Electrical detection of DNA using gold and magnetic nanoparticles
and bio bar-code DNA between nanogap electrodes
Tien-Li Chang
a, Chien-Ying Tsai
a, Chih-Chen Sun
a, Ramesh Uppala
a,
Chun-Chi Chen
b, Chun-Hung Lin
b, Ping-Hei Chen
a,*aDepartment of Mechanical Engineering, National Taiwan University, No. 1, Roosevelt Road, Sec.4, Taipei 106, Taiwan, ROC bNational Nano Device Laboratories, Hsinchu, Taiwan 300, ROC
Available online 17 February 2006
Abstract
This paper presents an electrical detection method on a DNA biochip that employs a novel approach for ultra sensitive detection of DNA using self-assembled gold nanoparticles and bio-bar-code-based amplification (BCA) DNA. The experimental study relies on three-components oligonucleotide-modified gold nanoparticles, single-component oligonucleotide-modified magnetic nanoparticles and subsequent detection of amplified target DNA in the form of bio-bar-code ssDNA (single strand DNA) using a chip-based detection method. In this study, the BCA technique measures the bar-code DNA rather than the target DNA. There withal, the DNA chips with nanogap electrodes are fabricated by electron-beam lithography. The gap distance and an electrode height are 300 and 65 nm, respec-tively. Here, the surface between the electrodes and multilayer of gold nanoparticles is established by the hybridization among single strand BCA, the second capture DNA (C2DNA) and the second probe DNA (P2DNA). Measurable current through nanogap electrodes can be obtained over multilayer gold nanoparticles. In this way, magnetic nanoparticles and bio-bar-code DNA are used to amplify obtainable current through nanogap electrodes from the extremely low concentration of target DNA. The detective concentration of target DNA with electrical DNA biosensor is as low as 1 fM for the analysis of current-voltage curves.
Ó 2006 Elsevier B.V. All rights reserved.
Keywords: Bio-bar-code DNA detection; DNA Biochip; Magnetic nanoparticles; Gold nanoparticles; Self-assembly; Electrical Detection
1. Introduction
So far the study of deoxyribonucleic acid (DNA) hybridization chips has evolved into a powerful tool, pro-viding complex informative data from nucleic acid sequences. Taking advantage of the acceleration of genom-ics discoveries, this technology has proven invaluable in
many fields of research and diagnostics [1]. Only few
attempts have so far been made at bio-bar-code-based amplification (BCA) DNA detection technique that is a novel detection for DNA chip to overtake polymerase chain reaction (PCR). Both AuNPs attached to bar-code DNA and magnetic particles have been developed by
Nam and coworkers [2]. Their BCA method utilizes the
two DNA-bound particles, one a gold nanoparticle and the other an iron-oxide containing magnetic microparticle, to detect a DNA sequence associated with anthrax lethal factor. AuNPs having a diameter of 30 nm are attached to two types of oligonucleotide sequences, one is comple-mentary to the target-DNA sequence and the other one is complementary to a bar-code DNA sequence chosen as a unique identification tag for the target. Reintroduction of the magnetic field removes all of the magnetic microparti-cles from solution, leaving bar-code DNA detection. It can be helpful to achieve different targets in one solution and decode the whole system by identifying the bar-codes. In their method, it is essential to use a silver enhancer
solu-tion of AgNO3 and hydroquinone to improve the
sensitivity.
In this experiment, a DNA chip of well-defined feature size and spacing is important for developing highly
sensi-0167-9317/$ - see front matter Ó 2006 Elsevier B.V. All rights reserved. doi:10.1016/j.mee.2006.01.117
*
Corresponding author. Tel.: +886 2 33662689; fax: +886 2 23670781. E-mail address:phchen@ntu.edu.tw(P.-H. Chen).
www.elsevier.com/locate/mee Microelectronic Engineering 83 (2006) 1630–1633
tive and selective immunoassays. Here, there are significant developments in the use of the electron-beam lithography technique for patterning surfaces with DNA on the nano-meter scale. Then, an electrical approach is used to detect DNA hybridization through the formation of a gold nano-structure in the gap surface between two electrodes. The gold nanostructure is established in such a way that it func-tions like single strand DNA as linking chemical. Employ-ing an electrical detection for DNA hybridization has some advantages over the optical approach in rapid clinical diag-nosis of genetic diseases. These advantages include easier operation, less instrumental cost, and denser measuring points. Nevertheless, the sensitivity and consistency of elec-trical detection should be improved significantly for practi-cal application.
In the present study, a DNA chip employs a novel approach for ultra sensitive detection of DNA to take account of self-assembled multilayer AuNPs and BCA with an electrical detection in nanogap electrodes. The major principle technique is the bar-code DNA rather than the target DNA. To begin with, the study employs magnetic nanoparticles (MNPs) to link an oligonucleotide comple-mentary to part of the target-DNA sequence. When both AuNPs and MNPs contact with the test sample, they attach themselves to any strands of the target-DNA sequence. Each target-DNA sequence sandwiches between a gold nanoparticle and a magnetic nanoparticle. Next, a magnetic field can separate the target-DNA sequences and unreacted MNPs from the other components. Furthermore, the use of bioconjugate consists of oligonucleotide probes and colloi-dal AuNPs for DNA hybridization detection via the con-struction of multilayer nanostructure on top of the bottom layer through a layer-by-layer process without a sil-ver enhancer. Afterwards, the oligonucleotides are conju-gated to colloidal AuNPs. They are used to selectively identify surface-confined target via the hybridization among single strand BCA and the second capture DNA (C2DNA) and the second probe DNA (P2DNA) within suitable detection. Here, the first AuNP monolayer is
formed by immobilizing AuNPs on a SiO2 substrate
between the nanogap electrodes using 3-aminopropyltri-methoxysilane (APTMS) molecules. One end of the APTMS compound is to silanize the substrate surface, while the thiol end of the APTMS compound is used to bind the gold nanoparticles. Finally, measurable current-voltage (I–V) behaviors between the AuNP monolayer and AuNP multilayer can be used to identify whether the sample con-tains the bar-code DNA. In addition, the purpose of this study is to develop an electrical detection method utilizing magnetic nanoparticles and bio-bar-code DNA that can amplify obtainable current through nanogap electrodes from the extremely low concentration of target DNA. 2. Experimental
There are five steps involved in the preparation of bio-bar-code DNA sensors, including the synthesis of MNPs
(see Fig. 1.), synthesis of AuNPs (see Fig. 2.), fabrication of nanogap gold electrodes (seeFigs. 3 and 4), conjugation of bar-code DNA with functional magnetic nanoparticles, and immobilization of thiol-modified DNA probes and
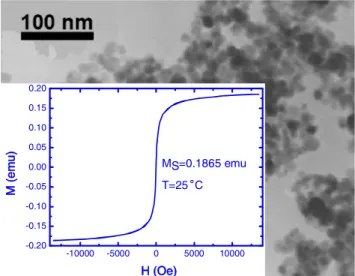
-10000 -5000 0 5000 10000 -0.20 -0.15 -0.10 -0.05 0.00 0.05 0.10 0.15 0.20 MS=0.1865 emu T=25
˚
C M (emu) H (Oe)Fig. 1. TEM image of MNPs with an average diameter of 27 nm. The inset shows the hysteresis curve measured by VSM.
350 400 450 500 550 600 650 700 750 800 0.0 0.2 0.4 0.6 0.8 1.0 1.2 523 nm Absorbance Wavelength (nm)
Fig. 2. TEM image of AuNPs with an average diameter of 12 nm. The inset shows the UV–Vis absorption spectrum of AuNPs solution, showing a strong surface plasma resonance at a wavelength of 523 nm.
Fig. 3. Fabrication sequence of nanogap gold electrodes onto DNA biochip using a lift-off process.
self-assembly multilayer of AuNPs onto the gold surfaces of substrate (see Fig. 5). For identifying the specificity of bar-code DNA, it is required to check whether the hybrid-ization among DNAs has any mismatch. To check it, the chip is washed with a 0.01 PBS buffer at room temperature [3]. If the hybridization among DNAs is not fully comple-mentary, then the top layer of AuNPs is washed away. As a result, the electrical current is reduced significantly back to a value that is obtained for self-assembly gold nanoparticle monolayer. The measurable I–V curves of the array bio-chip are obtained from a HP 4156A precision semiconduc-tor parameter analyzer in the voltage ranges from 1 to +1 V with a scan rate of 1 mV/s.
3. Results and discussion
The 1 and 100 fM T1DNA measurement with and
with-out BCA shows different results in Figs. 6 and 7. In
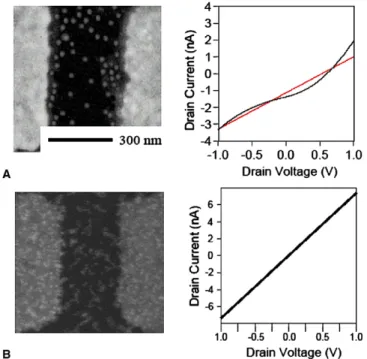
Fig. 6(A), the average density of the particles is shown
which is larger than that shown inFig. 6(B). I–V curve is as straight as the line shown inFigs. 6(B) and 7(B) that
cer-Fig. 4. SEM image of nanogap electrode with a magnification of 4000. The electrodes include gate, drain and source (thickness: 350 A˚ , the distance of gap: 300 nm).
Fig. 6. FE-SEM images of I–V curves of the AuNPs multilayer and nanogap electrode (A) without BCA and (B) with BCA by using T1DNA, the concentration of which is 1 fM.
Fig. 5. The procedure for the attachment of bar-code DNA detection is via nanogap electrodes. (A) Preparation of AuNP-DNA and MNP-DNA conjugates. (B) Hybridization among C1DNA of MNP-DNA conjugate, P1DNA of AuNP-DNA conjugate, and T1DNA. (C) Separation between unbinding AuNP-DNA conjugate and MNP-DNA conjugate with a 6000 Gauss permanent magnet. (D) Collection of bar-code DNA after denature between C2DNA and bar-code DNA of three-component MNP-DNA conjugate. (E) Establishment of second layer of AuNPs with hybridization between bar-code, P2DNA and C2DNA that is already immobilized on the first layer of AuNPs.
tainly correspond with Ohm’s law due to a similarly linear curve in the measurement of range from 1 to 1 V at room temperature. I–V curve appears to be Ohm’s behavior, indicating that it is helpful to integrate the electrical detec-tion of bio-sampling diagnostic chip in the future. How-ever, the I–V curves for the excessively low concentration at 1 and 100 fM are not straight and have a slight
oscilla-tion as shown inFigs. 6(A) and 7(A). From these pieces of evidence, it is very clear that MNPs and bio-bar-code DNA can amplify the current signal in a nanogap with extremely low concentration of T1DNA. Comparing without the BCA study, the FE-SEM images show that the particles distribution is different from with and without using BCA
method as shown in Figs. 6 and 7. The BCA procedure
described above provides a versatile and distinct I–V curve for DNA detection.
4. Conclusion
In this study, the analysis successfully demonstrates the feasibility of electrical detection to DNA biochip with MNPs and bio-bar-code DNA that helps the diagnostic biosensor in the speedy detection of the disease. he interre-lated application for the biological samples of DNA bio-chip with BCA method, to improve the commercial potential of the novel biochip platform, is foreseen. Acknowledgments
The authors thank National Nano Device Laboratories and Nation Science Council for this research (NSC 94-2212-E-002-049 and NSC 94-2212-E-002-017).
References
[1] F.F. Bier, F. Kleinjung, Fresenius J. Anal. Chem. 371 (2001) 151–156. [2] J.M. Nam, S.I. Stoeva, C.A. Mirkin, J. Am. Chem. Soc. 126 (2003)
5932–5933.
[3] S.J. Park, T.A. Taton, C.A. Mirkin, Science 295 (2002) 1503–1506. Fig. 7. FE-SEM images of I–V curves of the AuNPs multilayer and
nanogap electrode (A) without BCA and (B) with BCA by using T1DNA, the concentration of which is 100 fM.