The current progress and future prospects of personalized
radiogenomic cancer study
Juhn-Cherng Liua,b, Wu-Chung Shena,c, Tzu-Ching Shiha,c, Chia-Wen Tsaia, Wen-Shin Changa,d, Der-Yang Choa, Chang-Hai Tsaia and Da-Tian Baua,b
a Terry Fox Cancer Research Laboratory, China Medical University Hospital, Taichung,
Taiwan
b Graduate Institute of Clinical Medical Science, China Medical University, Taichung,
Taiwan
c Department of Biomedical Imaging and Radiological Science, China Medical
University, Taichung, Taiwan
Correspondence to: Da-Tian Bau, Terry Fox Cancer Research Lab, China Medical University Hospital, 2 Yuh-Der Road, Taichung, 404 Taiwan, R.O.C. E-mail address:
artbau 2 @ gmail .com
Keywords: Radiogenomics; Cancer study; Single nucleotide polymorphism; Genotype; Cancer therapy
ABSTRACT
During the last twenty years, mounting studies have supported the hypothesis that there is a genetic component which that plays an important role in clinically observed variability in individual tissue/organ toxicity after radiotherapy. We propose Tthe term “Personalized Radiogenomics” we proposed here for cancer therapy is the translational study of individual genetic variations which that may associate with or contribute to the responses of tissues to radiation therapy used for in the treatment of all types of cancer. The missions of personalized radiogenomic research are 1) to reveal the related genes, proteins, and biological pathways responsible for non-tumor
or tumor tissue toxicity resulting from radiotherapy that could be targeted with radio-sensitizing and/or radio-protective agents, and 2) to identify specific genetic markers that can be used in risk prediction and evaluation models before and after clinical cancer surgery. As fFor the members of the Terry Fox Cancer Research Lab in China Medical University and Hospital, the long-term goal is to develop SNP-based risk models that can be used to stratify patients to more precisely tailored radiotherapy protocols. Worldwide, Tthe field world widely has evolved over the last two decades in parallel with rapid advances in genetic and genomic technology, moving step by step from narrowly focused candidate gene studies to large-scale, collaborative genome-wide association studies. This article will summarize the candidate gene
association studies published so far from the Terry Fox Cancer Research Lab and as well as worldwide, on the risk of radiation-related cancers and highlight some whole-genome association studies showing feasibility in fulfilling the dream of personalized radiogenomic cancer therapy.
The current status of personalized radiotherapy
The golden ultimate goal of radiotherapy for cancer is to regain the control of the growth of the cancer, and at the same time, minimize the possible adverse effects of the radiotherapy. However, there is no denying that cancer patients receiving radiotherapy alone or in combination with chemotherapy display a lot of individual difference in tissue responses. Even though the amazing technological improvements
indeed have enabled a more precise focusing and killing of the tumor cells, there is still some normal tissue that is inevitably exposed to radiation and that becomes
abnormal [1]. In some cancer survivors, this radiation exposure leads to cytotoxicity, tissue toxicity or organ dysfunction and affects their quality of life. This is of particular importance for those patients diagnosed with early-stage localized cancer with a favorable prognosis who may live for a lotnumber of years after cancer therapy in the hospital. With increasing cure rates for common cancers such as nasopharyngeal carcinoma, oral, breast, lung, prostate and cervical cancers, ‘survivorship issues’ are attracting more and more attention and discussion among the
patients and their relatives [2]. Thus, it is critical for radiologists and doctors to re
-consider the morbidity that could result from radiotherapy and the necessary steps that might be undertaken to prevent these complications, as well as to consideror offering
between genomic variations and responses to radiation therapy, radiogenomic cancer study, is a multi-disciplinary and practically translational field of multi-disciplinary and practically translational research. If successful, the work should would not only lead to strategies to that reduce the negative impact of these toxicities for cancer patients, but also lessen the financial burden placed on the health care system to care for people suffering from radiation-induced injuries. Identification of sensitive patients would also permit safe dose escalation in more resistant individuals, leading to an increase in local control and cure for some cancers. Therefore, radiogenomic cancer therapy has the dual potential to both reduce toxicity and to permit dose escalation in personalizing radiotherapy [3].
The spirits of single nucleotide polymorphisms and genome-wide analysis studies
Single nucleotide polymorphisms (SNPs) account for most of the known genetic variation among individuals and are usually defined as polymorphisms in which the minor variant (allele) is present in at least 1% of an investigated population. Non-synonymous SNPs can affect protein function by altering the amino acid composition or by affecting various aspects of the sequential transcriptional and translational outcomes. However, most SNPs are located in regions without any apparent genes or yet-identified functional elements. About 11.8 million SNPs are now included in the
National Center for Biotechnology Information’s public database (www.ncbi.nlm.nih.gov/projects/SNP). Among them, less than half of the SNPs have been validated experimentally, whereas while the rest are yet-unconfirmed variations mainly identified through computational prediction and analysis. Like the oasis in the deserts, the SNPs are not uniformly distributed among the 3 billion base pair (bp) human genome and there are more than 10,000,000 of them identified. From the gene angle, theaverage length of a gene is about 27,000 bp. That is to say, about 100 SNPs are present nearby or within a typical gene. Most SNPs are dbSNP (about 80%) and
are single nucleotide variations, also known as “true” or “typical” SNPs. The remaining SNPs are mainly small insertions/deletions including 1one or a few nucleotides, and this type of SNP is called “INDELs”.
Now we turn to the radiogenomic studies about the annotation for the SNPs. In the Ffollowing we are going to discuss briefly and concisely about the two commonly used methodologies, candidate gene approach and genome-wide analysis study (GWAS). At the beginning, iInitial radiogenomic research focused on candidate gene studies, i.e. studying common SNPs in genes- encoding proteins associated with responses to radiation, such as those in charge of DNA repair and cytoskeleton remodeling. Thise approach by candidate gene studies has the advantage of a
involved in radiation response, thus it limitsing the scope of studies to a manageable number of investigated genes. Prior to recent improvements in the technology for large-scale genotyping and advances in the field of meta-analysis genetic statistics, this approach allowed oncological scientists to conduct affordable studies with modest sample sizes, often with less than one thousand individuals or less. For instance, the
Terry Fox Cancer Research lab has collected hundreds of lung, liver, gastric, colorectal, renal, bladder and prostate cancer patients in central Taiwan. We have also recruited thousands of patients for the most prevalent cancers we are interested in such as oral and breast cancer. Statistically, significant associations were reported for SNPs in a variety of candidate genes and many of these studies have been reviewed previously [4-12]. These associations have to be replicated in validation studies in both the same ethnicity and others to strengthen their contributions to the whole world, or they will have little of less value in clinical practice [13]. Part of the reasons
for the lack of reproducible results for most of the SNPs investigated is due to shortcomings in study design, inclusion and exclusion criteria, and experimental performing progresses such as failure to control for individual ancestry and failure to correct P-values for multiple-factor stratification and ad adjustment. These problems may be further compounded by publication bias. From a statistical point of view, the candidate gene approach considerably reduces the dimensionality of the data set and
therefore yields efficiently increasesd the statistical power in with a limited sample size. From a biological point of view, however, it may be too restrictive to search for SNPs only in the relatively limited genes that are widely accepted as being involved in radiation damage induction and processing.
The candidate gene approach has also been used to study the association between genetic variants and disease susceptibility and outcomes. Up to now, the genomics community has recognized several important limitations for this approach. One problem is that many studies assay just one or a few SNPs within a gene of interest, but many genes contain tens to hundreds of common SNPs [14]. While careful study design can take advantage of linkage disequilibrium between SNPs to capture a large proportion of variation within a gene region, most candidate gene studies have not performed this type of analysis. This is not in itself a limitation to the candidate gene approach, but it does reflects how studies have been performed until recently. A more serious biological limitation is that we are only just beginning to understand the effects that SNPs have on gene expression and protein function. It is often the case that some SNPs found to be associated with a disease or phenotype of interest are not themselves protein coding variants, but often candidates for affecting transcriptional regulation of a gene that may not be the nearest in proximity. For instance, the prostate cancer risk-associated SNP rs339331 lies within a HOXB13 transcription
factor binding site and affects the expression of a more distant gene, RFX6 [15]. MicroRNAs such as mir27 may affect the expression of HOXA10 [16]. Most of all, many of the SNPs selected for candidate gene studies of cancer risk were not replicated in larger GWAS or other populations [17,18]. So, while some of the candidate genes themselves proved to be of importance, thise limited approach to SNP selection prevented identification of the most important loci. In theAt present, in
the radiogenomic candidate gene studies, the SNPs are selected on the basis of changes in the amino acid sequence of the candidate gene, similar to other candidate gene studies. Potentially important SNPs that lie in regulatory regions affecting expression of the candidate genes have likely been missed using this limited approach. Finally, the candidate gene approach relies on the completeness and accuracy of our understanding of the biology behind normal tissue reactions to radiation exposure. Much knowledge of the biological response to radiation comes from cell-based studies, but less is known about what occurs in whole tissues, or interacting tissue systems, upon exposure to radiation.
In response to the lack of success of candidate gene SNP studies and recognition of the above-mentioned limitations inherent in this approach, the focus of radiogenomics has shifted towards GWAS. The Radiogenomics Consortium (RGC) was established [19,20] to enable the collaboration and data sharing needed to obtain
the large sample sizes and multiple studies required for effectivewell-powered
GWAS. The RGC is a National Cancer Institute/NIH-supported Cancer Epidemiology Consortium (http://epi.grants.cancer.gov/Consortia/single/rgc.html) that’s has an
overall aim is to provide a structure in which tissue samples and data can be pooled from investigators performing radiogenomics work on a global scale. The RGC has been essential in facilitating a shift in radiogenomics from narrowly focused single center studies to a multi-institutional approach and from candidate gene studies towards a broader, genome-wide approach.
With the rapid development in the technological and quality controlling systems, arrays in biochips have been made available that allow the simultaneous analysis (genotyping) of more than hundreds of thousands of SNPs in each sample. Using the available information provided by thefrom Human Genome Project and the customer-made service from the genotyping company, it is verymuch convenient now for scientists to design SNP arrays that span the entire genome. The HapMap project describes the statistical relatedness of SNPs by providing a catalog of how SNPs are organized on chromosomes and distributed among different populations (www.hapmap.org). Adjacent SNPs are often linked together into some boxes, and regions of linked variants that are inherited together are known as “haplotypes”. Typically, only about 5 different common haplotypes (with frequencies ~5%) are
found in most parts of human genome [21]. Within a given haplotype, it is therefore possible to identify particular variants that can predict or “tag” the presence of particular variants at other sites. These SNPs that can be used to uniquely identify haplotypes are known as “tag” SNPs. A good GWAS researcher has to pay more attention to the selection of the investigated samples, ensure a sufficient sample size,
andthe well control of the stratification criteria, not only tojust collect more and more
fundingbudgets for their research project. In Taiwan, some clinical researchers have aare naïve mindand applying for huge amounts of funding for a project with huge amounts of budgets with only the methodology of GWAS for a simple disease. The feasibility of such a project may be enhanced with the monopoly of the specific patients for the disease, but itthe project is lacks of spirits insight and prospective
forin the near future.
The efforts on toward revealing cancer susceptibility genotypes for Taiwan cancer
patients from the viewpoints point of view of radiologists
Environmental carcinogens such as radiations may induce DNA single and double strand breaks in the cells and those double strand breaks are a very severe type of DNA damage which should be repaired by the DNA double strand break repair subpathway as soon as possible [22,23]. Mechanically, if cells cannot remove them
immediately by means of homologous recombination (HR) and non-homologous end-joining (NHEJ), those DNA double strand breaks may induce precancerous lesions and cancer itself as well [24,25]. Thus, it is reasonable to propose a hypothesis that genetic polymorphisms in DNA double strand break repair genes influence DNA repair capacity and confer predisposition to several cancers, including skin [26], breast [4,27], liver [28], gastric [6] and oral cancer [7,8]. For this purposeBecause of this, we have devoted ourselves for years intoto the missions of finding potential cancer biomarkers for Taiwanese cancer patients as the cancer predictive and preventive targets. Please keep in mind that the results of a study with a candidate gene approach may not be as splendid as a GWAS, but the significance may be even more solid and useful in clinical practice. During From 2008 to 2011, we awere investigating the contribution of X-ray cross-complementing groups 4, 5 and 6 (XRCC4, 5 and 6) genotypes to Taiwan common cancers in Taiwan. XRCC4 is found to restore DNA double strand break repair and has the ability to support V(D)J recombination of transiently introduced substrates in the XR-1 CHO cell line [29]. The XRCC4 protein interacts directly with Ku70/Ku80, and it is hypothesized that XRCC4 may serve as a flexible tether between Ku70/Ku80 and its associated protein ligase 4 [30]. We have found that those who had G/T or G/G at XRCC4 G-1394T (rs6869366) showed a 3.79-fold (95% confidence interval = 1.47-9.82) increased risk
of gastric cancer compared to those with T/T [6]. As for oral cancer, those who had heterozygous del/ins at XRCC4 intron 3 (rs28360071) showed a 1.57-fold (95% confidence interval=1.12-2.21) increased risk of oral cancer compared to those with ins/ins [7]. The genotypes of the former SNP was were also associated with the personal susceptibility for other types of cancer, such as myoma [31], endometriosis [32], childhood leukemia [33], breast [34], prostate [35], bladder [36], lung [37], and colorectal cancer [38]; while the latter was also associated with childhood leukemia [33]. In addition to these two novel SNPs, the XRCC4 codon 247 (rs3734091) was also associated with increased oral cancer risk [39]. As for XRCC5 (also named Ku80), there were significant differences between oral cancer and control groups in the distributions of their genotypes (P=0.0038) and allelic frequencies (P=0.0044) in the Ku80 promoter G-1401T (rs828907) SNP [9]. After the stratification of personal betel nut chewing status, it was found that the G/T plus T/T genotypes at Ku80 promoter G-1401T significantly enhanced the risk only in the areca chewers (odds ratio=1.603; 95% confidence interval=1.053-2.011), not in the non-areca chewers [9]. This risk genotype is also associated with an increased risk of bladder [40], colorectal [41] and breast cancer [42] in Taiwan. Similarly, the genotype of XRCC6 (also named Ku70), could be associated with pterygium risk at T-991C (rs5751129) [43], oral [8], gastric [44], kidney [45], and liver cancer [28] as well asand childhood leukemia [46].
As for this important SNP at the promoter region, the genotype-phenotype correlation was proved at transcriptional and translational levels. The results showed that the mRNA and protein expression levels in HCC tissues had significantly lower XRCC6 mRNA and protein expressions in the HCC samples with TC/CC genotypes compared with those with the TT genotype (P=0.0037 and 0.0003, respectively) [28]. The most valuable literature produced from the candidate gene rationale in our mind is the findings that the A-2841T polymorphism of SLC2A1, one member of the facilitated glucose transporter family, was significantly associated with 2-[fluorine-18]-fluoro-2-deoxy-D-glucose-uptake in combination with the apurinic/apyimidinic endonuclease Asp148Glu (T to G) polymorphism in the squamous cell type of non-small-cell lung cancer [47]. As far as radiologists are concerned, the uptake of 18F-deoxyglucose
(FDG) of PET, expressed as the SUVmax, is largely dependent on glucose metabolism in lung cancer. SLC2A1 is reported to be the primary glucose transporter of glucose metabolism, and the literature has shown that overexpression of SLC2A1 plays a critical role in the survival and rapid growth of the cancer cells in a suboptimal environment [48]. It is also reported that high FDG uptake is associated with lower overall survival and disease-free survival among non-small cell lung cancer patients [49].
Other outstanding researches, especially those GWAS in radiogenomic cancer research
In the first few years, RGC was successful in enabling radiogenomic researchers to complete and publishing several GWAS onf radiotherapy toxicity. The first radiogenomic GWAS report was a small pilot study with thea sample size of only seventy-nine of patients with erectile dysfunction following radiotherapy for prostate cancer [50]. Despite the relatively small sample size, one SNP was identified with a very significant P-value (P=5.5*10-8) and several others were identified that were
suggestive of significance (P<1.0*10-6). While this study only included a discovery
set, and the SNPs identified must be validated, it highlights the potential of GWAS in surveying genes not previously known to be important for radiotherapy response [50]. The most important SNP identified in this study is locateds within the FSHR gene, which encodes the follicle stimulating hormone receptor involved in gonad development and function [51]. Not as expectedWhile it may have been expected to reveal a significant association of those genes involved in DNA damage recognition and/or relevant DNA repair pathways, this study also identified a SNP on a gene involved in normal cellular function. This finding does not mean that those genes known to be involved in routine radiation responses do not play a role in the etiology of radiation-induced toxicities, but suggests that other tissue-specific pathways may
also be of some importance and play a role in the radiogenomics. This is one of the
evidence that a rough GWAS with less SNPs investigated or too small of a sample size may have lost lots of information and got too many false-positive findings, even with significant P-values. This kind of findings, or other studies revealing those unknown SNPs without any gene nearby, may add the complexity and provide insight
too cancer genomics and radiogenomics.
A second GWAS study in radiogenomics we want to discuss here was another study of prostate cancer patients who were assessed for three toxicity endpoints following radiotherapy: urinary toxicity, erectile dysfunction, and rectal bleeding [52]. This study included a moderate population with about 800 prostate cancer patients, and the dataset was split into a discovery and replication sets. For the urinary toxicity and erectile dysfunction endpoints, the top SNPs fell short of genome-wide significance, but for each endpoint, several suggestive loci were identified. An 8-SNP haplotype block was associated with urinary toxicity, in which the top SNP rs17779457 was associated with a 2.7-fold increase on thein American Urological Association Symptom Score (AUASS) (95% CI 1.2, 4.1) in the discovery set and a 2.4-fold increase (95% CI 1.1, 3.6) in the replication set (P-value =6.5*10-7) [52]. This
haplotype block is locateds within IFNK, a member of the type I interferon family of immunological genes in charge of inhibiting IL-12 signaling and modulating the
cytokine release from innate immune system cells [53]. SNP rs11648233 was associated with erectile dysfunction (P-value=9.1*10-5), and this SNP is locateds
within the 17-beta-hydroxysteroid dehydrogenase II gene (HSD17B2) that catalyzes the oxidative metabolism of androgens [54]. The top locus associated with rectal bleeding contained two SNPs in linkage-disequilibrium, of which rs7120482 with a minor allele frequency (MAF) of 0.37, approached the strict cut-off for genome-wide significance with a P-value of 5.4*10-8, and thus represents a promising risk locus
[55]. This locus lies upstream of SLC36A4, which can modulate the activity of the mammalian target of the rapamycin complex 1 (mTORC1) signaling cascade that affects angiogenesis, proliferation, cell survival, and cellular radiosensitization [56-59]. The study revealed several SNPs associated with intracellular radiosensitivity from the radiogenomic viewpoint of radio-oncologists.
The A third GWAS study famous in radiogenomics was about three interesting SNPs among prostate cancer patients, and with each of them three wasbeing
associated with a syndrome of prostate cancer patients. They were associated with decreased urinary stream, rectal incontinence, and increased urinary frequency, respectively [60]. But the findings in the discovery database could not be representedative in validate populations. Similarly, in 1,194 breast cancer patients, one SNP neared genome-wide significance in the discovery set for association with
telangiectasia, but was not replicated in an independent dataset [61]. From this example we know that it is not easy to find a set of conclusive SNPs for either early predictors of radiosensitivity or prognosis outcomes.
In addition to these kinds of GWAS, recent studies have begun to investigate variation in mitochondrial DNA (mtDNA). There are compelling reasons to hypothesize that such variation could be important in radiation responses, including the role that the mitochondria and mtDNA-encoded genes have on apoptosis signaling and generation of reactive oxygen species. Two studies examined the association between mtDNA variation and radiotherapy responses. The first, in 32 nasopharyngeal carcinoma patients treated with radiotherapy, found that individuals with ≥ grade 2 fibrosis had more nonsynonymous mtDNA variants than individuals with ≤ grade 2 fibrosis [61]. This study also identified a significant association between the nonsynonymous A10398G variant in the NADH dehydrogenase subunit 3 gene and fibrosis. A second larger study of 606 prostate cancer patients evaluated for radiotherapy-related urinary and gastrointestinal toxicity, did not find any significant associations among a panel of mtDNA variants, including the A10398G variant [62]. Up to now we know that most of the findings areresearch is performed among prostate cancer patients since it is the most prevalent cancer among males in
nucleus and is possibly associated with most aging diseases including all types of cancer. OfF course, further large-scale studies of mtDNA are also in urgent need to provide definitive answers for radiogenomics.
Future directions for radiogenomic cancer research and therapy
Clinical radiosensitivity represents the response to a fairly well-defined exposure (i.e.,
radiotherapy), but it is in other respects a complex phenomenon, and future research efforts must appreciate be aware of this fact. The normal tissue response to radiotherapy is made up of a number of different distinctive types of normal tissue reactions. It has been a long standing discussion in radiogenomics whether we should expect genetic factors to have a general impact on radiosensitivity across different types of reactions or if the impact would be specific to certain reactions [63]. The GWAS recently published by Barnett et al. may shed light on this question. Even though this study had substantially more power to detect associations for overall toxicity, the strongest associations were in fact shown for individual endpoints [60]. This observation may seem at odds with the fact that most of the rare radiosensitive syndromes with Mendelian inheritance seem to inflict a general enhancement of clinical radioresponsiveness [64]. Nevertheless, the observation is consistent with clinical data indicating that no strong association exists between the risks of
developing different types of normal tissue toxicity [65,66]. Thus, the initial results of GWAS underline the importance of not only looking for associations with regard to overall toxicity but also addressing separate toxicity endpoints. Future efforts should take this into consideration when developing study designs and data analysis plans.
Going forward, three main directions are promising in fulfilling the dream of personalized radiogenomics. The first is to enlarge the sample size and expand the investigated cancer from prostate cancer to others as well. The second is to establish a worldwide bioinformatic databank for annotation of these novel SNPs for radiogenomics. The third is to establish individual cell lines from the patients to investigate their radiosensitivity and redioresponsiveness for in order to selectively killing of the cancer cells while protecting the non-tumor cells.
To substantially increase the size and number of studies has been proposed by the OncoArray consortium [67] and cancer genomics researchers from nearly 50 countries haves formed a collaborative group that is currently in the process of genotyping over 400,000 DNA samples, of which roughly 200,000 are from people diagnosed with either prostate, breast, lung, colorectal or ovarian cancer, using a custom SNP platform called the OncoArray. This SNP microarray has both a GWAS backbone consisting of 260,000 SNPs plus 440,000 SNPs that have been associated with one or more type of cancer or cancer-related outcome. This includes
approximately 2,000 SNPs nominated by investigators for which initial evidence was obtained linking them with variation in toxicity following radiotherapy. This worldwide collaboration will permit a large expansion in sample size for radiogenomic studies and shortening the time before revealing of required to discover
useful radiogenomic biomarkers.
As for establishing a worldwide bioinformatic databank for annotation of these novel SNPs for radiogenomics, the participations of the nurses, caretakers, patients,
and relatives are as important as thate of surgeons and scientists. The nurses should record all the behaviors and lifestyles with the help of the patients and their relatives, the caretakers should record the all responses and adverse effects of the patients to concurrent chemoradiotherapy or radiotherapy with or without nutrition supplements such as AVEMAR.
The Llast part is the culturing of the tumor and non-tumor cells from the patients during surgery and examining their individual sensitivity to those known or unknown combinations of radiation with radiosensitizers or radioprotectors. The ideal protocol for a radiotherapy is to kill most of the tumor cells while to protecting most of the non-tumor cells. However, the dose of both chemo- or radio-therapy should be fine-tuned according to the cells from each patients, and it is not easyily to reach the such an optimized condition. Only with the well establishmenting of all the three parts can
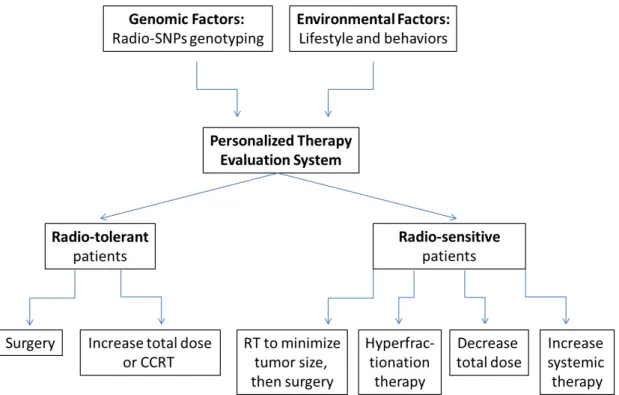
we complete the perfect predictive models for personalized radiogenomic cancer therapy. In the a perfect model, the records in part I for the genomic factors and in part II for the environmental factors will be used to stratify patients according to their predicted risk level, and then each patient will be arranged to an treatment protocol with the experimental data we have got ingathered from part III. For instance, individuals predicted to be at a high risk for developing normal tissue toxicity may experience better outcomes if given a more targeted form of radiotherapy. Alternatively, treatment protocols that avoid radiatherapy, such as the use of surgery or chemotherapy, may be more appropriate for this proportion of cancer patients. For the majority of individuals who are predicted to be at low risk for toxicity, dose escalation protocols could be tested with an aim to improve cancer cell killing rates without increasing toxicity to normal cells. In addition, we anticipate that the genes and pathways uncovered by the agnostic approaches of to genotyping studies will lead to novel hypotheses regarding the biology of radiation response. Investigations of these hypotheses could lead to the development of molecular interventions for preventing the adverse effects of radiation on normal tissues and organs. A personalized therapy evaluation system is shown in Figure 1.
Conclusion
be fulfilled in the near future. Before that, the predictive systems for cancer drug resistance, responses to radiotherapy or radiochemotherapy, and cellular toxicity to radiotherapy or radiochemotherapy should be as widely disseminated and carefully studiedconvenient and popular as possible. Although the ongoing development of radiogenomic cancer study research is promising, the participation of all of us is needed if we are to end the war against cancer.
Acknowledgements
This work was supported by the Terry Fox Cancer Research Foundation in China Medical University and Hospital. The authors deeply appreciate the great help in genotyping from a the following group of outstanding young scientists from the
Department of Biomedical Imaging and Radiological Science in at China Medical University:, Hong-Xue Ji, Chia-En Miao, Lin-Lin Hou, Tzu-Chia Wang, Yun-Ru Syu
REFERENCES
[1] Bentzen SM, Constine LS, Deasy JO, Eisbruch A, Jackson A, Marks LB, et al. Quantitative Analyses of Normal Tissue Effects in the Clinic (QUANTEC): an introduction to the scientific issues. Int J Radiat Oncol Biol Phys 2010; 76:S3-9.
[2] Siegel R, DeSantis C, Virgo K, Stein K, Mariotto A, Smith T, et al. Cancer treatment and survivorship statistics, 2012. CA Cancer J Clin 2012; 62:220-41.
[3] Barnett GC, West CM, Dunning AM, Elliott RM, Coles CE, Pharoah PD, et al. Normal tissue reactions to radiotherapy: towards tailoring treatment dose by genotype. Nat Rev Cancer 2009; 9:134-42.
[4] Bau DT, Fu YP, Chen ST, Cheng TC, Yu JC, Wu PE, et al. Breast cancer risk and the DNA double-strand break end-joining capacity of nonhomologous end-joining genes are affected by BRCA1. Cancer Res 2004; 64:5013-19. [5] Chiu CF, Tsai MH, Tseng HC, Wang CL, Tsai FJ, Lin CC, et al. A novel
single nucleotide polymorphism in ERCC6 gene is associated with oral cancer susceptibility in Taiwanese patients. Oral Oncol 2008; 44:582-86.
[6] Chiu CF, Wang CH, Wang CL, Lin CC, Hsu NY, Weng JR, et al. A novel single nucleotide polymorphism in XRCC4 gene is associated with gastric cancer susceptibility in Taiwan. Ann Surg Oncol 2008; 15:514-18.
[7] Chiu CF, Tsai MH, Tseng HC, Wang CL, Wang CH, Wu CN, et al. A novel single nucleotide polymorphism in XRCC4 gene is associated with oral cancer susceptibility in Taiwanese patients. Oral Oncol 2008; 44:898-902.
[8] Bau DT, Tseng HC, Wang CH, Chiu CF, Hua CH, Wu CN, et al. Oral cancer and genetic polymorphism of DNA double strand break gene Ku70 in Taiwan. Oral Oncol 2008; 44:1047-51.
[9] Hsu CF, Tseng HC, Chiu CF, Liang SY, Tsai CW, Tsai MH, et al. Association between DNA double strand break gene Ku80 polymorphisms and oral cancer susceptibility. Oral Oncol 2009; 45:789-93.
[10] Bau DT, Tsai MH, Tsou YA, Wang CH, Tsai CW, Sun SS, et al. The association of Caveolin-1 genotypes with oral cancer susceptibility in Taiwan. Ann Surg Oncol 2011; 18:1431-38.
[11] Bau DT, Tsai CW and Wu CN. Role of the XRCC5/XRCC6 dimer in carcinogenesis and pharmacogenomics. Pharmacogenomics 2011; 12:515-34. [12] Bau DT, Tsai CW, Lin CC, Tsai RY and Tsai MH. Association of alpha
B-crystallin genotypes with oral cancer susceptibility, survival, and recurrence in Taiwan. PLoS One 2011; 6:e16374.
[13] Barnett GC, Coles CE, Elliott RM, Baynes C, Luccarini C, Conroy D, et al. Independent validation of genes and polymorphisms reported to be associated with radiation toxicity: a prospective analysis study. Lancet Oncol 2012;
13:65-77.
[14] Sachidanandam R, Weissman D, Schmidt SC, Kakol JM, Stein LD, Marth G, et al. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 2001; 409:928-33.
[15] Huang Q, Whitington T, Gao P, Lindberg JF, Yang Y, Sun J, et al. A prostate cancer susceptibility allele at 6q22 increases RFX6 expression by modulating HOXB13 chromatin binding. Nat Genet 2014; 46:126-35.
[16] Yang Q, Jie Z, Ye S, Li Z, Han Z, Wu J, et al. Genetic variations in miR-27a gene decrease mature miR-27a level and reduce gastric cancer susceptibility. Oncogene 2014; 33:193-202.
[17] Easton DF, Pooley KA, Dunning AM, Pharoah PD, Thompson D, Ballinger DG, et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 2007; 447:1087-93.
[18] Long J, Cai Q, Sung H, Shi J, Zhang B, Choi JY, et al. Genome-wide association study in east Asians identifies novel susceptibility loci for breast cancer. PLoS Genet 2012; 8:e1002532.
[19] West C and Rosenstein BS. Establishment of a radiogenomics consortium. Radiother Oncol 2010; 94:117-118.
[20] West C, Rosenstein BS, Alsner J, Azria D, Barnett G, Begg A, et al. Establishment of a Radiogenomics Consortium. Int J Radiat Oncol Biol Phys
2010; 76:1295-96.
[21] Norman A, Kagan AR and Chan SL. The importance of genetics for the optimization of radiation therapy. A hypothesis. Am J Clin Oncol 1988; 11:84-88.
[22] Wood RD, Mitchell M, Sgouros J and Lindahl T. Human DNA repair genes. Science 2001; 291:1284-89.
[23] Yu Z, Chen J, Ford BN, Brackley ME and Glickman BW. Human DNA repair systems: an overview. Environ Mol Mutagen 1999; 33:3-20.
[24] Khanna KK and Jackson SP. DNA double-strand breaks: signaling, repair and the cancer connection. Nat Genet 2001; 27:247-54.
[25] Yang H, Lippman SM, Huang M, Jack Lee J, Wang W, Spitz MR, et al. Genetic polymorphisms in double-strand break DNA repair genes associated with risk of oral premalignant lesions. Eur J Cancer 2008; 44:1603-11.
[26] Han J, Colditz GA, Samson LD and Hunter DJ. Polymorphisms in DNA double-strand break repair genes and skin cancer risk. Cancer Res 2004; 64:3009-13.
[27] Bau DT, Mau YC, Ding SL, Wu PE and Shen CY. DNA double-strand break repair capacity and risk of breast cancer. Carcinogenesis 2007; 28:1726-30. [28] Hsu CM, Yang MD, Chang WS, Jeng LB, Lee MH, Lu MC, et al. The
Anticancer Res 2013; 33:529-35.
[29] Li Z, Otevrel T, Gao Y, Cheng HL, Seed B, Stamato TD, et al. The XRCC4 gene encodes a novel protein involved in DNA double-strand break repair and V(D)J recombination. Cell 1995; 83:1079-89.
[30] Mari PO, Florea BI, Persengiev SP, Verkaik NS, Bruggenwirth HT, Modesti M, et al. Dynamic assembly of end-joining complexes requires interaction between Ku70/80 and XRCC4. Proc Natl Acad Sci U S A 2006; 103:18597-602.
[31] Hsieh YY, Chang CC, Bau DT, Yeh LS, Tsai FJ and Tsai CH. X-ray repair cross-complementing group 4 (XRCC4) promoter -1394( *)T-related genotype, but not XRCC4 codon 247/intron 3 or xeroderma pigmentosum group D codon 312, 751/promoter -114, polymorphisms are correlated with higher susceptibility to myoma. Fertil Steril 2008; 90:1417-23.
[32] Hsieh YY, Bau DT, Chang CC, Tsai CH, Chen CP and Tsai FJ. XRCC4 codon 247*A and XRCC4 promoter -1394*T related genotypes but not XRCC4 intron 3 gene polymorphism are associated with higher susceptibility for endometriosis. Mol Reprod Dev 2008; 75:946-51.
[33] Wu KH, Wang CH, Yang YL, Peng CT, Lin WD, Tsai FJ, et al. Significant association of XRCC4 single nucleotide polymorphisms with childhood leukemia in Taiwan. Anticancer Res 2010; 30:529-33.
[34] Chiu CF, Wang HC, Wang CH, Wang CL, Lin CC, Shen CY, et al. A new single nucleotide polymorphism in XRCC4 gene is associated with breast cancer susceptibility in Taiwanese patients. Anticancer Res 2008; 28:267-70. [35] Chang CH, Chiu CF, Wu HC, Tseng HC, Wang CH, Lin CC, et al. Significant
association of XRCC4 single nucleotide polymorphisms with prostate cancer susceptibility in Taiwanese males. Mol Med Rep 2008; 1:525-30.
[36] Chang CH, Chang CL, Tsai CW, Wu HC, Chiu CF, Wang RF, et al. Significant association of an XRCC4 single nucleotide polymorphism with bladder cancer susceptibility in Taiwan. Anticancer Res 2009; 29:1777-82. [37] Hsu NY, Wang HC, Wang CH, Chang CL, Chiu CF, Lee HZ, et al. Lung
cancer susceptibility and genetic polymorphism of DNA repair gene XRCC4 in Taiwan. Cancer Biomark 2009; 5:159-65.
[38] Bau DT, Yang MD, Tsou YA, Lin SS, Wu CN, Hsieh HH, et al. Colorectal cancer and genetic polymorphism of DNA double-strand break repair gene XRCC4 in Taiwan. Anticancer Res 2010; 30:2727-30.
[39] Tseng HC, Tsai MH, Chiu CF, Wang CH, Chang NW, Huang CY, et al. Association of XRCC4 codon 247 polymorphism with oral cancer susceptibility in Taiwan. Anticancer Res 2008; 28:1687-91.
[40] Chang CH, Chiu CF, Liang SY, Wu HC, Chang CL, Tsai CW, et al. Significant association of Ku80 single nucleotide polymorphisms with bladder
cancer susceptibility in Taiwan. Anticancer Res 2009; 29:1275-79.
[41] Yang MD, Hsu YM, Kuo YS, Chen HS, Chang CL, Wu CN, et al. Significant association of Ku80 single nucleotide polymorphisms with colorectal cancer susceptibility in Central Taiwan. Anticancer Res 2009; 29:2239-42.
[42] Wang HC, Liu CS, Chiu CF, Chiang SY, Wang CH, Wang RF, et al. Significant association of DNA repair gene Ku80 genotypes with breast cancer susceptibility in Taiwan. Anticancer Res 2009; 29:5251-54.
[43] Tsai YY, Bau DT, Chiang CC, Cheng YW, Tseng SH and Tsai FJ. Pterygium and genetic polymorphism of DNA double strand break repair gene Ku70. Mol Vis 2007; 13:1436-40.
[44] Yang MD, Wang HC, Chang WS, Tsai CW and Bau DT. Genetic polymorphisms of DNA double strand break gene Ku70 and gastric cancer in Taiwan. BMC Cancer 2011; 11:174.
[45] Chang WS, Ke HL, Tsai CW, Lien CS, Liao WL, Lin HH, et al. The role of XRCC6 T-991C functional polymorphism in renal cell carcinoma. Anticancer Res 2012; 32:3855-60.
[46] Pei JS, Lee YM, Lo HH, Hsu YN, Lin SS and Bau DT. Association of X-ray repair cross-complementing-6 genotypes with childhood leukemia. Anticancer Res 2013; 33:5395-99.
of 18F-deoxyglucose (FDG) uptake of PET with polymorphisms in the glucose transporter gene (SLC2A1) and hypoxia-related genes (HIF1A, VEGFA, APEX1) in non-small cell lung cancer. SLC2A1 polymorphisms and FDG-PET in NSCLC patients. J Exp Clin Cancer Res 2010; 29:69.
[48] Brown RS, Leung JY, Kison PV, Zasadny KR, Flint A and Wahl RL. Glucose transporters and FDG uptake in untreated primary human non-small cell lung cancer. J Nucl Med 1999; 40:556-65.
[49] Hanin FX, Lonneux M, Cornet J, Noirhomme P, Coulon C, Distexhe J, et al. Prognostic value of FDG uptake in early stage non-small cell lung cancer. Eur J Cardiothorac Surg 2008; 33:819-23.
[50] Kerns SL, Ostrer H, Stock R, Li W, Moore J, Pearlman A, et al. Genome-wide association study to identify single nucleotide polymorphisms (SNPs) associated with the development of erectile dysfunction in African-American men after radiotherapy for prostate cancer. Int J Radiat Oncol Biol Phys 2010; 78:1292-300.
[51] Themmen APN and Huhtaniemi IT. Mutations of gonadotropins and gonadotropin receptors: elucidating the physiology and pathophysiology of pituitary-gonadal function. Endocr Rev 2000; 21:551-83.
[52] Kerns SL, Stone NN, Stock RG, Rath L, Ostrer H and Rosenstein BS. A 2-stage genome-wide association study to identify single nucleotide
polymorphisms associated with development of urinary symptoms after radiotherapy for prostate cancer. J Urol 2013; 190:102-08.
[53] Nardelli B, Zaritskaya L, Semenuk M, Cho YH, LaFleur DW, Shah D, et al. Regulatory effect of IFN-kappa, a novel type I IFN, on cytokine production by cells of the innate immune system. J Immunol 2002; 169:4822-30.
[54] Kerns SL, Stock R, Stone N, Buckstein M, Shao Y, Campbell C, et al. A 2-stage genome-wide association study to identify single nucleotide polymorphisms associated with development of erectile dysfunction following radiation therapy for prostate cancer. Int J Radiat Oncol Biol Phys 2013; 85:e21-28.
[55] Kerns SL, Stock RG, Stone NN, Blacksburg SR, Rath L, Vega A, et al. Genome-wide association study identifies a region on chromosome 11q14.3 associated with late rectal bleeding following radiation therapy for prostate cancer. Radiother Oncol 2013; 107:372-76.
[56] Heublein S, Kazi S, Ogmundsdottir MH, Attwood EV, Kala S, Boyd CA, et al. Proton-assisted amino-acid transporters are conserved regulators of proliferation and amino-acid-dependent mTORC1 activation. Oncogene 2010; 29:4068-79.
[57] Pillai SM and Meredith D. SLC36A4 (hPAT4) is a high affinity amino acid transporter when expressed in Xenopus laevis oocytes. J Biol Chem 2011;
286:2455-60.
[58] Ausborn NL, Le QT, Bradley JD, Choy H, Dicker AP, Saha D, et al. Molecular profiling to optimize treatment in non-small cell lung cancer: a review of potential molecular targets for radiation therapy by the translational research program of the radiation therapy oncology group. Int J Radiat Oncol Biol Phys 2012; 83:e453-64.
[59] Schiewer MJ, Den R, Hoang DT, Augello MA, Lawrence YR, Dicker AP, et al. mTOR is a selective effector of the radiation therapy response in androgen receptor-positive prostate cancer. Endocr Relat Cancer 2012; 19:1-12.
[60] Barnett GC, Thompson D, Fachal L, Kerns S, Talbot C, Elliott RM, et al. A genome wide association study (GWAS) providing evidence of an association between common genetic variants and late radiotherapy toxicity. Radiother Oncol 2014; 111:178-85.
[61] Alsbeih GA, Al-Harbi NM, El-Sebaie MM, Al-Rajhi NM, Al-Hadyan KS and Abu-Amero KK. Involvement of mitochondrial DNA sequence variations and respiratory activity in late complications following radiotherapy. Clin Cancer Res 2009; 15:7352-60.
[62] Fachal L, Mosquera-Miguel A, Gomez-Caamano A, Sanchez-Garcia M, Calvo P, Lobato-Busto R, et al. Evaluating the role of mitochondrial DNA variation to the genetic predisposition to radiation-induced toxicity. Radiother Oncol
2014; 111:199-205.
[63] Andreassen CN, Barnett GC, Langendijk JA, Alsner J, De Ruysscher D, Krause M, et al. Conducting radiogenomic research--do not forget careful consideration of the clinical data. Radiother Oncol 2012; 105:337-40.
[64] Pollard JM and Gatti RA. Clinical radiation sensitivity with DNA repair disorders: an overview. Int J Radiat Oncol Biol Phys 2009; 74:1323-31.
[65] Bentzen SM and Overgaard M. Relationship between early and late normal-tissue injury after postmastectomy radiotherapy. Radiother Oncol 1991; 20:159-65.
[66] Bentzen SM, Overgaard M and Overgaard J. Clinical correlations between late normal tissue endpoints after radiotherapy: implications for predictive assays of radiosensitivity. Eur J Cancer 1993; 29A:1373-76.
Figure 1. Personalized therapy evaluation system that considersing the individual genomic