Differential Expression and Geographic Variation of the
Venom Phospholipases A
2of Calloselasma rhodostoma
and Trimeresurus mucrosquamatus
1Inn-Ho Tsai,2
Yi-Hsuan Chen, Ying-Ming Wang, Ming-Yi Liau,* and Pei-Jung Lu
Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan; Biochemical Sciences Institute, National Taiwan University, Taipei, Taiwan; and *Center for Disease Control, Department of Health,
The Executive Yuan, Republic of China
Received July 11, 2000, and in revised form November 17, 2000; published online February 9, 2001
To investigate the geographic variations in venoms of two medically important pitvipers, we have purified and characterized the phospholipases A2(PLA2s) from
the pooled venoms of Calloselasma rhodostoma from Malaysia, Thailand, Indonesia, and Vietnam, as well as the individual venom of Trimeresurus
mucrosquama-tus collected from both North and South Taiwan.
En-zymatic and pharmacological activities of the purified PLA2s were also investigated. The complete amino
acid sequences of the purified PLA2s were determined
by sequencing the corresponding cDNAs from the venom gland and shown to be consistent with their molecular weight data and the N-terminal sequences. All the geographic venom samples of C. rhodostoma contain a major noncatalytic basic PLA2-homolog and
two or three acidic PLA2s in different proportions.
These acidic PLA2s contain Glu6-substitutions and
show distinct inhibiting specificities toward the plate-lets from human and rabbit. We also found that the T.
mucrosquamatus venoms from North Taiwan but not
those from South Taiwan contain an Arg6-PLA2
desig-nated as TmPL-III. Its amino acid sequence is reported for the first time. This enzyme is structurally almost identical to the low- or nonexpressed Arg6-PLA2from
C. rhodostoma venom gland, and thus appears to be a
regressing venom component in both of the Asian pitvipers. © 2001 Academic Press
Key Words: Trimeresurus mucrosquamatus; Callosel-lasma rhodostoma; snake venom; phospholipase A2;
cloning; complete sequence; platelet aggregation.
Calloselasma rhodostome (Malayan pitviper) is
widely distributed in Southeast Asia and is the most common cause of snakebites in Malaysia and Thailand (1, 2), while Trimeresurus mucrosquamatus is distrib-uted in Taiwan, China, Vietnam, Burma, Northern Bangladesh, and Assam (3). Both species are ground dwelling and primitive pitvipers. The venom of C.
rho-dostoma has been reported (2) to contain moderate to
low levels of phospholipases A2(PLA2s 3
; E.C. 3.1.1.4.). We found that T. mucrosquamatus venom contains relatively abundant PLA2s (4). Snake venom PLA2s usually exist in multiple isoforms of ⱖ50% sequence homology, which exert specific pharmacological effects such as neurotoxicity, myotoxicity, haemolytic activity, edema-induction, antiplatelet, or anticoagulating ac-tivity (5).
Recently, we have cloned and sequenced the cDNAs encoding the PLA2s from the venom gland of a single specimen of C. rhodostoma (Thailand origin) (6) and that of T. mucrosquamatus (South Taiwan) (4, 7). In-terestingly, the respective sequences of the Lys49- and the Arg6-PLA2s in these two venom species were found to be almost identical (6). On the other hand, electro-phoretic analyses on the venoms from about 200 indi-vidual geographic samples of C. rhodostoma revealed that the venom variations can be correlated with the diet or feeding ecology (8, 9). This is so far the only venomous snake whose diets and geographic variations have been investigated in details. We are thus
1
The novel cDNA sequence of TmPL-III has been deposited to Genbank under Accession No. X77646.
2To whom correspondence and reprint requests should be
ad-dressed. Fax:⫹886-2-23635038. E-mail: bc201@gate.sinica.edu.tw.
3Abbreviations used: CRV, Callosellasma rhodostoma venom;
ESI-MS, electrospray ionization mass; FPLC, fast protein liquid chromatography; PCR, polymerase chain reaction; PLA2,
phospho-lipase A2; PRP, platelet rich plasma; RP-HPLC, reversed phase high
performance liquid chromatography; TFA, trifluoroacetic acid; Tm,
Trimeresurus mucrosquamatus.
0003-9861/01 $35.00 257
Copyright © 2001 by Academic Press
All rights of reproduction in any form reserved.
prompted to compare the venom PLA2expression pat-terns of these two species from different regions and try to relate the intraspecies variation of venom pro-teins possibly to the snake feeding ecology.
Herein, we purified and characterized the PLA2s from the pooled venom samples of C. rhodostoma from five geographic regions, as well as 19 individual venom samples of T. mucrosquamatus collected from South and North Taiwan. The identification of venom PLA2 was made according to results of ESI-MS and auto-matic sequencing. Our results suggest that the major causes of geographic variations in both venom species are associated with differential expression of existing mRNAs of venom proteins. Moreover, we show that the functional specificities of certain venom proteins are likely to be related with the feeding ecology of the snake.
MATERIALS AND METHODS
Venoms and other materials. Venom powder of C. rhodostoma
from Vietnam, Thailand, and West-Java origins were purchased from Kentucky reptile Zoo (U.S.A.), Latoxan Co. (France), and Venom Supplies pty Ltd. (W. Australia), respectively. C. rhodostoma venom from Malaysia and origin-unidentified sample from the Hong Kong Zoo were gifts from Professors D. A. Warrel and R. D. G. Theakston (U.K.). T. mucrosquamatus venoms were also collected from nine and ten specimen of individual snakes from North and South Taiwan, respectively. Trifluoroacetic acid (TFA), acetonitrile were purchased from Merck. ADP, apyrase, and sodium citrate were obtained from Sigma. Other reagents and biochemical used are as those described previously (4, 7, 10).
Cloning and sequencing of PLA2s. The cDNA library of T. mu-crosquamatus venom gland was prepared as described (7). In order to
amplify the cDNA encoding venom PLA2, PCR was conducted using
cDNA synthesized from the venom gland mRNA as the template. A pair of mixed-base oligonucleotide primers (21 and 18 residues) was designed based on the highly conserved cDNA regions from other group-II PLA2s (6).
The PCR was performed with SuperTaq DNA polymerase (11). A 0.4-kb DNA fragment was specifically amplified. After being treated with polynucleotide kinase, it was inserted into the pGEM-T vector (Promega) and then transformed into Escherichia coli strain JM109. While transformants were picked-up and specific cDNA clones were selected. Both strands of the cDNA were sequenced by the dideoxynucleotide method (12) using a sequencing kit (Sequenase Version 2.0, U.S. Biochemical). A total of four PLA2-cDNAs were
cloned from venom glands of T. mucrosquamatus (4, 7) and 10 were cloned from that of C. rhodostoma (6).
Purification of PLA2s. About 15 mg of the C. rhodostoma venom
were dissolved in up to 0.3 ml of water. After centrifugation at 12,000-rpm aliquots of 100 l were injected into a gel filtration column (Superdex G75, HR10/30) on a FPLC system (Pharmacia). The column was preequilibrated with 0.1 M ammonium acetate (pH 6.5) at room temperature, and the sample was eluted at 1 ml/min. Fractions corresponding to proteins of 14 and 28 kDa were sepa-rately collected and lyophilized. The proteins were further purified by RP-HPLC on a Vydac C8column or Chemcosorb C18column (4.5⫻
250 mm). Similarly, 10 –15l of fresh venom collected from individ-ual T. mucrosquamatus specimen were diluted with 120l 0.07% TFA and centrifuged. The supernatant was filtrated through a 0.45 mini-filter before injected on the HPLC column. The elution
gra-dient was generated by a solvent delivery system (140B, Applied Biosystem–Perkin–Elmer) using both A-solvent (0.07% TFA) and B-solvent (0.07% TFA in CH3CN).
Protein determination and enzyme assay. Concentration of PLA2
was determined by the absorbance at 280 nm and assuming an extinction coefficient of 1.5 at 1.0 mg/ml. The chromatographic pro-cedures did not affect the PLA2activities. The hydrolysis toward
mixed micelles of dipalmitoyl phosphatidylcholine with either deoxy-cholate or Triton X-100 was measured on a pH-stat apparatus at pH 7.4 and 37°C (7). Assay of neurotoxicity using a neuromuscular preparation of 6 –10 days old chick was performed as previously described (7).
Molecular mass and N-terminal sequences. Purified PLA2s from
HPLC were dried in a vacuum-centrifuge device (Labconco). Their molecular weights were determined by electrospray ionization mass spectrometry (ESI-MS) on a PE-Sciex API100 mass analyzer (Per-kin–Elmer). The N-terminal amino acid sequences of PLA2s were
determined by an automatic gas-phase sequencer (model 477A, Ap-plied Biosystems) coupled with an on-line phenylthio-hydantion amino acid analyzer using the Normal-program.
Preparation of platelets and the aggregation assay. Blood was collected from rabbit and healthy human donor and mixed with 3.8% sodium citrate (9:1, v/v). The blood was centrifuged at 130g for 15 min at room temperature to prepare the platelet-rich plasma (PRP). Platelet aggregation was measured by an aggregometer (Payton, module 600B, Canada). The PRP was kept at 37°C and aliquots (0.45 ml) were preincubated with the PLA2 for 2 min in a siliconized
cylindrical glass cuvette under constant stirring. The aggregation was initiated by the addition of 10M ADP and followed for a period of 5 min. The IC50value of a PLA2, defined as the concentration to
inhibit 50% of the aggregation of PRP caused by 10M ADP, was determined from the dose-dependence curve (13).
RESULTS AND DISCUSSION
Cloning and Predicted Amino Acid Sequences of TmPL-III
We have reported the sequences of three distinct
PLA2s from T. mucrosquamatus venom, namely the
acidic PLA2-I (4), a basic neurotoxic trimucrotoxin (7), and a Lys49-PLA2(10). By cDNA cloning and sequenc-ing that use the venom gland of a southern Taiwan specimen, we have now solved the protein sequence of the fourth one, an acidic PLA2designated as TmPL-III (Fig. 1). This PLA2contains all the conserved Cys and the active site residues including the essential Ca2⫹ binding loop (14, 15). The calculated molecular weight (13,974, assuming seven disulfide bonds) of TmPL-III is also consistent with that determined by ESI-MS of the PLA2purified (peak 7, Fig. 2). The signal-peptide is 16 residues-long and highly similar to those in the cDNA of other of pitviper PLA2s (4, 6, 7).
Geographic Variation of TmPL-III in the Venom
Separated by a river and inhabiting in ranges less than 300 km apart, the southern and the northern T.
mucrosquamatus in Taiwan are not identical in their
skin patterns, e.g., there is an extra dark spot on cheek of the southern snakes. The gel filtration patterns of
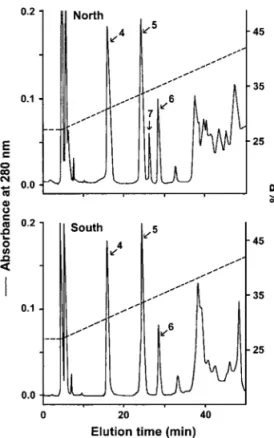
the geographic venom samples of T. mucrosquamateus are very similar but their HPLC elution patterns re-veal geographic differences (Fig. 2). We found that three of the four PLA2 isoforms are present at high abundance in all the Taiwanese habu venoms analyzed whereas TmPL-III is present in the northern but miss-ing in the southern habu venom. A venom sample of T.
mucrosquamatus from Hunan, China, also showed
similar HPLC pattern of PLA2s as those from the northern Taiwan samples (data not shown). By inte-grating the area under their UV-absorbing peaks, the average yields of the T. mucrosquamatu PLA2s were calculated. For the PLA2s with K49, N6, R6, or E6 substitution, the yields are about 10, 7, 2, and 3% of the total proteins in the northern habu venoms, while the corresponding PLA2yields were 10, 10, 0 – 0.3, and 3% in those of the southern venoms analyzed (Fig. 2), respectively.
Structure and Function of TmPL-III
TmPL-III showed very high catalytic activities to-ward the mixed micelles of 3 mM dipalmitoyl phos-phatidylcholine with sodium deoxycholate or Triton X-100, the rates were 690⫾ 26 and 228 ⫾ 17 mol/ min/mg, respectively. Nonlethal at a dose of 3 g/g body weight of the mice (i.p.), it showed antiplatelet activity but very low neurotoxicity toward chick biventer cervicis tissue (7). Addition of 8g/ml TmPL-III to the tissue bath during the electrophysiological assay, the twitch of chicken tissue was 90% inhibited in
163 ⫾ 15 min and 100% inhibited in 263 ⫾ 21 min
(average of four experiments). Thus, the toxicity is 110-folds weaker than trimucrotoxin (7), and the acidic subunit of crotoxin could not increase its toxicity. TmPL-III contains Arg6 and its calculated pI value is 5.3. However, Arg6-PLA2s from the venoms of the
ge-FIG. 1. The cDNA sequence and deduced amino acid sequence of TmPL-III. One-letter codes of amino acids are used and the numbering is shown below the sequences. PCR primers were underlined and the signal peptide was dash-underlined. The nucleotide sequences of CRV-R6 PLA2s differ from that of TmPL-III by 5 bases (shown in bold letter above the sequence) and result in the Lys121Arg substitution.
nus Gloydius (formerly the Old World Agkistrodon) (14) and the tree viper T. stejnergeri are usually much more basic (pI 9.2) (16). How would the differences affect their functions remain to be investigated further.
Common Arg6-PLA2and Lys49-PLA2Genes of Two
Asian Venoms Species
Molecular cloning and cDNA sequencing are very helpful to determine the complete sequences of venom PLA2s and also to find the un-translated PLA2mRNAs. Recent phylogenetic analyses based on mitochondrial DNA sequences suggest that T. mucrosquamatus and
C. rhodostoma are only loosely related (18), but some of
their venom PLA2s (mRNA) share surprising
similari-ties. Two PLA2s, designated as R6a and
CRV-R6b, cloned from the venom gland of C. rhodostoma (6) are structurally identical to TmPL-III except a semi-conserved substitution at residue 121 (Fig. 1). Their signal peptide sequences are also identical. However, both CRV-R6-PLA2s are not translated into venom pro-teins, a situation similar to the un-translated TmPL-III in the southern Taiwan habu.
Polymorphic mRNAs for venom PLA2s (19) and other venom proteins (20, 21) have been suggested to result from gene-duplication and accelerated evolution to ac-quire new functions or new specificities to preys. Venom components may be synergistic to each other and important for catching certain preys thus their mRNA tend to be retained for possible future use. However, the low-expressing Arg6-PLA2in both pitvi-per venoms is more likely examples of the regressing venom component. The presence of low level of certain PLA2 mRNAs in the venom glands possibly bears ad-aptation importance or is beneficial to snake survival.
Geographic Variations of C. rhodostoma Venom
Geographic variations in the electrophoretic pattern of crude venoms, diets, and mitochondrial DNAs of
FIG. 3. Gel-filtration of geographic venom samples of C.
rhodo-stoma. About 10 mg of the pooled crude venoms were dissolved and
fractionated on a Superdex G-75 column (1 cm⫻ 31 cm, Pharmacia, Sweden) in 0.1 M ammonium acetate, pH 6.5. Fractions of 0.5 ml were collected at a flow rate of 1 ml per min. Pooled fraction I and II contained proteins of 26 ⫾ 2 kDa and 14 kDa, respectively. The gel-filtration profile of the Thailand venom sample (6) was almost identical to that of the Malaysia venom.
FIG. 2. Purification of PLA2s of T. mucrosquamatus venoms by
HPLC. The habu enzymes from (A) North Taiwan and (B) South Taiwan were purified by a Chemcosorb RP-HPLC column (ODS-H, 5 m, 1 ⫻ 25 cm). Solvent A was 0.07% TFA in H2O, solvent B was
0.07% TFA in CH3CN, and the elution was by a linear gradient
(27– 42%) over 45 min at a flow rate of 1 ml/min. Peaks 4 –7 were identified as the PLA2isoforms with K49, N6, E6, and R6
substitu-tions, respectively.
more than 200 C. rhodostoma samples have been stud-ied (8, 9). Variations of the venoms have been attrib-uted to difference in snake diet since genetic difference could not be detected. After characterization of the purified C. rhodostoma PLA2s, we found that the anti-platelet acidic PLA2s contribute to the systemic bleed-ing symptom, while the nonhydrolytic basic PLA2 -ho-molog is responsible for edema and myonecrosis fol-lowed the snakebite (6). We now address the questions whether the proportional changes of venom PLA2s are related to the observed geographic variations of the snake and whether functional specificities of the venom PLA2s may possibly correlate with their prey types.
Although all the geographic samples of C.
rhodo-stoma venom we analyzed show very similar
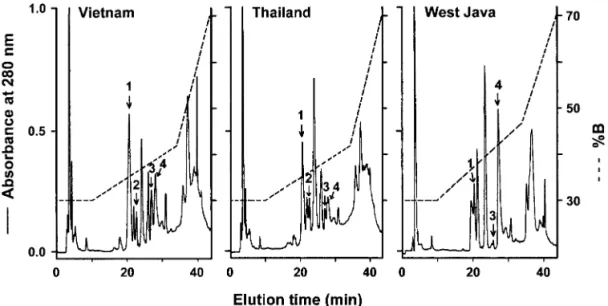
gel-filtra-tion pattern (Fig. 3), their RP-HPLC patterns revealed differences (Fig. 4). The basic PLA2-homolog CRV-W6D49 (6) was purified from the 14-kDa fractions, while two or three acidic PLA2s were purified from the 28-kDa fractions in variable proportions, depending on the venom locality. The N-terminal sequences and the masses of the purified PLA2s derived from ESI-MS analyses are consistent with those deduced from the corresponding cDNA sequences we recently published (6). The elution condition (% B) and the molecular weight determined are: W6 (36%, 13,674), H1E6 (37%, 13,904), S1E6a (38%, 14,071), and S1E6b (39%, 14,052), respectively. Their relative contents in the venoms were also estimated from the area under the UV-absorbance peak (Table I). Notably, the venom samples from Indonesia (West Java) do not contain the CRV-H1E6 in contrast to those from Vietnam and Thailand. CRV-S1E6a is more abundantly expressed
in the Vietnam samples (and the sample from Hong Kong Zoo) than those from the other regions. The con-tent of CRV-S1E6b is relatively higher in the Malaysia and Java samples.
FIG. 4. Purification of the C. rhodostoma venom PLA2s by direct HPLC of the venom. About 1.5 mg of the C. rhodostoma venom from each
region was dissolved in 0.1% TFA (in H2O) and purified after centrifugation by reversed-phase HPLC on a C8 silica gel column. Peaks 1– 4
denote the CRV-PLA2W6, H1E6, S1E6a, and S1E6b, respectively.
TABLE I
Summary of PLA2s in Different Geographic Samples of C.
rhodostoma Venom
Venom origin PLA2variants
Relative abundance (%)a
Unidentified
(Hong Kong Zoo) W6 56
H1E6 ⱕ4 S1E6a 15 S1E6b 25 Vietnam W6 55 H1E6 8 S1E6a 17 S1E6b 20 Thailand W6 63 H1E6 14 S1E6a 6 S1E6b 17 Malaysia W6 59 H1E6 ⱕ5 S1E6a 9 S1E6b 27 West Java W6 41 S1E6a 5 S1E6b 54
aRelative amount of the PLA
2was calculated from area under the
absorbance peak at 280 nm, and the total area of purified PLA2peaks
It was pointed out that the sources of intraspecies variations of pooled venoms include size/age or sex of the snakes, season, and geographic regions covered in the venom collection (22). However, collecting venoms from a broad geographic area and analyzing venoms by precise subcategories are laborious, expensive, and hazardous (23). The C. rhodostoma venoms we have studied are representative samples of the gross areas
or Nations. Apparently, differential expression of venom PLA2messengers, possibly under ecological in-duction, may contribute to the observed geographic variations of venom proteins. Quantitative variations in the proportion of venom PLA2s were also reported for Okinawa T. flavoviridis (24) and Mexican Crotalus
r. ruber (25). In some other cases, qualitative changes
in venom protein sequences were found to be respon-sible for the venom geographic variations (26 –28).
Comparison of the Acidic PLA2s of C. rhodostoma
Venom
The hydrolytic activities of the PLA2s toward two types of micellar substrates are listed in Table II. The activities toward the zweitter-ionic micelles (with Tri-ton X-100) are apparently lower than toward the cat-ionic micelles (with deoxycholate). CRV-H1E6 has 2–3-fold higher enzymatic activity than CRV-S1E6a and b which have similar activities and specificities toward the lecithin substrates.
Amino acid sequences of the acidic CRV-E6-PLA2s are aligned with the most similar sequences of other PLA2s in Fig. 5. Notably, CRV-E6 PLA2s are structur-ally more similar to the E6-PLA2s from American
pitvi-TABLE II
Enzymatic Activities of the Acidic PLA2s
of C. rhodostoma Venom
PLA2
Enzyme activity (mol/min/mg)a
DPPC⫹ deoxycholate DPPC⫹ Triton X-100
H1E6 584⫾ 9 291⫾ 26
S1E6a 245⫾ 27 83⫾ 4
S1E6b 269 82⫾ 6
aThe initial rate was measured by pH-stat method, using mixed
micelles of 3 mM L-dipalmitoyl glycero-3-phosphocholine (DPPC) with either detergents in the presence of 10 mM CaCl2and 0.1 M
NaCl at pH 7.4 and 37°C. Data shown are the average results of two independent assays.
FIG. 5. Multiple sequence alignment of representative pitviper venom PLA2s containing a Glu6. C. rhodostoma venom PLA2s are
abbreviated with CRV. The sequence data for CRV-H1E6, CRV-S1E6a, and CRV-S1E6b have been deposited in the GenBank database with the accession numbers AF104067, AF104068, and AF104069, respectively. Refer to (35, 36) for other sequences. Single-letter codes of amino acids are used. The numbering system follows that of Renetseder et al. (17). Residues identical to those in the top line are denoted with dots, gaps are marked with hyphens.
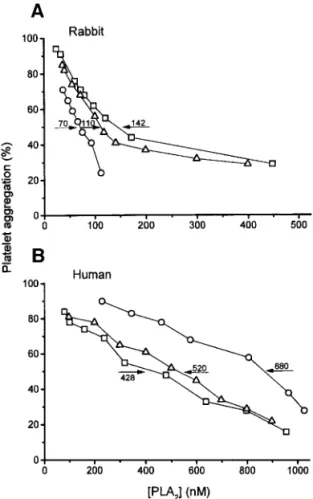
per venoms (sequences 4 and 5, Fig. 5) as compared with other Asian E6-PLA2s. According to previous crys-tallographic study on a venom E6-PLA2(29), the neg-ative-charged surface region involving acidic residues 6 and 115 and aromatic residues 20, 21, 113, and 119 of the PLA2s have been postulated to be essential for the antiplatelet activity. These residues are mostly con-served in the CRV-E6-PLA2s. The enzyme mechanism has been related with a change in the cytoskeleton structure and signal-transduction (30), and the lyso-phospholipid produced may also play a vital part (31). The platelet-rich-plasma (PRP) from human and rabbit were used to assay the inhibition specificities of the CRV-E6-PLA2s in ADP-induced platelet aggrega-tion (Fig. 6). Although CRV-H1E6 has about threefold higher hydrolytic activity than CRV-S1E6a or b (Table II), CRV-H1E6 is a weaker inhibitor (IC50 880 nM) than CRV-S1E6 (IC50 428 or 520 nM) toward the hu-man platelets but a much stronger inhibitor toward the rabbit platelets (IC50 70 nM) (Fig. 6). It is speculated that the lower activity of CRV-H1E6 than CRV-S1E6 for human platelets is due to the substitutions of
W21S, D114N (29), and N67D (26). CRV-S1E6a differs from CRV-S1E6b only by a single substitution (Fig. 5). The single change of Met8 to Leu8 in CRV-S1E6 may increase its antiplatelet potency toward the rabbit platelets but decrease that toward the human platelets by about 25% (Fig. 6). It is known that the platelets from different mammals may have different suscepti-bilities toward antiplatelet agents (13). Thus, expres-sion of the venom antiplatelet PLA2s possibly has been evolved to cope with the feeding ecology of the snakes (8, 9, 24). Interestingly, the venoms of many other pitvipers, e.g., T. flavoviridis (32), T. stejnergeri (33), and Agkistrodon p. picivorus (34), also contain two or more acidic E6-PLA2s. Before a final conclusion on structure–activity relationships of the antiplatelet PLA2s can be reached, the inhibitions of the platelets from more species (including reptiles and amphibians) by various PLA2isoforms or mutants need to be stud-ied in details.
Conclusion
This study demonstrates the convenient use of HPLC and ESI-MS to identify geographic and in-traspecies variations of venom proteins. In five geo-graphic venom samples of C. rhodostoma, the varia-tions were likely attributed to the acidic PLA2s ex-pressed in different proportions. We also showed that these PLA2s have different inhibition specificities to-ward the platelets from human and rabbit and presum-ably may distinguish platelets from various prey spe-cies. A novel venom PLA2(TmPL-III) was purified and completely sequenced and identified as a regressing protein, which is missing in the T. mucrosquamatus venom of southern Taiwan, and is transcribed but not translated in the venom glands of C. rhodostoma. Therefore, the geographic variations in snake venom proteins are usually quantitative rather than qualita-tive, and may result from differential expression of the venom mRNAs.
ACKNOWLEDGMENTS
We thank Dr. D. A. Warrell and Dr. R. G. D. Theakston for generous gifts of the venom of C. rhodostoma from Malaysia, Thai-land, and Hong Kong Zoo, and Professor W. Y. Wang of Kunming Zoology Research Institution for the venom sample of T.
mucrosqua-matus (Hunan Province, China). This study was supported by grants
from Academia Sinica, and National Science Council of Taiwan, ROC (Grant 89-B-FA01-1-4).
REFERENCES
1. Warrell, D. A. (1986) in Natural Toxins: Animal, Plant and Microbial (Harris, J. B., Ed.), pp. 25– 45, Clarendon Press, Ox-ford, UK.
2. Tan, N.-H., and Ponnudurai, G. (1996) J. Toxic. Toxin Rev. 15, 1–17.
FIG. 6. Dose-dependent inhibition of platelet aggregation by CRV-H1E6 (E), CRV-S1E6a (䊐), and CRV-S1E6b (‚). The ADP-induced platelet aggregation was studied with PRP from (A) rabbit and (B) human. Values of IC50(nM) are shown after arrows, and data points
3. Zhao, E.-M., and Adler, K. (1993) Herpetology of China. Society for the Study of Amphibians and Reptiles, Oxford, UK. 4. Wang, Y.-M., Lu, P.-J., and Tsai, I.-H. (1994) J. Chinese
Bio-chem. Soc. 23, 53–58.
5. Tsai, I.-H. (1997) J. Toxicol. Toxin Rev. 16, 79 –114.
6. Tsai, I.-H., Wang, Y.-M., Au, L.-C., Ko, T.-P., Chen, Y.-H., and Chu, Y.-F. (2000) Eur. J. Biochem. 267, 6684 – 6691.
7. Tsai, I.-H., Lu, P.-J., Wang, Y.-M., Ho, C.-L., and Liaw, L.-L. (1995) Biochem. J. (London) 311, 895–900.
8. Daltry, J. C., Wu¨ ster, W., and Thorpe, R. S. (1996) Nature 379, 537–540.
9. Daltry, J. C., Wu¨ ster, W., and Thorpe, R. S. (1998) J. Herpetology
32, 198 –205.
10. Liu, C.-S., Chen, J.-M., Chang, C.-H., Chen, S.-W., Teng, C.-M., and Tsai, I.-H. (1991) Biochim. Biophys. Acta 1077, 362–370. 11. Mullis, K. B., and Faloona, F. (1987) Methods Enzymol. 155,
335–350.
12. Maniatis, T., Fritsch, E. F., and Sambrook, J. (1982) in Molecu-lar Cloning, A Laboratory Manual, Cold Spring Harbor Labora-tory, Cold Spring Harbor.
13. Chen, Y.-L., Huang, T.-F., Chen, S.-W., and Tsai, I.-H. (1995)
Biochem. J. 305, 513–520.
14. Zhao, K.-H., Song, S.-Y., Lin, Z.-J., and Zhou, Y.-C. (1998) Acta
Cryst. D54, 510 –521.
15. Zhao, K.-H., Zhou, Y.-C., and Lin, Z.-J. (2000) Toxicon 38, 901– 916.
16. Lee, S.-Y., Wang, W.-Y., and Xiong, Y.-L. (1997) Toxion 35, 495. 17. Renetseder, R., Brunie, S., Dijkstra, B. W., Drenth, J., and
Sigler, P. B. (1985) J. Biol. Chem. 260, 11627–11634.
18. Kraus, F., Mink, D. G., and Brown, W. M. (1996) Copeia 1996, 763–773.
19. Ogawa, T., Kitajima, M., Nakashima, K. I., Sakaki, Y., and Ohno, M. (1995) J. Mol. Evol. 41, 867– 877.
20. Chang, L. S., Lin, S. K., Huang, S. B., and Hsiao, M. (1999)
Nucleic Acids Res. 27, 3970 –3975.
21. Deshimaru, M., Ogawa, T., Nakashima, K., Nobuhisa, I., Chijwa, T., Shimohigashi, Y., Fukumaki, Y., Niwa, M., Yamashina, I., Hattori, S., and Ohno, M. (1996) FEBS Lett. 397, 83– 88.
22. Warrell, D. A. (1997) in Venomous Snakes. Ecology, Evolution and Snakebite. (Thorpe, R. S., Wu¨ ster, W., and Malhotra, A., Eds.), pp. 189 –203, Clarendon Press, Oxford, UK.
23. Daltry, J. C., Wu¨ ster, W., and Thorpe, R. S. (1997) in Venomous Snakes. Ecology, Evolution and Snakebite. (Thorpe, R. S., Wu¨ ster, W., and Malhotra, A., Eds.), pp. 155–172, Clarendon Press, Oxford, UK.
24. Chijiwa, T., Deshimaru, M., Nobuhisa, I., Nakai, M., Ogawa, T., Oda, N., Nakashima, Ki., Fukumaki, Y., Shimohigashi, Y., Hat-tori, S., and Ohno, M. (2000) Biochem. J. 47, 491– 499. 25. Straight, R. C., Glenn, J. L., Wolt, T. B., and Wolfe, M. C. (1992)
Comp. Biochem. Physiol. 103B, 635– 639.
26. Chuman, Y., Nobuhisa, I., Ogawa, T., Deshimaru, M., Chijiwa, T., Tan, N.-H., Fukumaki, Y., Shimohigashi, Y., Ducancel, F., Boulain, J.-C., Me´nez, A., and Ohno, M. (2000) Toxicon 38, 449 – 462.
27. Tsai, I.-H., Wang, Y.-M., Chiang, T.-Y., Chen, Y.-L., and Huang, R.-J. (2000) Eur. J. Biochem. 267, 1359 –1367.
28. Iha, M., Qi, Z.-Q., Kannki, T., Tomihara, Y., and Yonaha, K. (1995) Toxion 33, 229 –239.
29. Wang, X.-Q., Yang, J., Gui, L.-L., Lin, Z.-J., Chen, Y.-C., and Zhou, Y.-C. (1996) J. Mol. Biol. 255, 669 – 676.
30. Yuan, Y., Jackson, S. P., Newnham, H. H., Mitchell, C. A., and Salem, H. H. (1995) Blood 86, 4166 – 4174.
31. Yuan, Y., Schoenwaelder, S. M., Salem, H. H., and Jackson, S. P. (1996) J. Biol. Chem. 271, 27090 –27098.
32. Ogawa, T., Oda, N., and Nakashima, K. I. (1992) Proc. Natl.
Acad. Sci. USA 89, 8557– 8561.
33. Fukagawa, T., Nose, T., Shimohigashi, Y., Ogawa, T., Oda, N., Nakashima, K., Chang, C.-C., and Ohno, M. (1993) Toxicon 31, 957–967.
34. Welches, W., Reardon, I., and Heinrikson, R. L. (1993) J. Protein
Chem. 12, 187–193.
35. Danse, J. M., Gasparini, S., and Me´nez, A. (1997) in Venom Phospholipase A2Enzyme: Structure, Function and Mechanism
(Kini, R. M., Ed.), pp. 29 –71, Wiley, UK.
36. Wang, Y.-M., Wang, J.-H., Pan, F.-M., and Tsai, I.-H. (1996)
Toxicon 34, 485– 489.