Volume 16,Number7,1997 MaryAnnLiebert,Inc.
Pp.
883-892Production of
Biologically
Active Recombinant
Tilapia
Insulin-Like Growth
Factor-II
Polypeptides
in
Escherichia coli
Cells
and
Characterization of the Genomic Structure
of the
Coding Region
JYH-YIH
CHEN,1
CHI-YAOCHANG,2
JIAN-CHYICHEN,2
SHIH-CHIEHSHEN,1
and JEN-LEIHWU2-3
ABSTRACT
Insulin-like
growth
factor-II
(IGF-II)
is
afetal
growth
factor
inhumans,
but has not beenclearly
identifiedin
fish
uptonow. Foradetailed
understanding
of
thephysiological
response offish
IGF-II,
the firststep
wasto clone
tilapia
IGF-II cDNA from
thebrain
cDNAlibrary, coding
theregion
of
genomic
DNA,
and alsoex-pressing tilapia
IGF-II
polypeptides
from Escherichia coli.
Tilapia
cDNA sequences total1,977
bp,
and pre-dictednucleotide
sequences andamino acid sequences of
tilapia
share77.9%
and90.7%
homology
identity
withrainbow
troutIGF-II,
respectively.
Thegenomic
structureof
thetilapia prepro-IGF-II coding region
is
very
difficult
tosequence in mammals andbirds.
The clonedtilapia
IGF-II
genecoding region
appears muchmore
complex
than
in other vertebrates. Intilapia
IGF-II,
thefirst
coding
exonIencoding
part
of
thesignal
peptide
sequence is 25amino
acids shorter thanthe first
coding
exonof
mammals andbirds.
The other23
amino acids
of
thesignal peptide,
and the firstamino acids of
the Bdomain
andC domain
are encodedby
tilapia coding
exon2.The
C, A,
and
Ddomains,
and
thefirst 20 amino acids of
the Fpeptide
areencodedby
tilapia coding
exon3.
The other Epeptides
and the3'
untranslatedregion
(UTR)
region
areencodedby tilapia
coding
exon4. Thesedata
show that theIGF-II genes
havesignificantly differing
structuresin vertebrateevo-lution,
and therearedifferences ofinterrupting
introns
in theIGF-I
genomic
structurecompared
withmam-mals. To obtain recombinant
biologically
activepolypeptides,
tilapia
IGF-II B-C-A-D domains wereampli-fied
using
thepolymerase
chain reaction
(PCR),
then
ligated
withglutathione
S-transferase
(GST,
pGEX-2T
vector).
Tilapia
recombinant IGF-II
protein
waspurified
and characterized in E.coli.
The fusionprotein
wasalso
digested
with thrombin
andappeared
as arecombinant IGF-II
polypeptide single
bandwith
amolecu-larmassof 7
kD. The recombinant
tilapia
IGF-II
protein biological
function
wasmeasuredby
stimulation of
[3H]thymidine
incorporation.
The assayconcentration
wassetupfrom 0
to120
nMtostimulate
tilapia
ovary cell line(TO-2)
significantly
touptake thymidine.
The
resultssuggest
that therecombinant IGF-II
protein
was dose
dependent.
Brickell,
1996),
Sparus
aurata(Duguay
etal., 1996),
sheep
(Brown
etal, 1990;
Demmer etal,
1993),
rainbow trout(Shamblott
andChen,
1992),
mice(Stempien
etal,
1986),
hu-mans
(Bell
etal, 1984;
Jansen etal,
1985),
andrats(Dull
etal,
1984).
IGF-II isthought
toplay
animportant
role inmam-malian fetal
development (Gray
etal,
1987;
Cohick andClem-mons,
1993);
thehighest expression
of IGF-II mRNA occursin the fetus andneonate
(Soares
etal, 1985),
and genetarget-'Instituteof
Zoology,
National TaiwanUniversity, Taipei,
Taiwan,R.O.C. instituteofZoology,
AcademiaSinica,Taipei,
Taiwan,R.O.C.instituteof FisheriesScience,National Taiwan
University, Taipei,
Taiwan,R.O.C.INTRODUCTION
Insulin-like
growth factor II(IGF-II)
is asingle-chain
polypeptide
that contains theNH2-B-C-A-D-COOH
domain. Thesignal peptide
and E domainareremoved untilmaturepep-tide
production.
IGF-II structure consists of three disulfide bonds(Blundell
etal,
1978),
and cDNA sequencesarehighly
conservedindifferent
animals,
including
chickens(Darling
anding experiments
have shown that IGF-II is akey
component
regulating
fetalgrowth
(DeChiara
etal,
1990),
soIGF-II is alsocalledafetal
growth
factor. Administration of IGF-II in the rat's central nervous system can increase food intake andchange
feeding
behavior(Lauerio
etal,
1987).
As mentioned
above,
IGF-II haspotent
mitogenic
andmeta-bolic effects ininvivo andinvitrosystems
(Humbel
1990;
Luthiet
al, 1992;
Jones andClemmons, 1995).
In mouseIGF-II,
mRNAis
strongly expressed throughout
embryogenesis;
abun-dant IGF-II mRNA wasfound in allmesodermally
andendo-dermally
derived organs, butwasnotdetected in thedevelop-ing
nervoussystem (Lee
etal,
1990).
Culturedsheep
choroidplexus epithelial
cells cansynthesize
and secrete IGF-II and IGFbinding protein-2,
whichsuggests
that the choroidplexus
epithelium
is theimportant
organsecreting
thesepolypeptides
(Holm
etal,
1994).
Inmammals,
IGF-II is animprinting
gene(DeChiara
etal, 1991;
Rappolee
etal, 1992;
Willison,
1991),
with IGF-II transcribed from the
paternal
copy and the IGF-II/mannose6-phosphate
receptor
transcribed from thematernal copy. Thebiological
purpose ofimprinting
issupported by
thesurvival
hypothesis
forplacental
mammals(Haig
andGraham,
1991),
butthe sameisnotknown about fishes.IGF-II has
positive
effectsonfetalgrowth
andregulation
inmammals. These
multiple
functions are reflected in thecom-plex
gene structure; the human IGF-II gene consists of 10 ex-onsof about 30 kb inlength
ofDNA(de
Pagter-Holthuizen
etal, 1987, 1988;
Holthuizenetal,
1990).
The sequenceencod-ing
the mature,circulating
70-amino-acidpolypeptides
arecon-tained withinexons
8, 9,
and 10. Rat andmouseIGF-II genesconsist of 6exonsand span about12kb ofDNA
(Rotwein
andHall, 1990;
Ikejri
etal, 1990,
1991).
Theprepropeptide
is also contained within exons4, 5,
and 6. The ovine IGF-II gene iscomprised
of 9exonsthat spanapproximately
25kb,
and thecoding region
is contained within exons8, 9,
and 10(Ohlsen
et
al,
1994).
Infish,
theIGF-II cDNA sequence from liver isonly
presentinSparus
aurata(Duguay
etal,
1996)
and rain-bow trout(Shammblott
andChen, 1992);
cloning
andse-quencing
ofthepiscine
IGF-II gene hasnotbeenreported,
andnopapers were found
reporting
ontheproduction
of thebio-logically
active fish IGF-II recombinantpolypeptides
fromEs-cherichia coli. In
fish,
IGF-II-likepeptides
arereported
inin-sulin cells of the elasmobranchian endocrine pancreas
(Reinecke
etal,
1994). Furthermore,
it isnotclear if the func-tion of IGF-II in fish brain cell is autocrineorparacrine.
Sore-combinant
tilapia
IGF-IIprotein
should beproduced
as soon aspossible
toinvestigate
thephysiological
functions ofpiscines,
especially
fish brain cellphysiological
functions. It is veryim-portant
toause aprotein
expression
system
thatproduces
high-quantity
andquality tilapia
recombinant IGF-IIproteins
foran-tibody preparation,
immunohistochemicalstudy, biological
activity
analysis,
radioimmunoassay
(RIA),
andenzyme-linked
immunosorbent assay
(ELISA).
In this paper, we first reportcloning
andsequencing
of the IGF-II cDNA gene from thetilapia (hybridized species)
brain cDNAlibrary
and thecloning
andsequencing
ofthetilapia
(Oreochromis
mossambicus)
IGF-II gene of the
coding region.
We found that thecoding
exonarrangementsarevery different from those of mammalian
IGF-II genes, and the
expression
of thetilapia
IGF-II maturepep-tide
by
theE. coli geneexpression
system.Biological activity
analysis
was conductedwith thetilapia
ovary cell line(TO-2
cell
line).
Thestimulatory
effectof recombinanttilapia
IGF-IIpolypeptides
wasconcentrationdependent
withhigh activity
in[3H]thymidine
incorporation,
however,
suggesting
recombinanttilapia
IGF-IIpolypeptides
canstimulatecellularproliferation.
MATERIAL AND
METHODS
Isolation
of tilapia
IGF-II
cDNA clonesAcanthopagrus
schlegeli
IGF-I cDNA(unpublished
data)
from the S domain to E domain was
amplified by
thepoly-merase chain reaction
(PCR).
The PCRproduct,
about a551-bp
DNAfragment,
waspurified by
electroelution and used as aprobe
forisolating
clones from thetilapia
(hybrid)
braincDNA
library
by
theplaque hybridization
method(Maniatis
etal,
1982).
About 1 millionrecombinantbacteriophages
wereseeded on 12LB
plates
and transferred tonylon
membranes. Afterdenaturing, renaturing,
andcross-linking,
the membraneswere
hybridized
to theprobe
ofAcanthopagrus
schlegeli
IGF-I S domain to E domain PCRproducts. Hybridization
buffer used was 40% formamidecontaining
NaDodS04 (7
grams/100
ml),
0.5 M EDTApH
8.0(100
/al/100
ml),
50% PEG8000(20
ml/100ml),
40%formamide(40
ml/100ml)
at37°C for 16 hr. After
hybridization
filterswere washedin 2XSSC,
0.1%NaDodSCv,
0.5XSSC,
0.1%NaDodS04;
0.1XSSC,
0.1%NaDodS04
at roomtemperature, 37°C,and
40°C.Positive
plaques
allowed in vivo excision of thepBluescript
phagemid
from theUni-ZAPvector.Isolation
of tilapia
IGF-II
genomic
clones
Approximately
1 million recombinantbacteriophages
fromthe
tilapia
(O. mossambicus)
genomic library
were screened with32P-labeled
tilapia
IGF-II cDNAfragments.
Thetilapia
ge-nomicDNAlibrary
wasconstructedinphage
charon 40cloning
vector.
Hybridization
buffer used was 50% formamide(hy-bridization buffercontentswerethesamewith the isolationof
tilapia
IGF-IIcDNAcloneconditions),
at42°C for 16hr;
afterhybridization,
filters were washed four times in 0. IXSSC,
0.1%
NaDodS04
at 65°C.Thepositive
plaques
werepurified
andrestriction
mapping
ofDNA fromplaque-purified positive
isolateswasdoneby
Southernblotting
with32P-labeled
tilapia
IGF-II cDNA
probes.
Nucleotide
sequencing
and
analysis
cDNA
clones,
containing pBluescript
double-strandedphagemids
with the cloned DNAinsert,
appeared
ontheLB-ampicillin
plate.
Atotal of four cloneswereobtained and used forlarge-scale plasmid preparation.
The entirecDNA,
digested
with Pst Irestriction enzyme, was subcloned intopUC18
and transformed intoJM109,
and thensequenced by
theSänger
dideoxy
chain-terminationmethod(Sanger
etal,
1977)
andse-quenase kit
(USB,
version2.0).
Thetilapia
IGF-IIphage
DNAswere
digested
with SacI,
then subcloned into thepBluescript
vectorand transformed intoXL1 BlueE. coli host cells. Next the
QIAGEN
plasmid
extraction mini-kit was used to extractDNA,
andonestrandwassequenced by
anABIautosequencer. Thenucleic acid sequences werecompared
with allpublished
Construction
of
recombinanttilapia
IGF-IIexpression
vectorTilapia
IGF-II,
fromB domaintoDdomain,
wasamplified
by
PCR and twooligonucleotide
primers:
5'-CG-GAATTCATATGGAAATGGCCTCGGGCGGAGACGC;
and 5'-CGGAATTCCTCATTCGGACTTGGCAGGTTTG-GCAC. Thesetwo
primers
containedtheEcoRIsite. TheATGinitiation codonwas
designed
in the front of thetilapia
IGF-IIB
domain,
andthestopcodonwasconnected after the finalse-quence ofthe
tilapia
IGF-IIDdomain. The PCRproducts
wereconstructed with the
glutathione-S-transferase
(GST)
gene fu-sion system(Pharmacia
Biotech)
ofexpression
vector(pGEX-2T).
We chose the GST gene fusionsystem
(pGEX-2T)
toexpress
tilapia
IGF-II as afusionprotein
withglutathione-S-transferase because the GSTgene fusion
protein
would beex-pected
tobeefficient,
the fusionproteins
tendtobesoluble,
it isnotnecessarytoisolatethemfrom inclusionbodies,
andthey
are
produced
inhigh yields.
The thrombinsite-specific
cleav-age of GST fusionprotein
thencanberapidly purified
by
glu-tathione-agarose affinity chromatography
(Koland
etal, 1990;
Hooveret
al, 1991;
Kwang
etal,
1991).
Theexpression
vec-torwastransformedinto
BL21(DE3)
E.coli cells andselectedby
ampicillin. Colony hybridization
andsequencing
wereusedto
identify
thecorrectdirection for insertion.Expression
andpurification of
tilapia
IGF-II recombinantprotein
A
single colony
ofBL21(DE3)
E.coli cellscontaining
are-combinant
pGEX-IGF-II
plasmid
was inoculated in 100 ml of2X YTA medium
(16
grams/liter
tryptone;
10grams/liter
yeastextract; 5
grams/liter
NaCl;
100pglm\ ampicillin)
andincubatedat37°C for 10hrwith
shaking.
Theculturewasthentransferredinto 500 ml of2X YTA medium and incubated at 37°C with
shaking
until the absorbanceat600nm was 1.1. Then 0.1 mA/isopropyl-thio-D-galactoside
was added to the culturemediumandthe culturewasincubatedat25°C for3 hrwith
shaking.
The culturewasthencentrifuged
at7,700
Xgfor 10 minat4°Candresuspended
by
50p\
of ice-cold 1Xphosphate-buffered
saline(PBS)
per milliliter of culture. The cellsweredisrupted by
son-ication and 1% Trition X-100
(final
concentration)
was added.The
lysed
cellswerecentrifuged
at12,000
Xgfor10 minat4°Cand the
supernatant
waspassed
through
a0.45-/nm
filter andthenaspirated
intoa column(Pharmacia Biotech).
Thecolumn waswashed
by
1X PBS and thenathrombin solutionwasloadedinto the columnat25°C for 16 hr. The reactionmixture,
whichcon-tainedthe IGF-II
protein,
was collected. The recombinantpro-teinwas run on
NaDodS04-PAGE gel,
and then transferredtoaPVDF membrane for amino acid
sequencing.
Recombinanttilapia
IGF-IIpolypeptides
and GSTprotein
wereseparated
on15%
polyacrylamide
gels,
and theprotein
wasblottedontoaHy-bond ECL nitrocellulose membrane
(Amersham
LifeScience).
The
protein
was detectedby
anti-IGF monoclonal antibodies(kindly
provided by
Dr. Chi-YaoChang
of the Institute ofZo-ology,
AcademiaSínica,
R.O.C).
The ECLWesternblotting
wasperformed
according
toAmersham Life Scienceprotocols.
Bioassay
The
bioactivity
oftilapia
IGF-IIprotein
wasmeasuredwithanin vitroassay.
[3H]Thymidine
incorporation
into DNAinaTO-2cell line
(Chen
etal,
1983)
wasstudied. TO-2 cells(3
X104 cells/well)
were seeded in 24-wellplates
in MEM/F12 mediumsupplemented
with 10%(BSA)
for24hr. After 24 hr of serum-freeincubation,
the cellswereincubated withorwith-outvariousamountsof IGF-II
(0-120
x\M)
for18 hr.The cellswerethen
pulse-labeled
with[3H]thymidine
(2
pCilmX)
for 2hrat25°C. Then 0.5 mlof 0.3NNaOHwas
added,
and after 5 min the mixture was transferredto scintillation vials. After addition of 5 ml ofaquasol,
the solutionwascounted inascin-gaattcgcggccgcctaactcacctgcaatcacaccaaccaaataattcccaacattttg 61 actactgccatctgacatggaaacccagcaaagatacggacatcactcactttgccacoc 1 METQQRYGHHSLCHT 121 ctgccggagaacgcagaacagcagaatgaaggtccagaggatgtcttcgacgagtcgggc 16 CRRTQNSRMKVQRMSSTSRA 181 gctgctctttgcactggccctgacgctctacgcagtggaaatggcctcggcggagacgct 36 LLFALALTLYVVEMASAETL 241 gtgtgggggagaactggtggatgcgctgcagtttgtctgtgaagacagaggc'ttttattt 56 CGGELVDALQFVCEDRGFYF 301 cagtaggccaaccagcaggggtaacaaccgacgcccccagacccgtgggatcgtagagga 76 SRPTSRGNNRRPQTRGIVEE 361 gtgttgtttccgtagctgtgacctcaacctactggagcagtactgtgccaaacctgccaa 96 CCFRSCDLNLLEQYCAKPAK 421 gtccgaaagggacgtgtcagccacctccctacaggtcataccggtgatgcccgcactaaa xl6 SERDVSATSLQVIPVMPALK 481 acaggaagttccgaagaagcaacatgtgaccgtgaagtactccaaatacgaggtgtggca 136 QEVPKKQHVTVKYSKYEVWQ 541 gaggaaggcggcccagcggctccggaggggtgtccccgccattctgagggccagaaagta !56 RKAAQRLRRGVPAILRARKY 601 taagaggcacgcggagaagattaaagccaaggagcaggctatcttccacaggcccctgat 176 KRHAEKIKAKEQAIFHRPLI 661 cagccttcctagcaagctgcctcccgtgttactcaccacggacaactttgtcagtcacaa 196 SLPSKLPPVLLTTDNFVSHK 721 atgagcccgctgccagccctttgcacagacaagagttttgagggtgaaaaaaagactagg 781 ggattatagctttgtctctgacgtcatttcagtggcagtcctctttgacctcccctgccc 841 tgtccgagctcaccaatccctccccctgcacatatccactacgtcttgaacccctggccc 901 ttttctaatgacccnnttaaacccgaactcccccctccccaccaacccaccctcctctgg 961 cacacagacatgccttcacattcttcctgtctgaactctttctctcccaccctctttcag 1021 tcactgatacaaaaggcacaaacacaaaacgtcgaacaaaaagttaacaatttggctgaa 1081 tgcggttcaggtggatccttaagcaaaagacaaaaagagaagggaaaaagaagatgaaag 1141 agatctgtcgtttgcaagtgtcaagaggacacctagcggaatgttttttgtccttgtgga 1201 agacaactgaaagtgaagagctgcttgcatgaaagaatccattccacctcattttcctga 1261 ggcaaaagaaaatctccgttagtcetttagtctgcacctctacctgtaatgggactteca 1321 cactgtaaggaattattttgtaaaattagattcctgttccagcaccttttgatcacaaac 1381 aaaaagcagaaaagagtctgcaaaattgcacattgccacggattacgtctttgtaagaaa 1441 aaaatgggcactattttttcatgaacaatgaacgtgtagcttaaaaaaatgtcacggtgc 1501 tagctttgggaatggactcaaagaagaggtggaaaagcacgtttttttttctttgaatta 1561 ataattaaagctttccgttttaaggaaagtgtgactttttaaaaaaaggaaaattttgga 1621 tatgggggagctctggcagtggcaatgtcaagggggaaagagtcactgaggaaaaatatg 1681 ggctgtgttggcatctaggctcatggtgagtnctagcggctgctatttactagtttgcca 1741 gcataagncagcaagggatgacccgagacctagtccctgttcctcctgtccctctgaggc 1801 tgctggacacatggagcactatggggacacatacgggacaccatggaccacctggattgg 1861 gacagtactatagttcggggacagtacaacctgtttgccatggctttgcggactgttctg 1921 gcaggaagtaacatggcatggactaagaacgagtggggcggccgcgaattc
FIG. 1. Nucleotide sequence of
tilapia (hybrid)
IGF-II cDNA and thepredicted
amino acid sequence of the hormone. The nucleotideswerenumberedbeginning
with the first nucleotideatthe 5' end.The numberonthe second line indicatesthe
or-der of the amino acid
position.
IGF-II contains asignal
pep-tide of 47amino acid residues
(1-47),
aB domainpeptide
of 32 amino acid residues(48-79),
a C domainpeptide
of 11amino acid residues
(80-90),
anAdomainpeptide
of 21 amino acid residues(91-111),
a D domainpeptide
of6 amino acidresidues
(112-117),
andanEdomainpeptide
of 99 amino acid residues(118-216).
The5' UTR sequence containedatotalof 76 nucleotides inlength.
The 3'UTRsequence containedato-tal of
1,247
nucleotides inlength.
Asterisk(*),
Startcodon; #,
Tilapia
Sparus
aurata Rainbow trout Chicken Human Rat MouseSheep
B domain C domainEMAS..AETLCGGELVDALQFVCEDRGFYFSRPT
SRGNNRRPQTREVAS..AETLCGGELVDALQFVCEDRGFYFSRPT
SRGNNRRPQNR
EVAS..AETLCGGELVDALQFVCEDRGFYFSRPT SRSNSRRSQNRAYGTAETLCGGELVDTLQFVCGDRGFYFSPRV
GRNN.RR.INRAYRPSETLCGGELVDTLQFVCGDRGFYFSRPA
SRVS.RR..SRAYRPSETLCGGELVDTLQFVCSDRGFYFSRPS
SRAN.RR..SRAYGPGETLCGGELVDTLQFVCSDRGFYFSRPS
SRAN.RR..SRAYRPSETLCGGELVDTLQFVCGDRGFYFSRPS
SRIN.RR..SR A domain DdomainGIVEECCFRSCDLNLLEQYCA
KPAKSEGIVEECCFRSCDLNLLEQYCA
KPAKSEGIVEECCFRSCDLNLLEQYCA
KPAKSE GIVEECCFRSCDLALLETYCA KSVKSE GIVEECCFRSCDLALLETYCA TPAKSE GIVEECCFRSCDLALLETYCA TPAKSE GIVEECCFRSCDLALLETYCA TPAKSE GIVEECCFRSCDLALLETYCA APAKSE Ancestral vertebrate??A????ETLCGGELVD?LQFVC?DRGFYFSR??
?R???RR???R GIVEECCFRSCDL?LLE?YCA ???KSETilapia
Sparus
aurata Rainbow trout Chicken Human Rat MouseSheep
signal peptide
METQQRYGHHSLCHTCRRTQNSRMKVQRMSSTSRALLFALALTLYVV
METQQRHGRHSLCHTCRRTESSRMKVKKMSSSSRALLFALALTLYVVMETQKRHEYHSVCHTCRRTENTRMKVKMMSSSNRVLVIALALTLYIV
MCAARQILLLLLAFLAYALDSAA
MGIPMGKSMLVLLTFLAFASCCIA MGIPVGKSMLVLLISLAFALCCIA MGIPVGKSMLVLLISLAFALCCIA MGITAGKSMLALLAFLAFASCCYATilapia
Sparus
aurata Rainbow trout Chicken Human Rat MouseSheep
EdomainRDVSATSLQVIPVMPALKQEVPKKQHVTVKYSKYEV1FQRKAAQRLRRGVPAILRARKYKRHAEKIKAKEQA.IFHRPLI
RDVSATSTQVLPVMPPLKQEVSRKQHVTVKYSKYEVWQRKAAQRLRRGVPAILRAKKYRRQAEKIKAQEQA.
IFHRPLIRDVSATSLQ11PMVPTIKQDVPRK.HVTVKYSKYEAIQRKAAQRLRRGVPAILRARKFRRQAVKIKAQEQA.MFHRPLI
RDLSATSLAGLPALN..KESFQKPSH..AKYSKYNVWQKKSSQRLQREVPGILRARRYRWQAEGLQAAEEARAMHRPLI
RDVS.TPPTVLP.DNFRR..YPVGKFFQYDTW.KQSTQRLRRGLPALLRARRGHVLAKELEAFREAKR.HRPLI
RDVS.TSQAVLP.DDFPR..YPVGKFFKFDTW.RQSAGRLRRGLPALLRARRGRMLAKELEAFREAKR.HRPLI
RDVS.TSQAVLP.DDFPR..YPVGKFFQYDTW.RQSAGRLRRGLPALLRARRGRMLAKELKEFREAKR.HRPLI
RDVS.ASTTVLP.DDFTA..YPVGKFFQSDTW.KQSTQRLRRGLPAFLRARRGRTLAKELEALREAKS.HRPLI
Tilapia
Sparus
aurata Raiinbow trout Chicken Human Rat MouseSheep
SLPSKLPPVLLTTDNFVSHK*.. SLGSKLPPVLLATDNYVNHK*.. TLPSKLPPVLPPTDNYVSHN*..SLPSQRPPAPRASPEATGPQE*.
ALPTQDPA.HGGAPPEMASNRK*VLPPKDPA.HGGASSEMSSNHQ*
VLPPKDPA.HGGASSEMSSNHQ*
ALPTQDPATHGGASSEASSD*..
FIG. 2.
Comparison
of theaminoacid sequence oftilapia
IGF-II,
S. aurataIGF-II,
rainbowtroutIGF-II,
humanIGF-II,
ratIGF-II,
mouseIGF-II,
sheep
IGF-II,
and chicken IGF-II.Sequences
start atthe first methioninepeptide
amino acid residue. TheIGF-II
prepropeptide
is divided intothesignal peptide,
and theB,C,
A, D, andEdomain. Adot(•) represents
agap/deletion.
Hypothetical
ancestral vertebrate IGF-II BdomaintoDdomain sequencesareshown below. dilation counter.Every
parameter
in thisexperiment
wasre-peated
three times.RESULTS
Isolation and
characterization
of tilapia
IGF-II cDNA genePrevious
analyses
ofSparus
aurata(Duguay
etal,
1996)
and rainbowtrout
(Shamblott
andChen,
1992)
IGF-IIcDNAgeneswerefromthe livercDNA
library.
Buthere,
wescreened about 1 million recombinantbacteriophages
from thetilapia
(hybrid)
brain cDNAlibrary,
andwefinally
obtained fourpos-itive colonies. The recombinant
plasmids
of each of these cloneswere excisedin
vivo, extracted,
and sizedby
1% agarosegel
electrophoresis.
One of the fourclones,
designated
as12-1,
waschosen for further studies. The size of the cDNA
appeared
tobe about 2 kb and
by sequencing
wasidentifiedastilapia
IGF-II. The nucleotide sequences were
originally
cloned into theEco RI site ofthe
phage
ZAP vector. The recombinant DNAEx 1
Ex
2
Ex
3
Ex 4
Tilapia
Lys
S25 ValS26 Ser B29Arg
B30Ex8
Ex9
Ex
10
Sheep
Ser B29 (A) SerB29(GC
Pro Ell
AspE12
FIG. 3.
Comparison
ofsheep
coding region
structureandtheorganization
of thetilapia
IGF-IIcoding region.
Exonsareshownby
boxes,
and introns andflanking
sequenceare shownby
thin lines. At the bottom of each structureare the relative locationsoftheexonand intron boundaries.
in
Fig.
1. IGF-II cDNA gene contains 76bp
in5' untranslatedregion
(UTR),
1,246
bp
in the 3'UTR,
and thecoding region
hasalength
of 645bp.
The B toDdomains of the IGF-IIma-ture
peptide
translated into a 70-amino-acid residue. The first47 amino acid residues
possibly comprise
thesignal
peptide,
whereas the last 98 amino acid residuescomprise
theEdomain.The IGF-II amino acid
comparison
of different animals isshown in
Fig.
2.Comparison
ofpredicted
amino acidtilapia
IGF-IIBto Ddomains with rainbowtroutIGF-II B toD do-mains shows 95.7%similarity
and 92.9%identity.
Inaddition,
the
length
of thetilapia
IGF-II StoEdomainscompared
tothat of rainbow trout S to E domains shows 90.7%similarity
and81.8%
identity.
However,
tilapia
IGF-IIBtoDdomainscom-pared
tochicken,
human,
rat,mouse,andsheep
IGF-II BtoDdomains,
possess similarities of83.1%,
79.1%, 80.6%, 83.6%,
and80.6%,
and identities of78.5%,
77.6%, 79.1%, 79.1%,
and79.1%,
respectively.
With thepredicted
amino acid sequencecomparison
between fishspecies,
weinferred that the ancestralfish IGF-IIBtoDdomainswere
highly
conserved,
andinamino acid sequencecomparisons
with mammalianIGF-IIBtoD do-mains hada 3-codoninsertion,
and in theB domain had a 2-codon deletion. Thesephenomena
also existed betweentilapia
and rainbowtrout.Figure
2 shows that thetilapia
matureIGF-II
peptide
has 5 amino acids different from the othertwopub-lished fish IGF-IImature
peptides. They
are locatedonB2(for
tilapia
it isMet;
inrainbowtrout and S.aurata,they
areVal),
C3
(for
tilapia
and S.aurata,they
areGly;
inrainbow trout, itis
Ser),
C5(for
tilapia
andS. aurata,they
areAsn;
inrainbow trout, it isSer),
C8(for
tilapia
and S. aurata,they
arePro;
inrainbow trout, it is
Ser),
and C10(for
tilapia,
it isThr;
in rain-bowtroutandS. aurata,they
areAsn).
Isolation and
characterization
of
tilapia
IGF-IIgenecoding
region
About 1 million recombinant
bacteriophages
fromatilapia
TilapiaIGF-IICodingExon1
1 TACACTGCGTAAACGTGGAAAA TGCCCA TGGAAGTCTTCCA TA TTTTGTGACTCTCACC 60 CTCTTATTTCTCCCTTCAAGCACTTTCA TAAAACGrCTCTCCGCCTTTTTTTTTTCATC 119 GGCGAAGAGGAGGAGCAAGGGGTGGGGTCGGTGTAAGGCGCGTGCTTTAGTATATAATA 178 CCTCTCCCTGAGAAGTTTTGCCTGTCGCCTAGTCTTTGGCACAGCTTCTCACTCACCA T 237 CrCTATACTTTAACCCAACTGGGAAACTfiKCTUCCTGCUTUCKCUtiCUkMkKI 296 TCCCAACATTTTGACTACTCCCATCTCACATCGAAACCCAGCAAAGATACCGACATCAC I METQQRYGHH 355 TCACnTGCCACACCTGCCGGAGAACGCAGAACAGCAGAATGAAGgtaaccaaagaaca II SLCHTCRRTQNSRMK 414 agcaaattgttttatactctccggctctgccgtgcgcgtaatgnaagagtat. .- 0.8 Kb -.
TilapiaIGF-IICodingExon 2
1 tgtacctcttcgtctgaaaaaaaaaaaaaaaatctggctgattttgattaaaaaaatgg 60 tatttaactgtcattaactgttattttgttaacgatttctgtatgccacaactttctgc 119 atatcatgggtacatttggtgaaccccatgcttcattccgcagGTCAAGAAGATGTCTT 26 V K K M S S 178 CCACGAATCCCGCGCTGCTCTTTGCACTCGCCCTGACGCTCTACCTAATGGAAATGGCC 32 TNPALLFALALTLYLMEMA 237 TCCGCGGAAACCCTGTnGGGGGAAAACTGGTGGATGCGCTGCAATTTGTCTGTGAAGA 51 SAETLFGGKLVDALQFVCED 296 CAGAAGCTTTTATTTCAgtaagtttcaaagcattacnagtttccccaatggctgcgtga 71 R S F V F S 355 ttgctcatttgcctgttgaatctctctgttgtgcccttgcacacatctgtttggagcaa 414 aagtgggaagttacccactacnaatacttcgttactgtactccagtatagttttcagtt 473 agaatttttgccccctacatttttaaacagatatctgtactttctactcc. .- 2.8Kb -.
TilapiaIGF-IICodingExon 3
1 gatgttgtgtttgcagtccctaacctntacgtcttcattcctttttgtgtttttcctca 60 gGTAGGCCAACCAGCAGGGGTAACAACCGACGCCCCCAGACCCGTGGGATCGTAGAGGA 77 RPTSRGNNRRPQTRG1VEV 119 GTGTCTTTTCTGTAGCTGTGACCTCAACCTACTGGAGCAGTACTGTGCCAAACCTGCCA 96 CLFCSCDLNLLEQYCAKPAK 178 AGTCCGAAAGGGACGTGTCAGCCACCTCTCTACAGGTCATACCGGTGATGCCCGCACTA 116 SERDVSATSLQV1PVMPAL 237 AAACAGgtacgtctaagcaacaacaacaacaggccagtatgggaaatagtgctaatccc 135 K Q 296 agctctatctgtcctcccatctcctgtgcccccattcacctctgaggctagcccctatg 355 tcactgactcUgagtagagtgtacccacgctaacgcagttatatctagUaattggcc 414 aatggaaagcactcaacttacaaagaaagtgctgacagtcatggaaaacattacaaaag 473 tcacaacacgttatattcaggaaaaggaatgtgttaagtgcgtatatgaaggaa. .- 1.3 Kb -.
TilapiaIGF-IICodingExon 4
1 attgaacaatatnttatnaccntaatgaatgatccttcttttccctttttcttctattt 60 tcgcccgcacgccacaatagGAAGTTCAGAAGAAGCAACATGTGACCGTGAAGTATTCC 137 E V Q K K Q H V T V K Y S 119 AAATACGAGGTGTGGCAGAGGAAGGCGGCCCAGCGGCTCCGGAGGGGTGTCCCCGCCAT 150 KYEVWQRKAAQRLRRGVPAI 178 TCTGAGGGCCAGAAAGTATAAGAGGCACGCGGAGAAGATTAAACCCAAGGAGCAGGCTA 170 LRARKYKRHAEKIKAKEQAI 237 TCTTCCACACGCCCCTGATCAGCCTTCCTAGCAAGCTGCCTCCCGTGTTGCTCACCACG 190 FHRPL1SLPSKLPPVLLTT 296 GACAACTTTGTCAGTCACAAATGAGCCCGCTGCCAGCCCTTTGCACAGACAAGAGTTTT 209 D N F V S H K * 355 GAGGGTGAAAAAAAGACTAGGGGATTATAGCTTTGGTCTTCTGACGTCATTTCTGTGGC 414 AGTCCTCTTTGACCTCCCCTCCCCTGTCCGAGCTC
FIG. 4. Partial nucleotide sequence of the O. mossambicus IGF-II gene
coding region.
The uppercase lettersrepresenttran-scribed
regions
and lowercase lettersrepresent
intronsorflank-ing
sequences. The italic lettersrepresent
theregions
thatre-main uncertain
by
cDNAsequencing.
Amino acids number:1^-7,
signal peptide
sequence;48-79,
B domain sequence;80-90,
C domain sequence;91-111,
A domain sequence;112-117,
D domain sequence;118-215,
E domain sequence. Thestopcodon is indicated withanasterisk(*).
positive
colonieswereobtained. The fourpositive
colonieswereextracted and restriction enzyme
mapping
of the DNA from the fourpurified plaques
wasperformed.
Thegenomic
DNAse-quences of the
tilapia coding region
weredivided into four ex-ons and weremapped
as showninFig.
3. Thetilapia coding
exons were foundto span a
region
ofapproximately
12.9 kb.In mammalian IGF-II genes, the
coding region
iscomprised
of threeexonsbutintilapia
thecoding region
iscomprised
of fourexons.The
tilapia
IGF-IIcoding
exon 1 wasfrom the 5' UTR(compared
cDNAsequence)
tothesignal
peptide
(S25);
cod-ing
exon2was frompartof thesignal peptide
(S26)
toB do-main(B28);
coding
exon 3was from the C domain(Cl)
toE domain(E20);
andcoding
exon 4was from the Edomaintopart
of the 3'UTR(compared
cDNAsequence).
Intilapia,
cod-ing
exon 1contains 25 aminoacids,
coding
exon2contains 51amino
acids,
coding
exon3contains60aminoacids,
andcod-ing
exon4 contains 80 amino acids.Betweenexon 1 andexon2,
there isaninterruption by
anintron of about 0.8kb;
between exon2 andexon3,
there isaninterruption by
anintron of about2.8
kb;
and betweenexon3 andexon4there isaninterruption
by
an intron ofabout 1.3 kb. The sequence ofthefourtilapia
coding
exons andpartof theflanking
sequence of intronsareshown in
Fig.
4. The sequences of the exon-intronjunctions
are often describedas
conforming
tothe GT-AG rule(Breath-nach et
al, 1978;
Breathnach andChambón,
1981).
Thecod-ing region
oftilapia
IGF-II genepredicts
aprepropeptide
of 215amino
acids,
including
a 47-amino-acidsignal peptide,
a70-amino-acidmature
peptide,
anda98-amino-acidEpeptide.
Thepredicted
amino acid sequences, 5' UTR and 3' UTRDNAse-quences for
tilapia
IGF-II,
arecontrasted with those determinedby
cDNA sequence data. Thegenomic
structure of thetilapia
IGF-IIcoding region
showsgreatvariationwithpreviously
pub-lished mammal IGF-II gene
organization.
Construction and
expression of
IGF-IIexpression protein
In the first step, PCR was used to
amplify
IGF-II maturepolypeptides
ofDNAfragments,
whichwerethendigested
withEco RI. Inthe second
step,
weconstructed PCRproducts
withthe
pGEX-2T
vectorof Eco RIdigestion.
Theligation
products
were transformed into
BL21(DE3)
E. coli cells.Colony
hy-bridization andDNAsequencing
wereusedtoidentify
the DNA sequence and orientation.Toexpress thetilapia
IGF-IIrecom-binant
polypeptides,
theBL21(DE3)
E. coli cellsincluding
re-combinant IGF-II
plasmids
wereinduced with0.1 mMIPTG and grown for 3 hrat22°C.Figure
5 shows the total cellproteins
extracted from induced and noninducedE. coli cellculture;
aclear band of 36kDwasdetected after 0.1mMIPTG induction for 30-180 min. The
36-kD
protein
was notdigested
by
thrombin,
as afusionprotein
with GST. Lanes 2 to 7 show theprotein
bandpresent
in E.coli cell
containing pGEX2T
vextorligated
withthetilapia
IGF-II mature
polypeptide
DNA sequence. Lane 1 shows the pro-tein bandcontaining pGEX-2T
vectoronly.
The finalpurified
IGF-IIprotein
wascompared
by
denaturedpolyacrylamide gel.
This
protein
is not foundabundantly
in inclusion bodies.In-duced with 0.1 mM
IPTG,
afusionprotein
wasproduced
with anapparentmolecularweight
of 36 kD. The final recombinant12
3 4
5
6 7
12 3 4 5 630kDa—
GST
+IGF-II
GST
FIG. 5.Expression
oftilapia
EGF-IImaturepeptide
inE.coliBL21(DE3).
Cells were cultured in 2YT medium with 100¿ig/ml ampicillin
at37°C until theOD0oo
reached 0.4-0.6. Then thetemperature
was shifted to 22°C and 0.1 mMIPTG wasadded for induction of
tilapia
IGF-Imaturepeptide synthesis.
The cells wereharvested after
30, 60, 90, 120, 150,
and 180min
induction,
and the totalprotein
was extractedby lysis
buffer and
analyzed by
NaDodSOa-PAGE
on a 10%gel
with Coomassie Bluestaining.
Lane1,
Proteinexpressed by
pGEX-2Tvectoralone;
lane2,
cellscontaining
IGF-I insert afterin-duction for30
min;
lane3,
cellscontaining
IGF-I insert after induction for 60min;
lane4,
cellscontaining
IGF-I insertaf-terinduction for 90
min;
lane5,
cellscontaining
IGF-I insert afterinductionfor 120min;
lane6,
cellscontaining
IGF-Iin-sertafter induction for 150
min;
lane7,
cellscontaining
IGF-Iinsert after induction for 180min.
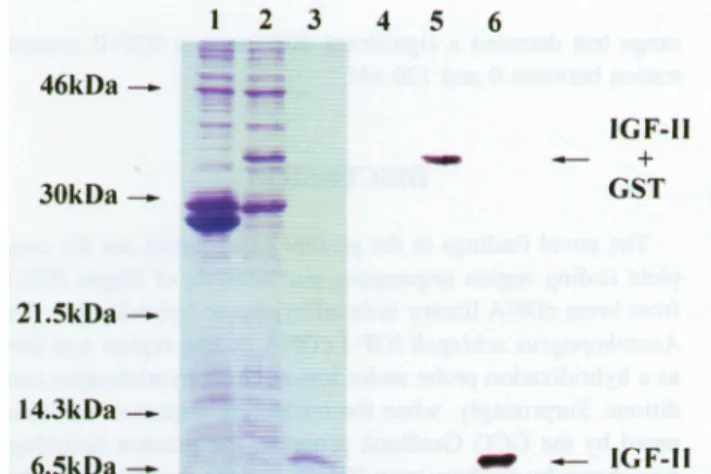
46kDa— 30kDa-~ 21.5kDa — 14.3kDa — 6.5kDa— IGF-II + GST — IGF-II
FIG. 6.
Expression
oftilapia
IGF-Imaturepeptide
inE.coliBL21(DE3).
The E. coli cell culture conditionswerethesame asinFig.
5. When the cell cultures wereharvested after 180 min ofinduction,
the fusionproteins
weredigested
with throm-bin andpurified by
RedPack Module(Pharmacia Biotech).
Theproteins
wereanalyzed by
NaDodSOa-PAGE
on a15%gel
with Coomassie Bluestaining
(A)
andimmunoblotting
(B).
Lane1,
E. coli culture
containing
thecloning
vector(pGEX-2T)
alone;
lane
2,
E. coli culturecontaining
thetilapia
IGF-IImature pep-tideDNAsequence constructed withpGEX-2T cloning
vector; lane3,
IGF-II maturepeptide digested by
thrombin andpuri-fied
by
RedPackModule;
lanes4—6,
monoclonal antibodies raisedagainst
thetilapia
IGF for ECL immunoreaction. Theloading
order of lanes 4-6 arethe same anin lanes1-3,
re-spectively.
single
band. ThepGEX-2T
vectorin cells alone and GST fu-sion IGF-IImaturepolypeptide
with and without thrombindi-gestion
were run onaNaDodS04-PAGE gel
as shown inFig.
6. ECLwestern
blotting analysis explained
thattheprotein
canbe detected with the
specifically
monoclonalanti-tilapia
IGFantibody
(Fig.
6).
The recombinant IGF-II
polypeptides
were identified with amino acidsequencing.
Theexpression
ofthe IGF-IIpolypep-tide and the
predicted
IGF-II maturepolypeptide comparison
are shown in Table 1. In Table 1, the
expression
of IGF-IIpolypeptide
shows anadditionalsevenamino acids before themature
polypeptides.
Of thesevenaminoacids,
sixare apGEX-2T multiclonal siteDNAsequencetotranslate amino
acids,
andoneis the ATGstartcodon.
Characterization
of tilapia
IGF-IIrecombinant
polypeptides
After thrombin
digestion,
thetilapia
IGF-II recombinant pro-tein revealedasingle
band of7 kDon adenaturedNaDodSOa
polyacrylamide gel.
Totestthebiological
function oftilapia
re-combinant IGF-II
polypeptides, they
wereanalyzed by
in vitroassay of
incorporation
of[3H]thymidine
intoDNA differentia-tion. The cell linedesignated
TO-2wasestablished from ovariesof
healthy
adulttilapia hybrids
(Tilapia
mossambicaX T.nilot-ica)
as anexperimental
cell line. Thetest concentrationswerebetween 0 and 120n/Vfand
incorporation ability significantly
increased over this concentration range(ANOVA;
F =4.46;
df=
6.14;
p<0.05).
Figure
7. shows that Duncanmultiple
Table 1. Different Amino Acid
Sequences
Betweenthe Predicted andActual ExpressionofRecombinant Polypeptides
amino
acid
sequence
Predicted
expression
amino
acid
EMASAETLCGGEL.
Actual
expression
amino acid
GSPGIHMEMASAETLCGGEL.
rangetestdetecteda
significant
difference in IGF-IIconcen-tration between0and 120nM.
DISCUSSION
The novel
findings
inthepresent
experiments
arethecom-plete coding region sequencing
andanalysis
oftilapia
IGF-II from braincDNAlibrary
isolatedby plaque hybridization.
TheAcanthopagrus schlegeli
IGF-I cDNAcoding region
wasusedas a
hybridization probe
underlow-strength hybridization
con-ditions.
Surprisingly,
when the nucleotide sequencewascom-pared by
the GCG GenBank program, thegreatest
homology
wasshown
by
rainbowtroutIGF-II,
and the secondwasby
hu-manIGF-II sequences. This extendsto
eight
the numberofan-imals
species
forwhichpublished
IGF-II sequence is available. Intilapia
and rainbow trout, thepeptides
arehighly
conserved. Predicted amino acids between fish and mammalsdifferedby
the addition of2 amino acids in the B
domain;
in the Cdo-main,
fish and mammals differedby
an increase of 3 amino acids.Achange
inthe IGF-I aminoacid sequenceatposition
B23-B25
(Phe-Tyr-Phe)
willgive
risetoadecreaseinability
of
binding
tothe IGF-Ireceptor
(Cascieri
etal,
1988).
These amino acid sequencesarealsopresent
atB26-B28inthetilapia
IGF-IImaturepeptide.
These
phenomena
existin all animals whoseIGF-II sequencehasbeen
published,
andthey
arestrictly
conserved.In mouseandrat
IGF-II,
position
B22 ischanged
from SertoGly
com-pared
withhumans;
Gly
is found atpositive
B22 in insulinsfrom all
species
except
hystricomorphs
(LeRoith, 1991).
But in10000r-1 . * 9000 L T
2
:J?
Q 8000 7/
1
.2 7000!"
*yS
W
jr 9 6000 -/ O * /YS
5000~:
7*^
M
4000-/
. / >ï 3000 -Öl*/
H : r V SC 2000 r/
1000 '-oF. , . i i i i i i . . i , . . i . . , i , 0 20 40 60 80 100 120 IGF-II(nM)
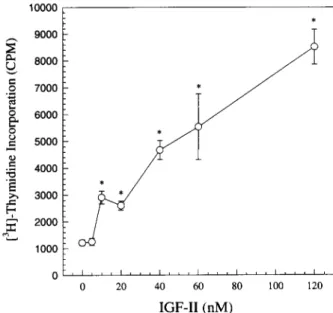
FIG.7. Effects ofrecombinant
tilapia
IGF-IIpolypeptide
onstimulated
tilapia
ovary cell(TO-2
cellline)
proliferation.
Thefollowing
effectsweremeasured with different concentrationsof recombinant
tilapia
IGF-IIstimulating incorporation
of[3H]thymidine
intoDNAsynthesis.
The data show that the TO-2cell membranemusthaveanIGFreceptor, andsorepresent
adose-dependent
effect.fish,
thisposition
B22 isGlu,
Why
is itnotGly
orSer? The real mechanism ispresently
unknown.Intilapia
IGF-I,
there isan inference A/-linked
glycosylation
site(Asn-X-Ser/Thr),
butit isnotfound in the
tilapia
IGF-IIpeptide.
The role of these IGF-IIpeptides,
whetherthey
haveaA/-linkedglycosylation
site ornot, remains for themostpartunknown in fish.InVitro,IGFs have very
important
functions and actionsonneuronal and
glial
cellfunction. The ribonucleaseprotection
as-say, insitu
hybridization,
andimmunohistochemistry
wereusedtodemonstrate thatIGF-IIis
synthesized predominantly
intheleptomeninges,
choroidplexus,
andparenchymal
microvascu-lature in rats, whichpresumably
represents
the site of IGF-IIbioactivity
within the brain(Logan
etal,
1994).
In adult rats, IGF-II mRNA canbe detected in brain and other organs, and alsocanbe detected in rainbowtrout.Except
in theliver,
lev-els of IGF-IImRNA in brain hasahigher
expression
thanto-talIGF-I in rainbowtrout
(Chen
etal,
1994).
So,
there is nodoubtthata
tilapia
IGF-II clonecanbeobtained fromabraincDNA
library.
Butinmostfishes,
theadenohypophysis
is dif-ferentiated intoarostralandaproximal
pars distalis andaparsintermedia. Whether IGF-II has any function infish
adenohy-pophysis
is still unclear.The exon
organization
of thetilapia
IGF-IIcoding region
gene is very dissimilartothat of mammalian and avian IGF-II genes. In
sheep
IGF-II genes, thepromoter
directs thetran-scription
of sixnoncoding
exonsandalternatively
splices
tothe sharedexons8, 9,
and 10(Ohlsen
etal,
1994). Up
to now,it has been determined that the IGF-II genes of mammals(hu-man, mouse,
sheep,
rat) (Frunzio
etal, 1986;
dePagter-Holthuizenet
al, 1987;
Soaresetal, 1986;
Rotwein andHall,
1990;
Holthuizen etal, 1990;
Ikejiri
etal, 1990, 1991;
vanDijk
etal, 1991;
Ohlsenétal,
1994)
and birds(chicken)
(Dar-ling
andBrickell,
1996)
have threecoding
exons of similarstructure; but in fish
(O.
mossambicus)
the IGF-II gene has fourcoding
exons. Aseparation
of IGF-IIgenestructurestrategy
issuggested
basedontherateof evolution of verterbrates.Com-mon
evolutionary history
for the insulin/IGFfamily
genes maybe duetothe
phylogeny
ofderived amino acid sequences.In-sulin and IGF genesarebelieved tohave evolved
by
repeated
duplication
anddivergence (Ellsworth
etal,
1994).
The IGF-IandIGF-II
separation
isconcludedtohavetakenplace
about 70million yearsago,which is aboutthesametime astheap-pearance of
placental
mammals(Rinderknecht
andHumbel,
1988).
Theseassumptions
arebasedonthepublication
of IGFsequences. It would seem that the
dissimilarity
ofthestruc-ture/function of the
coding
exon arrangement oftilapia
(Eu-teleostei)
compared
tobirdsandwarm-blooded vetebrates may have resulted fromhomoplastic
evolution.Tounderstand the IGF-II
protein
regulation
of fishphysiol-ogy, we have
developed
the GST-IGF-II fusionprotein
ex-pression
system. This isasingle-step purification
ofpolypep-tides
expressed
in E. coli as fusion withglutathione-S-transferase
(Smith
andJohnson,
1988).
Thelow-temperature
induction of fusion
protein synthesis
canimprove
solublepro-tein
production
(Hartman
etal,
1992).
Wetried manytemper-atureconditions and found that 22CCwassuitable for
purifica-tion of fusion
proteins.
The noveltilapia
IGF-IIprotein
wasexpressed
in E. coli andwashighly
activein the TO-2 cell line.Inrats, IGF-II
(50
ng/ml)
stimulatesoocyte
maturation(Feng
and IGF-Iweredetected
throughout sheep preimplantation
de-velopment
from the one-cellto theblastocyst
stages (Watson
etal,
1994).
IGF-II has
specific
andhigh-affinity binding
sites for IGF-IIreceptors
onwhole ovarian membranes andonovariansec-tions,
suggesting
the IGF-II/M6Preceptors
in ovarian tissuecanremodel and mediate IGF-II actionon
folliculogenesis
(Teissier
et
al,
1994).
In mostfish,
the structureof the ovarian follicle is similar.The ovary consists ofagranulosa
celllayer
andone ortwo outersublayers
of theca cells. The theca andgranulosa
layers
aredividedby
abasement membrane. Inhumans,
IGF-II mRNA isexpressed
in newborn ovarian stromaand in both newborn and adult ovaries(Zhou
andBondy,
1993).
Intheratovary, in situ
hybridization
and RNaseprotection
assayssug-gested
that the IGF-IIexpression
in theca-interstitial cells isspecific
tocelltype(Hernandez
etal,
1990).
In contrast, the amino acid sequences oftilapia,
avian,
and mammalian IGF-IImature
peptides
are very conserved(general
amino acidse-quences similaritiesare79% and
above).
Given theabove,
thesedata
suggest
thatmaturetilapia
IGF-IIpolypeptides
may have similar ovarian functions in fish ascompared
with those in mammals and birds. Thisexplains
why
the recombinanttilapia
IGF-IIprotein
canstimulatethymidine incorporation
andpre-sent
dose-dependent
effects in thetilapia
ovary cell line.ACKNOWLEDGMENTS
We thank Dr. Thomas T.
Chen,
Dr.Ching-Ming
Kuo,
andDr. Cho-Fat Hui for their
appropriate
and concise commentsaboutthis
experiment
andmanuscript.
We thank Dr. Wei-YuanChow for
kindly providing
thetilapia (hybrid)
brain cDNAli-brary
and Oreochromis mossambicusgenomic
DNAlibrary.
We thank Mr.Hung-Chih
Chen forproviding
theAcanthopa-grus
schlegeli
IGF-I cDNAplasmid.
We thank Dr. I-Chiu Liaofor his
support
andencouragement.Thisproject
wassupported
by
NSCgrants
NSC 85-2321-B-001-007-A15(R.O.C),
and NSC 86-2311-B-001-048-B24(R.O.C).
to Dr. Jen-Leih Wu.REFERENCES
BELL, G.I.,MERRYWEATHER, J.P., SÁNCHEZ-PESCADOR, R.,
STEMPIEN, M.M., PRIESTLEY, L., SCOTT, J.,andRALL, L.B. (1984).
Sequence
ofacDNA cloneencoding
humanpreproinsulin-like
growth
factor II. Nature310,775-777.BLUNDELL, S., BERDARKA, E., RINDERKNECHT, and
HUM-BEL,R.E. (1978). Insulin-like
growth
factor: A model fortertiary
structure
accounting
forimmunoreactivity
and receptor binding. Proc. Nati. Acad. Sei. USA75, 180-184.BREATHNACH, R.,andCHAMBÓN,P.(1981).
Organization
andex-pression
ofeucaryotic split
genescoding
forproteins.
Annu. Rev. Biochem.50,349-383.BREATHNACH, R., BENOIST, C, O'HARE, K., GANNON, F.,and
CHAMBÓN,P.(1978). Ovalbumingene:evidence foraleader
se-quence in mRNA and DNA sese-quencesatthe exon-intron boundaries.
Proc. Nati. Acad. Sei. USA75,4853-4857.
BROWN, W.M., DZIEGIELEWSKA, K.M., FOREMAN, R.C., and
SAUNDERS,N.R.(1990).The nucleotide and deducedaminoacid sequences of insulin-like
growth
factor II cDNAs from adult bovine and fetalsheep
liver. Nucleic Acids Res. 18,4614.CASCIERI, M.A., CHICCHI, G.G., APPLEBAUM, J., HAYES, N.S., GREEN, B.G.,andBAYNE,M.L. (1988).Insulin-like
growth
fac-torIwith reduced
affinity
for thetype 1 insulin-likegrowth
factorreceptor.
Biochemistry
27,3229-3223.CHEN, S.N., CHI-UENO, S.C.,andKOU,G.H.(1983).Acell line
de-rived from
Tilapia
ovary.Fish.Pathol.18(1), 13-18.CHEN, T.T., MARSH, A., SHAMBLOTT, M., CHAN, K.M.,TANG,
Y.L., CHENG, CM.,andYANG, B.Y.(1994).Structureand
evo-lutionof fish
growth
hormone and insulinlikegrowth
hormone andinsulinlike
growth
factorgenes. In FishPhysiology,vol. Xin. N.M. Sherwood and C.L. Hew, eds. (Academic Press, New York) pp.179-209.
COHICK, W.S., and CLEMMONS, D.R. (1993). The insulin-like
growth
factors.Annu. Rev.Physiol.
55,131-153.DARLING, D.C.,andBRICKELL,P.M.(1996).Nucleotide sequence
and
genomic
structureof the chicken insulin-likegrowth
factor-II (IGF-II)coding
region.Gen.Comp.
Endocrinol. 102,283-287.DeCHIARA, T.M., EFSTRATIADIS, A., and ROBERTSON, E.J. (1990). A
growth deficiency phemotype
inheterozygousmicecar-rying
an insulin-likegrowth
factorII genedisrupted by targeting.
Nature345,78-80.
DeCHIARA, T.M., ROBERTSON, E.J., and EFSTRATIADIS, A. (1991). Parental
imprinting
ofthemouseinsuline-likegrowth
factorHgene. Cell64,849-859.
DEMMER, J., HILL, D.F.,andPETERSEN,G.B.(1993). Characteri-zation oftwo
sheep
insulin-likegrowth
factor-II cDNAs withdif-ferent5'-untranslated
regions.
Biochim.Biophys.
Acta1173,78-80. De PAGTER-HOLTHUIZEN, P., JANSEN, M., VAN SCHAIK,F.M.A., VAN DER KÄMMEN, R.A., OOSTERWIJK, C, VAN DENBRANDE, J.L.,andSUSSENBACH,J.S.(1987).The human
insulin-like
growth
factorIIcontainstwodevelopment-specific
pro-moters.FEBSLett.214,259-264.
DE PAGTER-HOLTHUIZEN, P., JANSEN, M., VANDER
KÄM-MEN, R.A., VAN SCHAIK, R.M.A., and SUSSENBACK, J.S. (1988). Differential
expression
of the human IGF-II gene: charac-terizationof theIGF-IImRNAs andanmRNAencoding
aputative
IGF-II associated
protein.
Biochim.Biophys.
Acta. 950,282-295.DUGUAY, S.J., LAI-ZHANG,J.,STEINER,D.F., FUNKENSTEIN, B.,andCHAN,S.J.(1996).
Developmental
andtissue-regulated
ex-pression
ofIGF-I and IGF-II mRNAs inSparus
aurata.J. Mol. En-docrinol. 16,123-132.DULL, T.J., GRAY, A., HAYFLICK, J.S.,andULLRICH,A.(1984). Insulin-like
growth
factorIIprecursorgeneorganization
in relationtoinsulin gene
family.
Nature310,
777-781.ELLSWORTH, D.L., HEWETT-EMMETT, D.,andLI,W.H.(1994). Evolution of base
composition
in the insulin and insulin-likegrowth
factor genes. Mol. Biol. Evol. 11,875-885.FENG, P., CATT, K.T., and KNECHT, M. (1988).
Transforming
growth
factor-beta stimulates meiotic maturation of theratoocyte.Endocrinology
122,181-186.FRUNZIO, R„ CHIARIOTTI, L., BROWN, A.L., GRAHAM, D.E., RECHLER,M.M.,andBRUNI,C.B. (1986).Structure and
expres-sion of theratinsulin-like
growth
factorII(rIGF-II)gene. rIGF-IIRNAs are transcribes from two promoters. J. Biol. Chem. 261, 17138-17149.
GRAY, A., TAM, A.W., DULL, T.J., HAYFLICK, J., PINTAR, J., CAVENEE, W.K., KOUFOS, A.,andULLRICH,A.(1987).
Tissue-specific
anddevelopmentally
regulatedtranscription
ofthe insulin-likegrowth
factor 2 gene. DNA6,283-295.HAIG, D., and GRAHAM, C. (1991). Genomic
imprinting
andthestrangecaseoftheIGF-IIreceptor.Cell64,1045-1046.
HARTMAN, J., DARAM, P., FRIZZELL, R.A., RADO, T., BEÑOS, DJ., andSORSCHER,E.J.(1992).
Affinity purification
ofinsolu-blerecombinantfusion
proteins containing glutathione-S-transferase.
Biotechnol
Bioeng.
39,828-832.D.,andADASHI,E.Y.(1990). Ratovarian insulin-like
growth
fac-torII gene
expression
istheca-interstitial cell-exclusive:Hormonalregulation
andreceptordistribution.Endocrinology
127,3249-3251.HOLM, N.R., HANSEN, L.B.H.,NILSSON,C,andGAMMELTOFT,
S. (1994).Gene
expression
and secretion of insulin-likegrowth
fac-tor-II and insulin-likegrowthfactor binding protein-2
from culture sheepchoroidplexus epithelial
cells. Brain Res. Mol. Res.21,67-74.HOLTHUIZEN, P., VAN DER LEE, F.M., IKEJIRI, K.,
YA-MAMOTO, M„andSUSSENBACH,J.S.(1990). Identification and initial characterization ofafourth leaderexonandpromoterof the
human IGF-II gene.1087,341-343.
HOOVER, D., FRIEDMANN, M., REEVES, R., andMAGNUSON,
N.S.(1991).Recombinant human
pim-1
exhibits serine/threoninelá-ñase
activity.
J.Biol. Chem.266,14018-14023.HUMBEL, R.E. (1990). Insulin-like
growth
factors I and II. Eur JBiochem.
190,
445^162.IKEJRI, K., UENO, T., MATSUGUCHI, T., TAKAHASH1, K., ENDO, H.,andYAMAMOTO,M.(1990).The
primary
structureoftheratinsulin-likegrowthfactorII gene
region.
Biochim.Biophys.
Acta1049,350-353.IKEJRI, K., FURUICHI, M., UENO, T., MATSUGUCHI, T., TAKA-HASHI,K.,ENDO,H,andYAMAMOTO,M. (1991).The
pres-ence andactive
transcription
of threeindependent
leaderexons inthemouseinsulin-like
growth
factor II gene. Biochim.Biophys.
Acta 1089,77-82.JANSEN, M., VAN SCHAIK, F.M., VAN TOL, H, VAN DEN
BRANDE, J.L., and SUSSENBACH, J.S. (1985). Nucleotide
se-quencesofcDNAs
encoding
precursorsofhuman insulin-likegrowth
factorII(IGF-II)andanIGF-II variant. FEBSLett. 179,243-246.
JONES, J.I.,andCLEMMONS,D.R.(1995).Insulin-like
growth
fac-torsand their
binding proteins: Biological
actions. Endocrine Rev.16,
3-34.KOLAND, JG., O'BRIEN, K.M., andCERIONE, R.A. (1990).
Ex-pression
ofepidermal growth
factorreceptorsequenceasE. coli fu-sionproteins:
Applicationin thestudy
oftyrosine
kinase function. Biochem.Biophys.
Res. Commun. 166,90-100.KWANG, J., BOLINE, S., LITTLEDIKE, E.T., DUBOVI, E.J., and DONIS, R.O.(1991).
Expression
of thep80
region
of bovine viral diarrhea virus and identification ofspecific
antibodies to thisre-combinant
protein
inbovinesera.Biochem.Biophys.
Res. Commun. 178, 1326-1334.LAUTERIO, T.J., MARSON, L„ DAUGHADAY, W.H.,andBAILE, C.A.(1987). Evidencefor the role ofIGF-II in the control offood intake.
Physiol
Behav.40,755-758.LEE, J.E., PINTAR, J.,andEFSTRATIADIS,A.(1990).Patternofthe insulin-like
growth
factor II geneexpression during early
mouseem-bryogenesis. Development
110, 151-159.LeROITH,D.(1991).Insulin-like GrowthFactors:Molecularand Cel-lularAspects. (CRCPress,BocaRaton,FL).
LOGAN, A., GONZALEZ, A.M., HILL, D.J., BERRY, M.,
GREG-SON, N.A.,andBAIRD,A.(1994).Coordinatedpattern of
expres-sion and localization of insulin-like growth factor-II (IGF-II) and
IGF-binding protein-2
in the adult rat brain.Endocrinology
135, 2255-2264.LUTHI, C,ROTH,B.V.,andHUMBEL,R.E.(1992).Mutants of
hu-manIGF-II. Eur. J.Biochem.205,483^190.
MANIATIS, T., FRITSCH, E.F.,andSAMBROOK,J. (1982).
Mole-cularCloning:A
Laboratory
Manual. ColdSpring
HarborLabora-tory,Cold
Spring
Harbor,NewYork).OHLSEN, S.M., LUGENBEEL, K.A.,andWONG,E.A.(1994).
Char-acterizationofthe linked ovine insulin and insulin-like
growth
fac-torII genes. DNA Cell Biol.13,377-388.
RAPPOLEE, D.A., STURM, K.S., BEHRENDTSEN, O., SCHULTZ, G.A., PEDERSEN, R.A.,andWERB,Z.(1992).Insulin-like
growth
IIactsthroughanendogenous growth pathway regulated by
imprit-ing
inearly
mouseembryos.
Genes & Dev.6,939-952.RINDERKNECHT, E., andHUMBEL,R.E.(1978).The amino acid sequenceofhuman insulin-like
growth
factor I and its structuralho-mology
withproinsulin.
J.Biol. Chem. 253,2769-2776.REINECKE,M., WEIMAR, E., MAAKE, C, DRAKENBERG, K., FALKMER, S.,andSARA,V.R.(1994).IGF-2-like
peptides
arepre-sentin insulin cellsoftheelasmobranchianendocrine pancreas: An immunohistochemical and
Chromatographie study. Histochemistry
102,365-371.ROTWEIN, P.,andHALL,L.J. (1990).Evolution of IGF-II: charac-terization of themouseIGF-II gene andidentificationoftwo
pseudo-exons.DNACell Biol.
9,
725-735.SANGER, F., NICKLIN, S., andCOULSEN, A.E. (1997). DNA
se-quencing
withchain-terminating
inhibitors. Proc. Nati. Acad. Sei.USA74,5463-5467.
SHAMBLOTT, M.J.,andCHEN,T.T.(1992).Identification ofa
sec-ondinsulin-like
growth
factor inafishspecies.
Proc. Nati. Acad. Sei.USA89, 8913-8917.
SMITH, D.B.,andJOHNSON,K.S.(1988).
Single-step purification
ofpolypeptides expressed
in Escherichia coliasfusions withglutathione
S-transferase. Gene67,31-40.SOARES, M.B., ISHII, D.N., and EFSTRATIADIS, A. (1985).
De-velopment
andtissue-specific expression
ofafamily
oftranscripts
relatedto ratinsulin-likegrowth
factor II mRNA. Nucleic Acids Res. 13, 1119-1134.SOARES, M.B.,TÜRKEN,A., ISHII, D„ MILLS, L., EPISKOPOU, V., COTTER, S., ZEITLIN, S.,andEFSTRATIADIS,A.(1986).Rat insulin-like
growth
factor II gene: Asingle
gene withtwopromot-ers
expressing
amultitranscript family.
J. Mol. Biol.192,737-752.STEMPIEN, M.M., FONG, N.M., RALL, L.B.,andBELL,G.I.(1986).
Sequence
of aplacental
cDNAencoding
the mouse insulin-likegrowth
factor II precursor. DNA.5,357-361.TEISSIER, M.P., MONGET, P., MONNIAUX, D„andDURAND,P. (1994).
Changes
in insulin-likegrowth
factor-II/mannose-6-phos-phate
receptorduring growth
and atresia of ovineovarian follicles.BiolReprod.
50, 111-119.VAN DIJK, M.A., VAN SCHAIK, F.M.A., BOOTSMA, H.J.,
HOLTHUIZEN, P.,andSUSSENBACH,J.S.(1991).Initial charac-terization of the fourpromotersofthe human insulin-like
growth
fac-torIIgene. Mol. CellEndocrinol.81,81-93.
WATSON, A.J., WATSON, P.H., ARCELLANA-PANLILIO, M., WARNES, D., WALKER, S.K., SCHULTZ, G.A., ARMSTRONG, D.T.,andSEAMARK,R.F.(1994).A
growth
factorphenotype
map forovinepreimplantation development.
Biol.Reprod.
50,725-733.WILLISON,K. (1991).
Opposite imprinting
ofthemouseIGF-II and IGF-IIreceptorgenes. Trends Genet.7,107-109.ZHOU, J.,andBONDY,C.(1993). Anatomyof the insulin-like
growth
factor systemin the human testis. FértilSteril.60,897-904.
Address
reprint
requests
to:Dr. Jen-Leih Wu
Institute
of Zoology
Academia Sínica
Nankang, Taipei
Taiwan