Computer-aided Diagnosis Based on Speckle Patterns for
Ultrasound Images
Woo Kyung Moon1, Chung-Ming Lo2, Chiun-Sheng Huang3, Jeon-Hor Chen4,5, and Ruey-Feng Chang2,6*
1Department of Radiology, Seoul National University Hospital, Seoul, Korea 2Department of Computer Science and Information Engineering
National Taiwan University, Taipei, Taiwan
3Department of Surgery, National Taiwan University Hospital and National Taiwan University College of Medicine, Taipei, Taiwan
4Center for Functional Onco-Imaging and Department of Radiological Science University of California Irvine, California, USA
5Department of Radiology, China Medical University Hospital, Taichung, Taiwan 6Graduate Institute of Biomedical Electronics and Bioinformatics
National Taiwan University, Taipei, Taiwan
* Corresponding Author:
Professor Ruey-Feng Chang
Department of Computer Science and Information Engineering National Taiwan University
Taipei, Taiwan 10617, R.O.C. Telephone: 886-2-33664888~331 Fax: 886-2-23628167 E-mail: [email protected] 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28
Abstract
For breast ultrasound, the scatterer number density from backscattered echo was
demonstrated in previous research to be a useful feature for tumor characterization. To
take advantage of the scatterer number density in B-mode images, spatial compound
imaging was obtained, and the statistical properties of speckle patterns were analyzed
in this study for use in distinguishing between benign and malignant lesions. A total
of 137 breast masses (95 benign cases and 42 malignant cases) were used in the
proposed computer-aided diagnosis (CAD) system. For each mass, the average
number of speckle pixels in a region of interest (ROI) was calculated to utilize the
concept of scatterer number density. In addition, the first-order and second-order
statistics of the speckle pixels were quantified to obtain the distributions of the pixel
values and the spatial relations among the pixels. The performance of the speckle
features extracted from each ROI was compared with the performance of the
segmentation features extracted from each segmented tumor. As a result, the proposed
CAD system using the speckle features achieved an accuracy of 89.1% (122/137), a
sensitivity of 81.0% (34/42), and a specificity of 92.6% (88/95). All of the differences
between the speckle features and the segmentation features are not statistically
significant (p>0.05). In a receiver operating characteristic (ROC) curve analysis, the 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46
Az value of the speckle features was significantly better than the Az value of the
segmentation features (0.93 vs. 0.86, p=0.0359). The performance of this approach
supports the notion that the speckle patterns induced by the scatterers in tissues can
provide information for classifying tumors. The proposed speckle features, which
were extracted readily from drawing an ROI without any preprocessing, also provide
a more efficient classification approach than tumor segmentation.
Keywords: Speckle, Breast cancer, Ultrasound, Spatial compound imaging,
Computer-assisted diagnosis 47 48 49 50 51 52 53 54 55
Introduction
Breast ultrasound (US), including spatial compound imaging (SCI), is being
explored for distinguishing between benign and malignant lesions (Stavros et al.
1995; Cha et al. 2005; Cha et al. 2007). The sonographic appearances of breast tumors
are constructed by means of acoustic transmission and are interpreted by radiologists
upon clinical examination for a diagnosis. To standardize the terminology used for
describing tumors , the BI-RADS lexicon was proposed by the American College of
Radiology (American College of Radiology 2003). According to the descriptors
defined in BI-RADS descriptive categories, the dominant sonographic findings of a
tumor are classified and analyzed by radiologists to evaluate the likelihood of
malignancy. The descriptive categories include shape, orientation, margin, lesion
boundary, echo pattern, and posterior acoustic features. With the quantification of the
descriptors used clinically, various computer-aided diagnosis (CAD) systems
(Rangayyan et al. 2000; Shen et al. 2007a; Shen et al. 2007b; Nie et al. 2008) were
developed to provide an efficient procedure to diagnose breast tumors automatically.
By tumor segmentation, the tumor characteristics were extracted and quantified to
distinguish between benign and malignant lesions. The quantitative features utilized in
these CAD systems can be classified as morphology or texture features. Morphology 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73
features are used to describe the tumors’ shape characteristics, and texture features are
used to show the echogenicity properties through the correlations among pixels. Furthermore, the speckle patterns from backscattered echo were analyzed to
provide information for tissue characterization (Tuthill et al. 1988). Speckle is
generated by the constructive and destructive interference of US waves backscattered
from tissue scatterers, which are tissues with equal or smaller structures than the US
wavelength. From observing the backscattered echoes, the effects of the scatterer
number density can be shown by the statistical properties when the number per
resolution cell is small (i.e., < 10). The amplitude histogram of the signal with a
sparse scatterer number density has a pre-Rayleigh distribution. With the increasing
number of scatterers, the speckle is fully developed, and the corresponding histogram
approaches a Rayleigh distribution. Based on the analyzed results, the backscattered
echo was further modeled by the Nakagami parameter to distinguish between benign
and malignant lesions (Shankar et al. 2003; Chang et al. 2010). The Nakagami
parameter effectively quantified the statistics of the backscattered echo from different
scattering conditions, including pre-Rayleigh, Rayleigh and post-Rayleigh. In the
previous studies, the speckle patterns from backscattered echo signal were
demonstrated to be useful in classifying tumors. However, the signal data are not easy
to acquire and are less familiar to radiologists who generally use B-mode images to 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92
evaluate tumor characteristics.
In this study, we analyzed the speckle patterns in B-mode images obtained with
SCI to develop more useful features for tumor classification. For speckle extraction, a
tumor and its surrounding tissues were included in a region of interest (ROI) that was
cropped from a B-mode image. Next, the statistical properties of the speckle pixels
extracted from an ROI were quantified as features for distinguishing between benign
and malignant lesions. Moreover, the diagnostic performance of the speckle features
was compared with the performance of the morphology and texture features obtained
by tumor segmentation.
Materials and Methods
US acquisition
The breast US images collected in this study were acquired using an ATL HDI
5000 scanner (Philips, Bothell, WA). The applied L12-5 probe was a 192-element
linear array transducer with variable frequency ranging from 5 to 12 MHz and a scan
width of 38 mm. We obtained image data sets with and without SCI (SonoCT™,
Philips). The SCI method combined nine frames produced from different
transmit angles to reduce speckle and shadowing. In our experiment, conventional
US images were first obtained without SCI mode, and SCI images were subsequently 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111
obtained in an identical plane without changing depth, focus position, or gain settings.
The gray-level value of the image pixels ranges from 0 to 255.
A total of 137 masses were examined based on the B-mode images produced by
the scanner from April 2002 to May 2003. Two breast radiologists with 5 and 15
years of experience, respectively, classified the lesions into BI-RADS assessment
categories according to the observation of the B-mode images before biopsy. There
were 69 lesions (50%) in RADS 3 (probably benign), 53 lesions (39%) in
BI-RADS 4 (suspicious abnormality), and 15 lesions (11%) in BI-BI-RADS 5 (highly
suggestive of malignancy).
The lesions of all patients that were pathologically proven by core needle biopsy
or fine-needle aspiration cytology included 95 benign lesions (69%) and 42 malignant
lesions (31%). The lesion sizes ranged from 0.4 to 3.0 cm (mean: 1.3 ± 0.6 cm). In the
benign lesions, there were 74 cases of fibroadenoma and 21 cases of fibrocystic
changes. The size was 0.4-2.5 cm (mean: 1.2 cm). In the malignant lesions, there
were 40 cases of invasive ductal carcinoma, 1 case of invasive tubular carcinoma, and
1 case of invasive papillary carcinoma. The size was 0.6-3.0 cm (mean: 1.6 cm). The
patients with benign lesions had a mean age of 42 (range 22–64), and the patients with
malignant lesions had a mean age of 49 (range 34–77). For this study, approval was
obtained from our institution review board, and informed consent was waived. 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130
ROI
A subimage of a B-mode image was cropped into an ROI. To specify a tumor
region, a rectangular bounding box was used to enclose a tumor. In principle, there
was a tumor in the center of the ROI, leaving a distance of 5–10 pixels between the
tumor boundary and the bounding box. According to the tumor size, the smallest ROI
was 55 × 42 pixels (0.57 cm × 0.44 cm), and the largest one was 290 × 160 pixels
(3.02 cm × 1.67 cm). The illustration of ROI selection is shown in Fig. 1 (a).
Speckle features
A US B-mode image is composed of the backscattered echoes reflected from
tissues. With the transmission of the US pulses, tissues generate various responses to
the pulses, such as absorption, reflection, and scattering, according to their physical
properties. The scatterers with microstructure contained in tissues, such as tissue
parenchyma, scatter the US pulses and produce an interference pattern called speckle.
In this study, the speckle pixels in B-mode images were extracted to generate useful
features in tissue characterization.
The speckle patterns have granular appearances with small difference in gray
level. For fully developed speckle, the number of scatterers is considerable. The
intensity image should have an exponential distribution and a constant ratio of the 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150
mean to the standard deviation (SD) of 1.0 (Tuthill et al. 1998). According to this
speckle property, the B-mode images were first log decompressed to obtain the raw
intensity images (Smith and Fenster 2000). The decompression procedure was
performed to extract the speckle pixels more precisely. The pixel value of the
acquired image data was log compressed to a 0-255 log scale to reduce the dynamic
range. To obtain the raw intensity value, the log decompression converted the 0-255
log scale back to a linear scale. The gray value of the pixels was converted by the
equation
I(x , y)=10G (x , y) ∕ G0
(1) where G(x,y) is the pixel value in B-mode images and G0, which is a linearization
factor related to the frequency of the transducer (Smith and Fenster 2000), converts
G(x,y) to a linear scale. The G0 value was used to determine the range of the intensity values. After the conversion, I(x,y) is obtained as the acoustic intensity. Next, a
moving 5×5 window is used to find a region with a range of ratios of mean intensity
to SD of 0.8–1.2 in the raw intensity image. If a region satisfies the condition, the
center pixel of the region is defined as speckle.
After the extraction of speckle pixels, the statistical properties of the speckle
pixels were converted into parameters for tissue characterization. In a previous study
(Tuthill et al. 1988), the scatterer number density from backscattered echo is useful 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169
for tumor characterization. For B-mode images, the average number of speckle pixels
in an ROI is calculated as
Savgnum=Snum ∕ ROInum (2)
where S_num is the number of total speckle pixels in an ROI, and ROI_num means
the number of all pixels in an ROI. An illustration is presented in Fig. 1. The ROIs of
a benign tumor and a malignant tumor are in Fig. 1 (b) and (d), respectively. The
speckle pixels extracted from the ROIs are shown in Fig. 1 (c) and (e) by a white
appearance (with a pixel value of 255 on the 8 bit gray-level image) for visualization.
Clearly, the density of speckle pixels in the ROI of the benign tumor (Fig. 1 (c)) is
higher than that of the malignant tumor (Fig. 1 (e)).
In addition to S_avgnum, the first-order statistics and second-order statistics of
the extracted speckle pixels in the ROI are utilized. Among the first-order statistics,
the mean and SD of the qualified mean/SD (0.8–1.2) in a moving window are defined
as Smean=
(
∑
P∈S mSD (P))
∕ Snum (3) SSD=√
(
∑
P∈ S (mSD ( P)−Smean) 2)
∕ Snum , (4)where mSD(P) is the mean/SD value of a speckle pixel. Also, the mean and SD of the
extracted speckle pixels in the 256 gray-level are defined as 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187
Sgmean=
(
∑
P∈ S G(P))
∕ Snum (5) SgSD=√
(
∑
P∈ S (G ( P)−Sgmean)2)
∕ Snum , (6) where G(P) is the gray-level value of a speckle pixel.With regard to the spatial relations between speckle pixels, textures, as the
second-order statistics, are assumed to describe the regional information. In this study,
the gray-level co-occurrence matrices (GLCM) (Haralick et al. 1973) are employed to
provide texture features. In this work, 21 texture features, which were implemented to
predict the likelihood of malignancy of the tumors in a classifier, are listed in Table 1.
Segmentation features
In this study, the speckle features obtained from an ROI were compared with the
features extracted from a segmented tumor. To segment a tumor, the level set method
was implemented (Huang et al. 2011). The quantitative features obtained from
segmentation can be classified into two categories: morphology features and texture
features (Rangayyan et al. 2000; Shen et al. 2007a; Nie et al. 2008; Chang et al.
2011). Morphology features were proposed to describe the tumor shape, and texture
features were proposed to represent the tumor echogenicity.
Morphology features 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206
After segmentation, the tumor contour is delineated by the level set method.
Next, the geometric characteristics of the tumor contour can be described by
quantitative morphology features. In past research (Rangayyan et al. 2000; Shen et al.
2007a; Nie et al. 2008; Chang et al. 2011), a variety of morphology features have
been suggested to estimate tumor shape, orientation, and margins. Instead of
measuring tumors directly, the best-fit ellipse was utilized for approximating the size
and position of a tumor (Shen et al. 2007b). By analyzing the properties of the best-fit
ellipse, the close measurement of tumor characteristics was accomplished. For
example, the angle of the major axis of the ellipse was used to calculate the tumor
orientation. Comparing the ellipse perimeter and the tumor boundary helped to
determine the smoothness of the tumor.
Additionally, the inherent properties of tumors were utilized to develop other
morphology features. Rangayyan et al. (2000) used the tumor perimeter and area to
estimate the compactness of a tumor. Nie et al. (2008) described the roundness of a
tumor by the normalized radial length (NRL), which was defined as the Euclidean
distance between the tumor center and the pixels on the tumor boundary normalized
by the maximum distance.
Texture features 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225
Texture features were used to describe the characteristics of the specific pattern
in a region. After segmentation, the tumor region is defined as the pixels surrounded
by the tumor contour. Therefore, the analyses of the pixel values inside the tumor
region were used as the texture features. In B-mode images, different tissues inside
the tumor reflect different echogenicity patterns that result in various distributions of
gray-level values. GLCM (Haralick et al. 1973), which calculates the spatial
correlations among pixels, was used in this study to provide the texture information
inside the tumors. Furthermore, the average intensity of the tumor was compared with
that of surrounding tissues and posterior shadowing to describe the tumor
characteristics (Shen et al. 2007b). In Table 2, a total of 38 quantitative features
mentioned above were collected for predicting the likelihood of tumor malignancy in
a classifier.
Statistical analysis
The quantitative features mentioned above were implemented in our experiment
to distinguish between benign and malignant lesions. For this purpose, the speckle
features were evaluated if they exhibited discriminability. According to the
distribution of the feature values, different test methods were employed. At first, the
distribution was determined by the Kolmogorov-Smirnov test (Field 2009). If the 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244
distribution of a feature was normal, then Student’s t-test (Field 2009) was applied.
Nonnormal distributions were analyzed using the Mann-Whitney U test (Field 2009).
The test result was quantified using p values. A p value of less than 0.05 was
considered to be statistically significant. For classifying masses, only the significant
features were used in the classifier.
The classifier used in this study was the binary logistic regression model
(Hosmer 2000). To classify the tumors as malignant or benign, the significant features
were gathered in the classifier. Next, backward elimination was applied to select
features from the significant features in the stepwise procedure. While the lowest error
rate was achieved in the trained classifier, a subset of features was selected as the
most relevant for classifying tumors. The performance of the classifier was examined
by the leave-one-out cross-validation method (Alpaydin 2004). If there were K cases
involved in the validation, the cases were trained K times. In every instance, one case
was left out of the K cases and was used to test the result trained using the remaining
K-1 cases.
According to the biopsy-proven pathology, the classification result was
evaluated based on five performance indices: accuracy, sensitivity, specificity,
positive predictive value (PPV), and negative predictive value (NPV). Using the
chi-square test, the performance indices of the speckle features, which were drawn out 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263
from ROIs, were compared with those of the segmentation features. Moreover, the
trade-offs between sensitivity and specificity achieved by the two feature sets were
compared using the receiver operating characteristic (ROC) curve. The Az value,
which represents the normalized area under the curve, was measured for comparison
via the z-test. ROCKIT software (C. Metz, University of Chicago, Chicago, IL, USA)
was used to analyze the ROC curve, and all other test methods were performed with
SPSS software (version 16 for Windows; SPSS, Chicago, IL, USA).
Results
For detecting speckle pixels, the performances of different window sizes were calculated and listed in Table 3. The detected speckle pixels were zero in most cases while using window sizes bigger than 9 × 9. Therefore, the performances of 3 × 3, 5 × 5, and 7 × 7 were compared. Although the differences were not significant, the result achieved by 5 × 5 was better than 3 × 3 and 7 × 7. We adopted 5 × 5 as the window size for the following calculations.
To evaluate the effect of speckle reducing techniques, the speckle features
extracted from conventional US and SCI were compared. Figure 2 (a) shows an ROI
of a B-mode image generated by conventional US, and Fig. 2 (c) shows the identical
case generated by SCI. The extracted speckle pixels of the ROIs are depicted with a 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282
white appearance in Fig. 2 (b) and (d). The illustration reveals that SCI reduced a little
speckle in the B-mode images.
By Student’s t-test or the Mann-Whitney U-test, the speckle features and the
segmentation features were evaluated to determine whether they were significant in
distinguishing between benign and malignant lesions. The significant speckle features
and the significant segmentation features are shown in Table 4 and Table 5,
respectively. Next, the significant features were used in the classifier to predict
whether the tumors are benign or malignant. In Table 6, the performances achieved by
different feature sets are listed. First, the differences of all performance indices
between the speckle features extracted from conventional US and SCI are not
significant (p>0.05). Note that the significant speckle features listed in Table 4 were
evaluated based on SCI. For conventional US, the number of significant speckle
features is one less than SCI (Inertia ave). In further comparisons, the speckle features
extracted from ROIs with SCI were used.
Using the chi-square test, the differences in the five performance indices
between the speckle features and the segmentation features are not significant
(p>0.05). Significant differences were observed in the comparison of Az using the
z-test (p=0.0359). The performance of the combined feature set including the speckle
features and the segmentation features was also calculated. This comparison showed 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301
that the Az of the combined feature set was slightly better than the Az of the speckle
features and was significantly better than the Az of the segmentation features
(p=0.0219). To illustrate, the ROC curves of the speckle features extracted from
conventional US and SCI are shown in Fig. 3 (a). In Fig. 3 (b), the ROC curves of the
speckle features with SCI, the segmentation features, and the combined feature set are
presented.
Two cases are illustrated to show the performance of the speckle features. The
malignant tumor in Fig. 4, which has an S_avgnum value of 0.13, was classified
correctly by the speckle features and was misclassified by the segmentation features.
Another example of a benign tumor is shown in Fig. 5. A tumor with an S_avgnum
value of 0.17 was classified correctly by the speckle features and was misclassified by
the segmentation features.
In order to refine the features, the cases misdiagnosed by the speckle features
were analyzed in terms of margin, echogenicity, calcification, and shadowing.
Echogenicity was related to the misclassification. First, if the surrounding tissues of a
tumor included in an ROI were hypoechogenic, the image composition may affect the
classification result. Second, while a malignant tumor and the surrounding tissues had
similar levels of isoechogenicity, the tumor was regarded as benign. Tumor margin,
calcification, and shadowing did not affect the performance. 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320
Discussion
Various US CAD systems were developed based on the features used by
radiologists on clinical examination. The features can be observed by human vision
and thus are used to describe the appearances of tumors, such as their shape,
orientation, and margin. In these CAD systems, the feature extraction and
quantification procedure, which affects the final performance of the classification, is
highly dependent on the segmentation result. However, the computation required in a
segmentation procedure is considerable, and it is inevitable that a well-segmented
result will be necessary.
With respect to the signal features suggested in other research studies, the
analysis of the scatterer number density from the backscattered echo is also useful in
classifying tumors (Shankar et al. 2003; Chang et al. 2010). Nevertheless, general US
scanners are not designed to obtain the signal data conveniently. In this study, the
speckle pixels in common B-mode images were extracted from ROIs to utilize the
scatterer properties. The first-order and second-order statistics of the extracted speckle
pixels were quantified into the features for tumor classification and were compared
with the features from segmentation. To the best of our knowledge, this study 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338
represents the first attempt to use the speckle density and the first order statistics of
speckle pixels in common B-mode images for breast tumor classification. Furthermore, GLCM texture features (Haralick et al. 1973) were utilized in this study. Conventionally, texture features were used to characterize the correlations among pixels inside the tumor. In our CAD system, only the speckle pixels detected in a ROI were considered. The diagnostic accuracy of proposed speckle features was better than the accuracy of the Nakagami parameter in the literature (89% vs. 82%)(Chang et al. 2010). Considering the trade-offs between sensitivity and specificity, the proposed speckle features was also better in the comparison of Az (0.93 vs. 0.81). The proposed speckle features were based on B-mode image which is the basic function of widely available US scanners. The convenience of the proposed CAD system can make it be widely used for every common US scanner. We obtained SCI and conventional images with and without
the use of the scanner’s SonoCT feature.
In our study, the speckle features extracted from B-mode images obtained by
using both conventional US and SCI achieved good performance (Az=0.89, Az=0.93),
thereby suggesting that the speckle features can be used in these two settings of the
US scanner. For comparison with the segmentation features, the speckle features
obtained with SCI were used. The reason for this choice is that a relatively large 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357
amount of noise appeared in the conventional US and caused the failure of tumor
segmentation. In comparison, the performance of five indices achieved by the speckle
features obtained with SCI is close to the performance achieved by the segmentation
features. Considering the trade-offs between sensitivity and specificity, the Az of the
speckle features is significantly better than the Az of the segmentation features (0.93
vs. 0.86, p=0.0359). By combining the speckle features and the segmentation features,
the combined feature set also performed significantly better than the segmentation
features (p=0.0219). The result indicates that the proposed speckle features provide
equal or better diagnostic information than the conventional segmentation features.
For the likelihood of malignancy, the combined feature set can determine the
classification result with greater accuracy. That is, incorporating the morphological
properties of a tumor with the underlying scatterer characteristics improves the
reliability of the CAD system.
For clinical use, the proposed CAD system, which employs the speckle features
to classify tumors, is expected to be a more efficient procedure than tumor
segmentation. In our experiment, calculating the speckle features of a total of 137
cases took 121 seconds using a computer with an Intel® Core™2 Quad CPU Q9400
at 2.66 GHz 2.67 GHz and 3.25 GB memory (Intel, Santa Clara, CA, USA). After
acquiring the ROIs, the average processing time for the proposed CAD system to 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376
classify each case was 0.88 seconds. In clinical examination, the speckle density can
immediately be displayed visually by a white appearance on the screen. The suggested
classification result can be shown almost in real-time while the radiologists specify
the tumor area. Generally, the routine procedure for radiologists to mark the tumor
size on a B-mode image is sufficient to specify the tumor area. The extra time for the
processing of the underlying CAD system would not be noticeable.
The limitation of this study is that the B-mode images were acquired from one
US scanner (ATL HDI 5000). In our experiment, we have shown the difference
between conventional US and SCI for the proposed speckle features. The result demonstrated that the performances achieved by these two techniques were very close. The B-mode images used in the experiment were acquired from the manipulation of the experienced radiologist (15 years). For each patient, the adaptive configuration of the US scanner was customized for generating each US image. Similarly, the proposed CAD system can be adjusted for individual US systems. By training the up-to-date data, the classifier generates the customized parameters for optimizing performance. So far, the available image data collected from the clinical examinations was obtained from the US scanner. Applying the proposed CAD system to different US platforms will be investigated in the future.Various pieces of US equipment from different
377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395
manufacturers may generate various gray-level distributions. For calibration, the difference in dynamic range or contrast can be diminished by preprocessing filters. Applying the proposed CAD system to different US platforms will be investigated in the future. At present, the proposed CAD system can be adjusted for individual US systems. By training the up-to-date data, the classifier can generate the customized parameters for optimizing performance. Another method for showing the advantage of the proposed CAD system would be to collect various nonmass types of lesions as the
specimens for diagnosis. Nonmass lesions are abnormal tissues without distinct
boundaries. Because of the indistinct boundary between the lesion and its background
tissues, it is not practical to utilize segmentation for extracting features. Lesions that
are close to a nipple or that have posterior shadowing are the specimens that we will
use to evaluate the proposed CAD system.
Acknowledgments
The authors would like to thank the National Science Council of the Republic of China and National Taiwan University for financially supporting this research under Contract No. NSC 98-2221-E-002 -172 -MY3 and 10R80919-6. This study was also supported by a grant from the Innovative Research Institute for Cell Therapy, Republic of Korea (A062260).
396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415
References
Alpaydin E. Introduction to machine learning. Cambridge, Mass: MIT Press, 2004. American College of Radiology. Breast Imaging Reporting and Data System, 4th ed.
American College of Radiology, 2003.
Cha JH, Moon WK, Cho N, Chung SY, Park SH, Park JM, Han BK, Choe YH, Cho
G, Im JG. Differentiation of benign from malignant solid breast masses:
Conventional US versus spatial compound imaging. Radiology 2005;237:841-6. Cha JH, Moon WK, Cho N, Kim SM, Park SH, Han BK, Choe YH, Park JM, Im JG.
Characterization of benign and malignant solid breast masses: Comparison of
conventional US and tissue harmonic imaging. Radiology 2007;242:63-9.
Chang CC, Tsui PH, Yeh CK, Liao YY, Kuo WH, Chang KJ, Chen CN. Ultrasonic
Nakagami Imaging: A Strategy to Visualize the Scatterer Properties of Benign
and Malignant Breast Tumors. Ultrasound Med Biol 2010;36:209-17.
Chang RF, Moon WK, Shen YW, Huang CS, Chiang LR. Computer-Aided Diagnosis
for the Classification of Breast Masses in Automated Whole Breast Ultrasound
Images. Ultrasound Med Biol 2011;37:539-48.
Field AP. Discovering statistics using SPSS, 3rd ed. Los Angeles: SAGE
Publications, 2009.
Haralick RM, Shanmuga.K, Dinstein I. Textural Features for Image Classification. 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435
IEEE Trans Syst Man Cybern 1973;Smc3:610-21.
Hosmer DW. Applied logistic regression. 2nd edition. New York: Wiley, 2000. Huang CS, Moon WM, W. K., Chang SC, Chang RF. Breast Tumor Classification
Using Fuzzy Clustering for Breast Elastography. Ultrasound Med Biol
2011;37:700-8.
Nie K, Chen JH, Yu HJ, Chu Y, Nalcioglu O, Su MY. Quantitative Analysis of
Lesion Morphology and Texture Features for Diagnostic Prediction in Breast
MRI. Acad Radiol 2008;15:1513-25.
Rangayyan RM, Mudigonda NR, Desautels JEL. Boundary modelling and shape
analysis methods for classification of mammographic masses. Med Biol Eng
Comput 2000;38:487-96.
Shankar PM, Dumane VA, George T, Piccoli CW, Reid JM, Forsberg F, Goldberg
BB. Classification of breast masses in ultrasonic B scans using Nakagami and K
distributions. Phys Med Biol 2003;48:2229-40.
Shen WC, Chang RF, Moon WK. Computer aided classification system for breast
ultrasound based on breast imaging reporting and data system (BI-RADS).
Ultrasound Med Biol 2007a;33:1688-98.
Shen WC, Chang RF, Moon WK, Chou YH, Huang CS. Breast ultrasound
computer-aided diagnosis using BI-RADS features. Acad Radiol 2007b;14:928-39.
Smith WL, Fenster A. Optimum scan spacing for three-dimensional ultrasound by 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455
speckle statistics. Ultrasound Med Biol 2000;26:551-62.
Stavros AT, Thickman D, Rapp CL, Dennis MA, Parker SH, Sisney GA. Solid breast
nodules: Use of sonography to distinguish between benign and malignant lesions.
Radiology 1995;196:123-34.
Tuthill TA, Krucker JF, Fowlkes JB, Carson PL. Automated three-dimensional US
frame positioning computed from elevational speckle decorrelation. Radiology
1998;209:575-82.
Tuthill TA, Sperry RH, Parker KJ. Deviations from Rayleigh Statistics in Ultrasonic
Speckle. Ultrason Imaging 1988;10:81-9. 456 457 458 459 460 461 462 463 464 465
Figure Captions
Fig. 1 Illustration of the ROI selection and speckle extraction. (a) A tumor is selected
by drawing an ROI from the B-mode image. (b) The ROI of a benign tumor.
(c) The extracted speckle pixels inside the ROI of (b) are shown by a white
appearance. (d) The ROI of a malignant tumor. (e) The extracted speckle
pixels inside the ROI of (d) are indicated by a white appearance.
Fig. 2 The comparison between conventional US and SCI for speckle pixels. (a) The
ROI of a B-mode image with conventional US. (b) The extracted speckle
pixels inside the ROI of (a) are shown by a white appearance. (c) The ROI of a
B-mode image with SCI. (d) The extracted speckle pixels inside the ROI of (c)
are indicated by a white appearance.
Fig. 3 The ROC curves of feature sets. (a) The ROC curves of the speckle features
with conventional US and with SCI. (b) The ROC curves of the speckle
features with SCI, segmentation features, and the combined feature set. Note
that the combined feature set includes the speckle features with SCI and the
segmentation features.
Fig. 4 A malignant tumor that was classified correctly by the speckle features but was
misclassified by the segmentation features. (a) The ROI of the tumor. (b) The 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485
S_avgnum of the tumor is relatively small (0.13). (c) The segmentation result
of the tumor.
Fig. 5 A benign tumor that was classified correctly by the speckle features but was
misclassified by the segmentation features. (a) The ROI of the tumor. (b) The
S_avgnum of the tumor is relatively large (0.17). (c) The segmentation result
of the tumor. 486 487 488 489 490 491 492
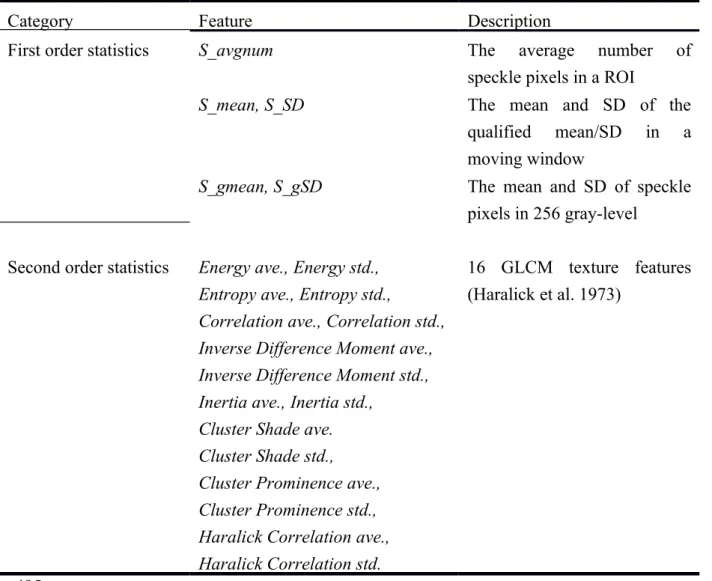
Table 1 21 speckle features.
Category Feature Description
First order statistics S_avgnum The average number of speckle pixels in a ROI
S_mean, S_SD The mean and SD of the qualified mean/SD in a moving window
S_gmean, S_gSD The mean and SD of speckle pixels in 256 gray-level
Second order statistics Energy ave., Energy std., Entropy ave., Entropy std.,
Correlation ave., Correlation std., Inverse Difference Moment ave., Inverse Difference Moment std., Inertia ave., Inertia std.,
Cluster Shade ave. Cluster Shade std., Cluster Prominence ave., Cluster Prominence std., Haralick Correlation ave., Haralick Correlation std.
16 GLCM texture features (Haralick et al. 1973)
Table 2 38 segmentation features.
Category Feature Description
Morphology Tumor_a, Tumor_p Tumor area and perimeter
Ellipse_a, The length of the major axis of the best-fit ellipse
Ellipse_b, The length of the minor axis
Ellipse_a/b, Ellipse_a / Ellipse_b
Ep/Tp, The ratio of the ellipse perimeter and the tumor perimeter
Ellipse_compactness, The overlap between the ellipse and the tumor 493
494
495
Ellipse_theta The angle of the major axis of the ellipse (Shen et al. 2007a)
NRL entropy, NRL variance NRL features(Nie et al. 2008)
Compactness Tumor roundness(Rangayyan et al. 2000)
Undulation, Sharp, MU Features about undulations on the tumor boundary(Shen et al. 2007b)
NS The number of spicules on the tumor boundary
MNS NS×Compactness
MaxSpicule The length of the longest spicule of NS
Texture LB The average intensity difference between the inner and outer bands around the tumor boundary (Shen et al. 2007b)
PS The average intensity difference between the tumor and the region under the tumor (Shen et al. 2007b)
PS_diff The average intensity difference between the surrounding tissues and the region under the tumor
EPc The average intensity difference between the 25% brighter pixels and whole tumor pixels (Shen et al. 2007b)
EP_diff The average intensity difference between the tumor and the surrounding tissues
Energy ave., Energy std., Entropy ave., Entropy std.,
Correlation ave., Correlation std., Inverse Difference Moment ave., Inverse Difference Moment std., Inertia ave., Inertia std.,
Cluster Shade ave. Cluster Shade std., Cluster Prominence ave., Cluster Prominence std., Haralick Correlation ave., Haralick Correlation std.
16 GLCM texture features (Haralick et al. 1973)
Table 3 The comparison of different window sizes for detecting speckle pixels. 3 × 3 5 × 5 7 × 7 ≥ 9 × 9 5 × 5 VS. 3 × 3 (p-value) 5 × 5 VS. 7 × 7 (p-value) Accuracy 83.9% (115/137) (122/137)89.1% (116/137)84.7% N/A 0.2160 0.2833 Sensitivity 73.8% (31/42) 81.0% (34/42) 76.2% (32/42) N/A 0.4340 0.5949 Specificity 88.4% (84/95) 92.6% (88/95) 88.4%. (84/95) N/A 0.3217 0.3217 PPV 73.8% (31/42) 82.9% (34/41) 74.4% (32/43) N/A 0.3136 0.3421 NPV 88.4% (84/95) 91.7% (88/96) 89.4% (84/94) N/A 0.4537 0.5875 Az 0.89 0.93 0.89 N/A 0.0657 0.0762
N/A: not available.
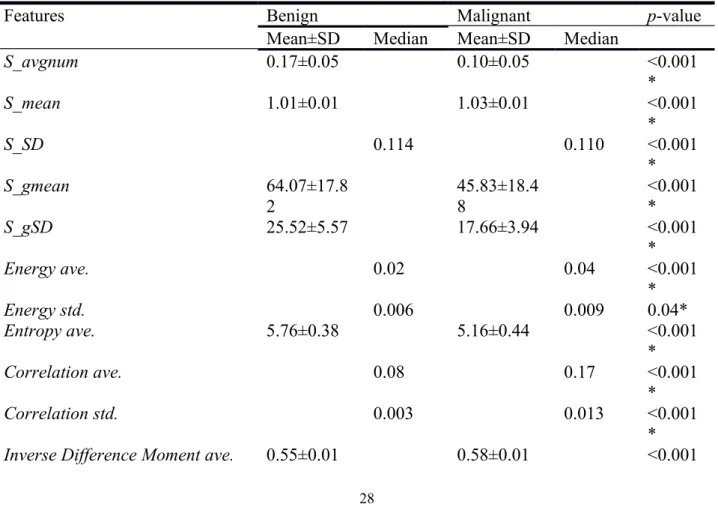
Table 4 The mean, standard deviation (SD), median, and p-value (Student’s t-test or
Mann-Whitney U-test) of significant speckle features with SCI.
Features Benign Malignant p-value
Mean±SD Median Mean±SD Median
S_avgnum 0.17±0.05 0.10±0.05 <0.001 * S_mean 1.01±0.01 1.03±0.01 <0.001 * S_SD 0.114 0.110 <0.001 * S_gmean 64.07±17.8 2 45.83±18.48 <0.001* S_gSD 25.52±5.57 17.66±3.94 <0.001 * Energy ave. 0.02 0.04 <0.001 * Energy std. 0.006 0.009 0.04* Entropy ave. 5.76±0.38 5.16±0.44 <0.001 * Correlation ave. 0.08 0.17 <0.001 * Correlation std. 0.003 0.013 <0.001 *
Inverse Difference Moment ave. 0.55±0.01 0.58±0.01 <0.001 498
499 500 501
*
Inertia ave. 1.82±0.19 1.57±0.17 <0.001 *
Inertia std. 0.69±0.1 0.57±0.1 <0.001 *
Cluster Shade ave. 121.95 40.23 0.002*
Cluster Shade std. 13.14 4.02 <0.001
*
Cluster Prominence ave. 4399.12 909.71 <0.001 *
Cluster Prominence std. 333.57 79.64 <0.001 *
Haralick Correlation ave. 33598.5 5
9968.95 <0.001 *
Haralick Correlation std. 1164.68 418.58 <0.001 * * p-value<0.05 indicates a statistically significant difference.
Table 5 The mean, standard deviation (SD), median, and p-value (Student’s t-test or
Mann-Whitney U-test) of significant segmentation features.
Features Benign Malignant p-value
Mean±SD Median Mean±SD Median
Tumor_a 4676 12839 <0.001* Tumor_p 344 616 <0.001* Ellipse_a 57.05±22.22 70.92±23.73 <0.001* Ellipse_b 30.92 54.77 <0.001* Ellipse_a/b 1.63 1.34 <0.001* Ep/Tp 0.79±0.09 0.67±0.10 <0.001* Ellipse_theta 0.08 0.20 0.002* NRL entropy 2.56±0.37 2.31±0.42 <0.001* NRL variance 0.14±0.03 0.11±0.03 <0.001* Undulation 2 4 <0.001* Sharp 2 3.5 <0.001* MU 5 7.5 <0.001* MNS 1.69±0.73 2.15±1.06 0.01* LB 34.91±8.95 26.15±8.97 <0.001* EPc 0.41 0.64 0.004* EP_diff 36.05±9.27 28.88±9.09 <0.001* Energy ave. 0.08 0.09 0.02* Entropy std. 0.20±0.03 0.18±0.03 <0.001* Correlation std. 0.02 0.01 0.04*
Inverse Difference Moment ave. 0.72±0.04 0.75±0.05 <0.001*
Inverse Difference Moment std. 0.05±0.01 0.04±0.01 <0.001* 503
504 505
Inertia ave. 0.80±0.24 0.67±0.23 0.003*
Inertia std. 0.23±0.08 0.18±0.08 <0.001*
Cluster Shade ave. 9.60 17.18 0.005*
Haralick Correlation ave. 3529.22 2166.20 0.01*
Haralick Correlation std. 54.12 27.39 <0.001* * p-value<0.05 indicates a statistically significant difference.
Table 6 The comparison of five performance indices and p-values (chi-square test)
between the speckle features and the segmentation features. Speckle (with SCI) Speckle (with convention al US) Segmentat ion Combined Speckle (with SCI) VS. Speckle (with convention al US) (p-value) Speckle (with SCI) VS. Segmentat ion (p-value) Combined VS. Speckle (with SCI) (p-value) Combined VS. Segmentat ion (p-value) Accuracy 89.1% (122/137) 88.3% (121/137) 84.7% (116/137) 89.8% (123/137) 0.8487 0.2833 0.8443 0.2052 Sensitivity 81.0% (34/42) 78.6% (33/42) 73.8% (31/42) 81.0% (34/42) 0.7860 0.4340 1.0000 0.4340 Specificity 92.6% (88/95) (88/95)92.6% (85/95)89.5%. (89/95)93.7% 1.0000 0.4458 0.7738 0.2960 PPV 82.9% (34/41) 82.5% (33/40) 75.6% (31/41) 85.0% (34/40) 0.9595 0.4138 0.7994 0.2886 NPV 91.7% (88/96) 90.7% (88/97) 88.5% (85/96) 91.8% (89/97) 0.8168 0.4684 0.9827 0.4541 Az 0.93 0.89 0.86 0.94 0.2513 0.0359* 0.9573 0.0219*
* p-value<0.05 indicates a statistically significant difference.
Combined feature set includes the speckle features with SCI and the segmentation
features. 507 508 509 510 511 512 513