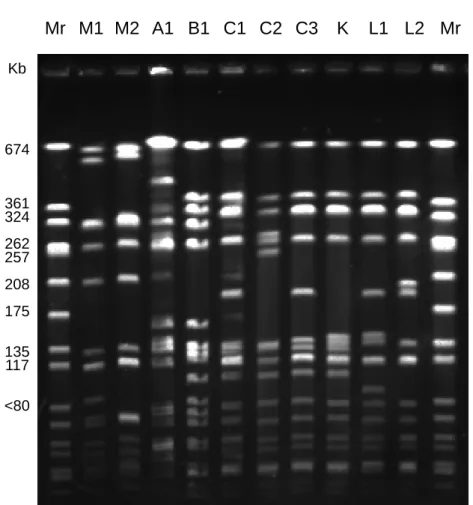
The PFGE banding pattern of the 60 MRSA isolates obtained in 2002 revealed that the type C remained the predominant PFGE type, comprising 21 isolates (35%) (belonging to two subtypes, C1, and C2). Thirteen isolates (21.6%) belonged to type K, six isolates (13.3%) belonged to two subtypes (L1 and L2) of type L, nine isolates (15%) belonged to two subtypes (M1 and M2) of type M, and the other nine isolates belonged to six other minor types (types N, O, P, Q, R. and S). This result was similar to our previous study (22). The PFGE banding patterns of type A, B, C1, C2, C3, K, L1, L2, M1, and M2 are shown in Figure 1.
Distribution of the qacA/B gene
The qacA/B gene was detected in 0, 7 (11.7%), 32 (53.3%), and 44 (73.3%) isolates of the MRSA isolates obtained in 1990, 1994, 1995, and 2002, respectively. Nucleotides sequence determinations revealed that all of the isolates obtained in 1994 and 1995, and 35 of 44 isolates from 2002 harbored qacA, while nine of the isolates from 2002 harbored qacB.
Relationship between PFGE banding patterns and chlorhexidine susceptibility
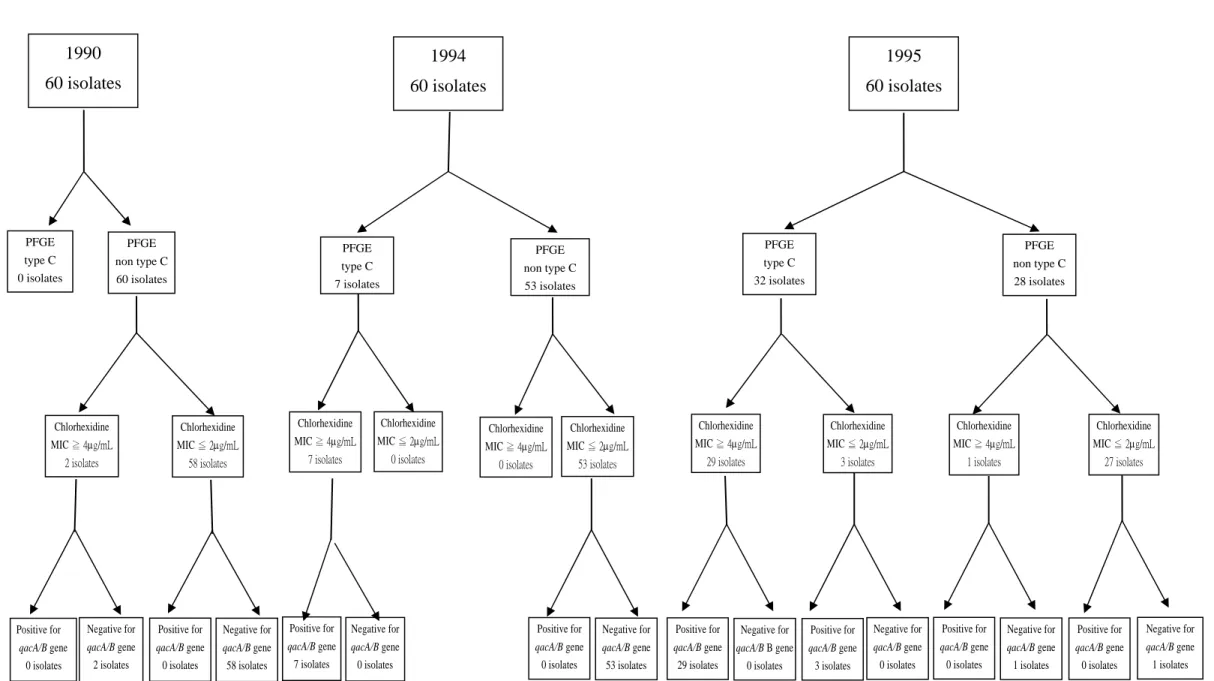
In 1990, there were only two MRSA isolates with the high chlorhexidine MIC of 4 μg/mL (both not belonging to type C, L, K, or M). All seven of the MRSA isolates with high chlorhexidine MIC obtained in 1994 belonged to PFGE type C. In 1995, 29 (96.7%) out of the 30 of MRSA isolates with high chlorhexidine MIC belonged to type C. In 2002, 40 isolates had high MICs of chlorhexidine, 14 of which (35%) belonged to type C, 11 (27.5%) belonged to type K, eight (20.0%) belonged to type M,
six (15%) belonged to type L, and one belonged to type N. The relationship between the PFGE banding patterns and chlorhexidine susceptibility is shown in Figure 2.
Relationship between PFGE banding patterns and qacA/B distribution
A total of 83 MRSA isolates collected in 1994, 1995, and 2002 were found to carry the qacA/B gene. Fifty-six isolates (75.7%) carrying the qacA gene belonged to PFGE type C, 11 isolates (14.9%) carrying the qacA gene belonged to type K and 6 isolates (8.1%) carrying the qacA gene belonged to type L. Only one isolate carrying the qacA gene belonged to type N. All isolates carrying the qacB gene belonged to PFGE type M. Figure 2 shows the relationship between PFGE banding patterns and the distribution of the qacA/B gene.
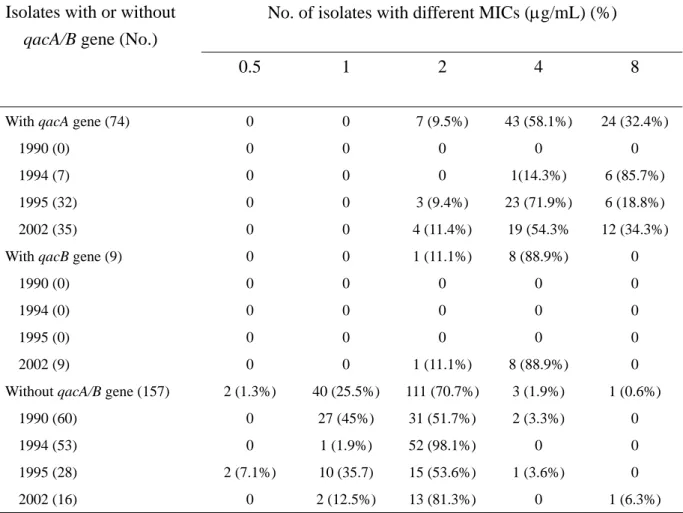
Relationship between the qacA/B gene and chlorhexidine susceptibility
In general, all of the MRSA isolates carrying the qacA/B gene had MICs equal to or higher than 2 μg/mL and 90.4% of them had MICs equal to or greater than the defined ‘high’ MIC. For those MRSA isolates that were devoid of the qacA/B gene, only 2.1% had high MICs. The relationship between qacA/B gene and chlorhexidine susceptibility is shown in Table 2.
Discussion
The present study demonstrates that the prevalence rate of MRSA isolates expressing high chlorhexidine MICs at the NTUH, a hospital using chlorhexidine as a major antiseptic, increased rapidly (from 3.3% to 66.7%) during the 13 years from 1990 through 2002. The prevalence rate of isolates collected in 2002 was similar to that reported from Japan (72%) (14).
Antiseptic resistant S. aureus is a global problem (12-16). An association between the intensity of chlorhexidine use and prevalence of micro-organisms with reduced susceptibility to chlorhexidine has been reported (25). To our best knowledge, however, our present study is the first study documenting the secular trend of
chlorhexidine susceptibility in MRSA.
Chlorhexidine resistance in S. aureus is caused by a multidrug efflux pump in the bacterial cell membrane, which is encoded by two major groups of resistant genes,
qacA/B and smr (14). The qacA/B gene confers a high-level resistance and is thus
worthy of more concern clinically than the smr gene, which confers only a low-level resistance. Previous reports demonstrated that MRSA isolates carrying the qacA/B gene usually expresses chlorhexidine MIC levels equal to or higher than 4 μg/mL (15, 20, 21). Consistent with this, we presently observed that 90.4% of MRSA isolates carrying the qacA/B gene demonstrated chlorhexidine MICs matching or above the 4 μg/mL benchmark. Furthermore, nearly all our MRSA isolates that exhibited high chlorhexidine MICs (75/77, 97.4%) carried the qacA/B gene. Thus, we concluded that the high chlorhexidine MIC in the MRSA isolates in our institute was overwhelmingly due to qacA/B-mediated resistance. Our finding that the resistance was associated with the presence of qacA in 87% of these isolates differs from a report from Japan, which documented the presence of qacB in about 61% of the isolates (15). This genetic dichotomy, if indeed significant, remains to be clarified.
Prior to the present findings, the clinical significance of elevated chlorhexidine MICs in MRSA has been contentious. Although some authors proposed that
chlorhexidine appeared to be as effective as a hand-washing agent for MRSA
regardless of the MIC levels and thus the phenomenon of elevated chlorhexidine MIC levels in MRSA should not be interpreted as clinical resistance (20). However, others reported that hand disinfectants containing only chlorhexidine (e.g., Hibiscrub) were less effective against MRSA isolates with high chlorhexidine MIC levels, and disinfectants containing both alcohol and chlorhexidine (e.g., Hibisol) were more preferred (17). In addition, some studies published recently has demonstrates that alcohol-based hand rubs, even without the presence of chlorhexidine, were more effective than chlorhexidine-based products in terms of spectrum of activities and rapidness of killing (26, 27). Chlorhexidine-based products were noted more easily to
lead to dryness of skin, impairment of skin barrier, skin irritation, and allergic reaction than alcohol-based hand rubs (26). It had also been demonstrated that the acceptance of using alcohol-based hand rubs by HCWs was better than using chlorhexidine-based disinfectants, which in turn increased the adherence of HCWs to hand hygiene (27). Taking all above factors into consideration, it is suggested to consider changing the antiseptic policy at a hospital with a high prevalence of chlorhexidine-resistant pathogens, such as NTUH, from using chlorhexidine-based hand disinfectant to alcohol-based hand rubs.
Type C MRSA isolates appeared at the NTUH in 1994 and became the
predominant strain by 1995 (22). The differences of banding patterns between type C and type K, and L were less than six bands (Figure 1). According to the report of Tenover et al, the banding patterns of PFGE of MRSA isolates originating from a single clone might vary by up to six bands even over a period of only six months (28). In addition, MRSA isolates belonging to PFGE types K and L express the same
pattern of antibiotics resistance as those of type C (data not shown). Therefore, MRSA isolates belonging to PFGE types K and L could be considered as belonging to
subtypes of type C, with their greater genetic variation being explained by a more prolonged evolution time.
Analyzed in this way, 77.5% (31/40) of our MRSA isolates with high
chlorhexidine MICs in 2002 belonged to the same PFGE type (Type C). Therefore, nearly all of the MRSA isolates with high chlorhexidine MIC levels originated from PFGE type C (type K and L were considered as subtypes of type C) (67/77, 87.0%), with the majority harboring qacA gene (Figure 2). The relationship between the prevalence rate of MRSA isolates with high chlorhexidine MIC and the prevalence rate of type C MRSA in four different years are shown in Figure 3.
As stated above, following their detection at the NTUH in 1994, the type C MRSA isolates had become predominant only one year later (22). Before the era of type C MRSA isolates, the prevalence rate of MRSA isolates with high chlorhexidine MIC levels was only 3.3% at the same hospital. All of the type C MRSA isolates in 1994 carried the qacA gene and expressed high chlorhexidine MIC levels (Figure 2). Therefore, the possibility that type C MRSA isolates acquired the qacA gene from other bacteria after their introduction into the NTUH is very low. It is much more likely that type C MRSA harbored the qacA gene and expressed high chlorhexidine MIC levels before they were introduced into the NTUH. In our previous study, type C MRSA isolates were also demonstrated to be highly resistant to many antibiotics and only susceptible to glycopeptide and rifampin (10). Because of their resistance to multiple antibiotics and expression of high chlorhexidine MIC levels, type C MRSA isolates more easily survived than other MRSA strains under the specific selective
pressure in this hospital where chlorhexidine and antibiotics were widely used. Once introduced into this hospital, type C MRSA rapidly became the predominant MRSA strain and the prevalence rate of MRSA isolates with high chlorhexidine MIC levels increased rapidly.
MRSA isolates that belonged to the PFGE type M, which were first noted in 2002, were also interesting. They carried the qacB gene and also expressed
chlorhexidine resistance. They were additionally resistant to erythromycin, clindamycin, and fluoroquinolones but had been shown to be susceptible to
trimethoprim/sulfamethoxazole, rifampin, tetracycline, and glycopeptides. Although they are more susceptible to some antimicrobial agents than type C strains, the possibility of their emergence as a significant problem cannot be discount.
In conclusion, the present study demonstrates that the prevalence rate of MRSA isolates with high chlorhexidine MICs at the NTUH has increased rapidly from 1990 to 2002. The introduction and rapid predominance of an endemic MRSA strain carrying the qacA gene and expressing high chlorhexidine MICs as well as multi-antibiotic resistance may partially explain why a rapid increase of MRSA isolates with high chlorhexidine MICs has been observed in the NTUH. With the consideration of decrease of chlorhexidine susceptibility in MRSA isolates, the shortage of chlorhexidine-based product in spectrum of activities and toxicity to human skin, and the benefits of alcohol-based hand rubs, the antiseptic policies at a hospital with a high percentage of chlorhexidine resistance in some important pathogens, such as NTUH, maybe need changed.
Acknowledgement
The authors thank Mr. Hao-Chun Chang for his assistance in laboratory work in this study.
Biography of first author
Dr. Jann-Tay Wang is an attending staff with specialty in infectious diseases at National Taiwan University Hospital.
REFERENCES
1. Chang SC, Hsu LY, Luh KT, Hsieh WC. Methicillin-resistant Staphylococcus
aureus infection. J Formos Med Assoc 1988;87:157-63.
2. Chang SC, Sun CC, Yang LS, Luh KT, Hsieh WC. Increasing nosocomial
infections of methicillin-resistant Staphylococcus aureus at teaching hospitals in Taiwan. Int J Antimicrob Agents 1997;8:109-14.
3. Hsueh PR, Teng LJ, Chen WH, Pan HJ, Chen ML, Chang SC, Luh KT, Lin FY.
Increasing prevalence of methicillin-resistant Staphylococcus aureus causing nosocomial infections at a university hospital in Taiwan from 1986 to 2001. Antimicrob Agents Chemother 2004;48:1361-4.
4. Haley RW, Hightower AW, Khabbaz RF, Thornsberry C, Martone WJ, Allen JR,
et al. Emergence of methicillin-resistant Staphylococcus aureus infections in United States hospitals: possible role of the house staff-patient transfer circuit. Ann Intern Med 1982;97:297-308.
5. Wang JT, Lin SF, Chiu HL, Wang LC, Tai HM, Jiang CF, Chang SC, Chu SH.
Molecular epidemiology and control of nosocomial methicillin-resistant
Staphylococcus aureus infection at a teaching hospital. J Formos. Med Assoc
2004;103:32-6.
6. Kunin CM. Resistance to antimicrobial drugs--a worldwide calamity.Ann Intern
Med 1993;118:557-61.
7. Weinstein RA. Controlling antimicrobial resistance in hospitals: infection control
and use of antibiotics. Emerg Infect Dis 2001;7:188-92.
8. Boyce JM. Methicillin-resistant Staphylococcus aureus in hospitals and
long-term care facilities: microbiology, epidemiology, and preventive measures. Infect Control Hosp Epidemiol 1992;13:725-37.
9. Cohen SH, Morita MM, Bradford M. A seven-year experience with
methicillin-resistant Staphylococcus aureus. Am J Med 1991; 91(Suppl. 3B):S233-7.
10. Wang, JT, Chen YC, Yang TL, Chang SC. Molecular epidemiology and antimicrobial susceptibility of methicillin-resistant Staphylococcus aureus in Taiwan. Diagn Microbiol Infect Dis 2002;42:199-203.
11. Hospital Infection Control Practices Advisory Committee. 18 February 1997. Recommendations for isolation precautions in hospitals. 1997. [cited February 18, 1997]. Available from: http://www.cdc.gov/ncidod/hip/isolat/isopart2.htm. 12. Emsile KR, Townsend DE, Grubb WB. Isolation and characterization of a family
of small plasmids encoding resistance to nucleic acid-binding compounds in
Staphylococcus aureus. J Med Microbiol 1986;22:9-15.
and quaternary ammonium compounds in methicillin-resistant Staphylococcus
aureus. FEMS Microbiol Lett 1986;34:47-51.
14. Noguchi N, Hase M, Kitta M, Sasatsu M, Deguchi K, and Kona M. Antiseptic susceptibility and distribution of antiseptic-resistant genes in methicillin-resistant Staphylococcus aureus. FEMS Microbiol Lett 1999;172:247-53.
15. Alam MM, Kobayashi N, Uehara N, Watanabe N. Analysis on distribution and genomic diversity of high-level antiseptic resistance gene qacA and qacB in human clinical isolates of Staphylococcus aureus. Microb. Drug Resist 2003;9:109-121.
16. Mayer S, Boos M, Beyer A, Fluit AC, Schmitz FJ. Distribution of the antiseptic resistance genes qacA, qacB and qacC in 497 methicillin-resistant
and –susceptible European isolates of Staphylococcus aureus. J Antimicrob Chemother 2001;47:896-7.
17. Kampf G, Jarosch R, Ruden H. Limited effectiveness of chlorhexidine based hand disinfectants against methicillin-resistant Staphylococcus aureus (MRSA). J Hosp Infect 1998;38:297-303.
18. National Committee for Clinical Laboratory Standards. Performance standards for antimicrobial disc diffusion susceptibility tests. Seventh edition. National Committee for Clinical Laboratory Standards, Wayne, Pa. 2000.
19. National Committee for Clinical Laboratory Standards. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically. Approved standards. Fifth edition. M7-A4. National Committee for Clinical Laboratory Standards, Wayne, Pa. 2000.
20. Cookson BD, Bolton MC, Platt JH. Chlorhexidine resistance in
methicillin-resistant Staphylococcus aureus or just an elevated MIC? An in vitro and in vivo assessment. Antimicrob Agents Chemother 1991;35:1997-2002. 21. Mcdonnell G, Russell AD. Antiseptics and disinfectants: activity, action, and
resistance. Clin Microbiol Rev 1999;12:147-179.
22. Chen ML, Chang SC, Pan HJ, Hsueh PR, Yang LS, Ho SW, Luh KT.
Longitudinal analysis of methicillin-resistant Staphylococcus aureus isolates at a teaching hospital in Taiwan. J Formos Med Assoc 1999;98:426-32.
23. Bannerman TL, Hancock GA, Tenover FC, Miller JM. Pulsed-field gel electrophoresis as a replacement for bacteriophage typing of Staphylococcus
aureus. J Clin Microbiol 1995;33:551-5.
24. Jorgensen M, Givney R, Pegler M, Vickery A, Funnell G. Typing
multidrug-resistant Staphylococcus aureus: conflicting epidemiological data produced by genotypic and phenotypic methods clarified by phylogenetic analysis. J Clin Microbiol 1996;34:394-403.
25. Block C. Furman. M. Association between intensity of chlorhexidine use and micro-organisms of reduced susceptibility in a hospital environment. J Hosp Infect 2002;51:201-6.
26. Kampf G, Kramer A. Epidemiologic background of hand hygiene and evaluation of the most important agents for scrubs and rubs. Clin Microbiol Rev
2004;17:863-93.
27. Boyce JM, Pitter D. Guideline for hand hygiene in health-care settings: recommendations of the Healthcare Infection Control Practices Advisory Committee and the HICPAC/SHEA/APIC/IDSA Hand Hygiene Task Force. MMWR Morb Mortal Wkly Rep 2002;51(RR-16):1-44.
28. Tenover FC, Arbeit RD, Goering RV, Mickelsen PA, Murray BE, Persing DH, et al. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol
Table 1. The susceptibility of 240 MRSA isolates to chlorhexidine, stratified by the year of isolation.
No. of isolates with different MIC* levels (μg/mL) (%) Year of isolation (No. of isolates) 0.5 1 2 4 8 MIC90 (μg/mL) 1990 (60) 0 27 (45%) 31 (51.7%) 2 (3.3%) 0 2 1994 (60) 0 1 (1.7%) 52 (86.7%) 1 (1.1%) 6 (10%) 4 1995 (60) 2 (3.3%) 10 (16.7%) 18 (30%) 24 (40%) 6 (10%) 4 2002 (60) 0 2 (3.3%) 18 (30%) 27 (45%) 13 (21.7%) 8
Table 2. Correlation between qacA/B gene and chlorhexidine resistance in 240 MRSA isolates isolated in 1990, 1994, 1995, and 2002 at NTUH.
No. of isolates with different MICs (μg/mL) (%) Isolates with or without
qacA/B gene (No.)
0.5 1 2 4 8
With qacA gene (74) 1990 (0) 1994 (7) 1995 (32) 2002 (35) 0 0 0 0 0 0 0 0 0 0 7 (9.5%) 0 0 3 (9.4%) 4 (11.4%) 43 (58.1%) 0 1(14.3%) 23 (71.9%) 19 (54.3% 24 (32.4%) 0 6 (85.7%) 6 (18.8%) 12 (34.3%) With qacB gene (9)
1990 (0) 1994 (0) 1995 (0) 2002 (9) 0 0 0 0 0 0 0 0 0 0 1 (11.1%) 0 0 0 1 (11.1%) 8 (88.9%) 0 0 0 8 (88.9%) 0 0 0 0 0 Without qacA/B gene (157)
1990 (60) 1994 (53) 1995 (28) 2002 (16) 2 (1.3%) 0 0 2 (7.1%) 0 40 (25.5%) 27 (45%) 1 (1.9%) 10 (35.7) 2 (12.5%) 111 (70.7%) 31 (51.7%) 52 (98.1%) 15 (53.6%) 13 (81.3%) 3 (1.9%) 2 (3.3%) 0 1 (3.6%) 0 1 (0.6%) 0 0 0 1 (6.3%)
Figure 1 Mr M1 M2 A1 B1 C1 C2 C3 K L1 L2 Mr 674 361 324 262 257 208 Kb <80 175 135 117
Figure 2A 1990 60 isolates 1994 60 isolates 1995 60 isolates PFGE type C 0 isolates PFGE non type C 60 isolates PFGE type C 32 isolates PFGE non type C 28 isolates PFGE type C 7 isolates PFGE non type C 53 isolates Chlorhexidine MIC ≦ 2μg/mL 0 isolates Chlorhexidine MIC ≧ 4μg/mL 7 isolates Chlorhexidine MIC ≧ 4μg/mL 29 isolates Chlorhexidine MIC ≧ 4μg/mL 1 isolates Chlorhexidine MIC ≦ 2μg/mL 27 isolates Chlorhexidine MIC ≦ 2μg/mL 3 isolates Chlorhexidine MIC ≧ 4μg/mL 2 isolates Chlorhexidine MIC ≦ 2μg/mL 58 isolates Chlorhexidine MIC ≦ 2μg/mL 53 isolates Chlorhexidine MIC ≧ 4μg/mL 0 isolates Positive for qacA/B gene 0 isolates Negative for qacA/B gene 2 isolates Positive for qacA/B gene 0 isolates Negative for qacA/B gene 58 isolates Positive for qacA/B gene 7 isolates Negative for qacA/B gene 0 isolates Positive for qacA/B gene 0 isolates Negative for qacA/B gene 53 isolates Positive for qacA/B gene 29 isolates Negative for qacA/B B gene 0 isolates Positive for qacA/B gene 0 isolates Negative for qacA/B gene 1 isolates Negative for qacA/B gene 0 isolates Negative for qacA/B gene 1 isolates Positive for qacA/B gene 0 isolates Positive for qacA/B gene 3 isolates
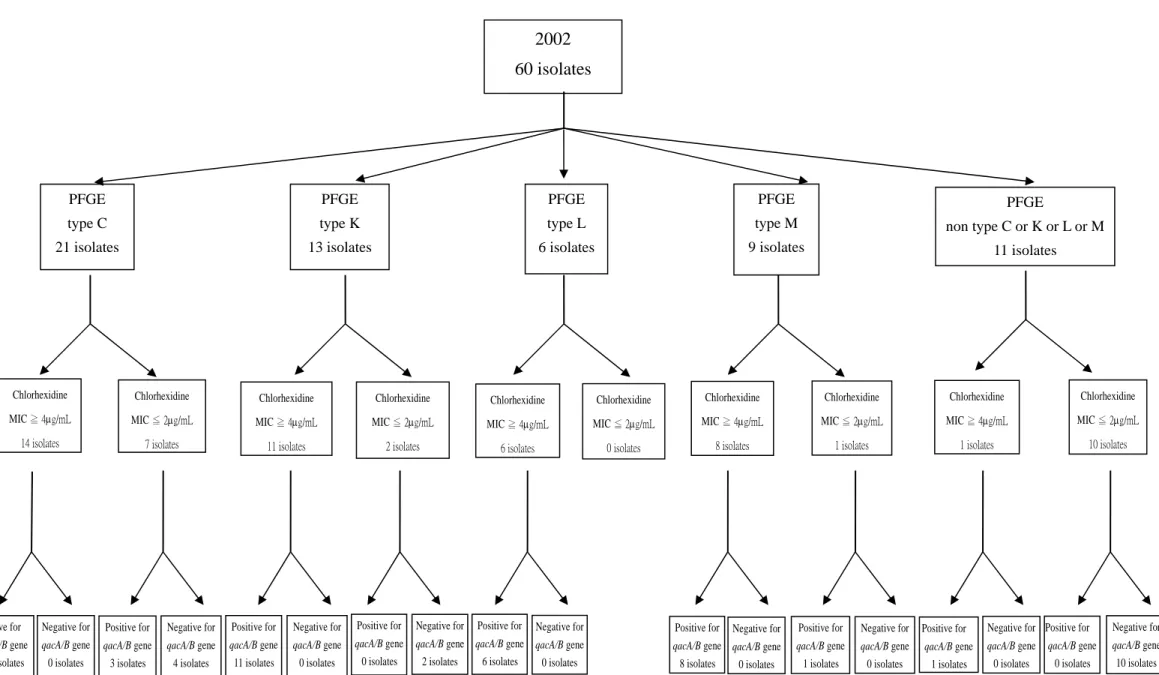
Figure 2B PFGE type C 21 isolates PFGE type K 13 isolates PFGE type L 6 isolates PFGE type M 9 isolates PFGE non type C or K or L or M 11 isolates Chlorhexidine MIC ≧ 4μg/mL 14 isolates Chlorhexidine MIC ≦ 2μg/mL 7 isolates Chlorhexidine MIC ≧ 4μg/mL 11 isolates Chlorhexidine MIC ≦ 2μg/mL 2 isolates Chlorhexidine MIC ≧ 4μg/mL 6 isolates Chlorhexidine MIC ≦ 2μg/mL 0 isolates Chlorhexidine MIC ≧ 4μg/mL 8 isolates Chlorhexidine MIC ≦ 2μg/mL 1 isolates Chlorhexidine MIC ≧ 4μg/mL 1 isolates Chlorhexidine MIC ≦ 2μg/mL 10 isolates Positive for qacA/B gene Negative for qacA/B gene Positive for qacA/B gene Negative for qacA/B gene Positive for qacA/B gene Negative for qacA/B gene Positive for qacA/B gene Negative for qacA/B gene Positive for qacA/B gene Negative for qacA/B gene Positive for qacA/B gene Negative for qacA/B gene Positive for qacA/B gene Negative for qacA/B gene Positive for qacA/B gene Negative for qacA/B gene Positive for qacA/B gene Negative for qacA/B gene 2002 60 isolates
Figure 3 0 10 20 30 40 50 60 70 80
Year 1990 Year 1994 Year 1995 Year 2002
0 10 20 30 40 50 60 70 80 % %
Legends of figures
Figure 1. Pulsed-field gel electrophoresis banding patterns of type M1, M2, A1, B1, C1, C2, C3, K, L1, and L2 MRSA isolates. Lane Mr: molecular weight marker (Staphylococcus aureus NCTC 8325).
Figure 2. (A) The relationship between pulsed-field gel electrophoresis banding patterns, chlorhexidine MIC, and distribution of qacA/B gene in MRSA isolates
obtained in 1990, 1994, and 1995 at the NTUH. (B) Relationship between pulsed-field gel electrophoresis banding patterns, chlorhexidine MIC, and distribution of qacA/B gene in MRSA isolates obtained in 2002 at the NTUH.
Figure 3. Prevalence rate of type C MRSA isolates and MRSA isolates with high chlorhexidine MIC at the NTUH. In 2002, the types K and L were also considered as subtypes of type C. Abbreviation: HCMIC, high chlorhexidine minimum inhibitory concentration.