Association of Rheumatoid Arthritis Risk with EGFR Genetic
Polymorphisms in Taiwan’s Han Chinese population.
Sui-Foon Lo,1,6 Lei Wan,2,7 Hsiu-Chen Lin,3 Chung-Ming Huang,4,8 Shih-Yin Chen,2,7
Su-Ching Liu2 and Fuu-Jen Tsai,2,5,7,9
1Department of Physical Medicine and Rehabilitation, 2Department of Medical
Genetics; 4Division of Immunology and Rheumatology, Department of Internal
Medicine; 5Department of Pediatrics; China Medical University Hospital.
3Department of Physical therapy; 6Department of Chinese Medicine, 7Graduate
Institute of Chinese Medical Science, 8 Graduate Institute of Integrated Medicine;
China Medical University, Taichung, Taiwan. 9Department of Biotechnology, Asia
University, Taichung, Taiwan.
Address correspondence to: Fuu-Jen Tsai,
Department of Medical Genetics, China Medical University Hospital, No 2, Yuh Der Road, Taichung, Taiwan
TEL: 886-4-22052121 ext 2041 Fax: 886-4-22033295
ABSTRACT
The involvement of the epidermal growth factor receptor (EGFR) in the pathogenesis of cancer is well documented. In contrast, its role in rheumatoid arthritis (RA) development is not that well defined although previous studies suggested the possible link between autoimmune diseases and malignancy. Therefore, we aimed to examine if there is a link between the EGFR genetic polymorphisms and RA.
Our study gauged the effects of EGFR (rs11543848 and rs17337023) single nucleotide polymorphisms (SNPs) on RA among Taiwan’s Han Chinese population. Polymorphism of EGFR gene was analyzed in 188 RA patients and 128 control subjects. Genotyping for EGFR SNPs was performed by restriction fragment length polymorphism (RFLP) assay. Our data confirmed statistically significant increased risk of RA development in subjects with A carrier at rs17337023 SNP (p=2.232E-07), and subjects with A allele at rs17337023 SNP (odds ratio [OR] = 1.52; 95% confidence interval [CI] = 1.10-2.09). Furthermore, comparison of haplotype frequencies between patients and controls suggested GA and AT haplotypes were more “at-risk” for RA development (p=3.333E-05 and p=1.3E-03, respectively). However, comparisons of the clinical features of RA patients according to different genotypes and haplotypes revealed no significant difference.
(rs11543848 and rs17337023) with the susceptibility of RA development and polymorphism revealed by this study merit further investigation.
Keywords: Rheumatoid Arthritis (RA); Epidermal growth factor receptor (EGFR); Single nucleotide polymorphisms (SNPs); Haplotype.
Running Title:
EGFR polymorphisms and Rheumatoid Arthritis
Introduction:
Rheumatoid arthritis (RA) is a chronic autoimmune inflammatory disease characterized by immune activation and hyperplasia of the synovium leading to progressive destruction of the affected joints. Activated synovial fibroblasts recruit inflammatory cells and cause granulomatous tissues (pannuses) formation and angiogenesis in the synovial membrane of the RA patients (1). These activated synoviocytes demonstrate the invasive behavior of metastatic malignant cells and show resistance to apoptosis (2,3). Similar to transformed cells, the rheumatoid fibroblasts have augmented tyrosine-phosphorylated proteins and manifest tumor-like behavior. Several cytokines that activate these fibroblasts mediate their action through tyrosine kinase growth factor receptors. Mechanisms of signal transduction via such tyrosine kinases are important in the development of rheumatoid lesions (4-6). Of the
receptor tyrosine kinases, the epidermal growth factor (EGF) /epidermal growth factor receptor (EGFR) families play a crucial role in the development, physiology, and cancer (7). Epidermal growth factor has been found as an important factor that stimulates the release of inflammatory mediators and enhances the growth of synovial cells (8). Among the members of the EGF family, amphiregulin and transforming growth factor–alpha (TGF-α), which are the main ligands for EGFR, have been found to have increased expression in the synovial tissues of RA patients (9,10). In addition, EGFR has been found to be the predominantly expressed receptors in synovial fibroblasts (9). The EGFR is encoded by a gene located in the short arm of chromosome 7 (7p12.1-12.3) (11). The EGFR 2133A/T polymorphism (rs17337023) was found to be associated with malignant oral keratinocytes (12), systemic lupus erythematosus (SLE) (13), endometriosis, and leiomyomas (14) in previous studies. Another polymorphism, the EGFR 1808G/A polymorphism (rs11543848) was found to be associated with acute coronary syndrome (15), the prognosis of colorectal carcinoma (16), and advanced lung cancer (17). In view of the tumor-like behavior of synovial fibroblasts and chronic inflammatory process in rheumatoid arthritis, we hypothesized that the EGFR gene polymorphism may be associated with the risk of rheumatoid arthritis development. To verify our hypothesis, we compared the genotypic and allelic frequencies of the EGFR polymorphisms (rs11543848 and
rs17337023) between RA patients and normal control participants of the Han Chinese population living in Taiwan. Furthermore, we compared the genotypes among RA patients with various clinical variables to find out if there is a relationship between EGFR polymorphisms and the clinical manifestation of RA.
Materials and Methods:
Patient selection
The study subjects included 188 RA patients and 128 healthy subjects, recruited from China Medical University Hospital in Taiwan. Patients with RA according to the revised America College of Rheumatology criteria(18) were enrolled. Nephelometry detected rheumatoid factor (RF), values ≧ 30 IU/ml defined as positive. Presence or history of extra-articular manifestations in patients with RA was recorded (19). Radiographs of hands, wrists, and feet of patients were taken, and the presence or absence of joint erosion was evaluated by a rheumatologist and a radiologist. The gender-age-matched unrelated healthy controls from the general population were selected by health examination. All individuals’ samples were collected by venipuncture for genomic DNA isolation. Informed consent was from all participants and was approved by the local Ethics Committee.
Polymerase chain reaction (PCR) was used to identify the EGFR polymorphisms including rs11543848 and rs17337023. Polymerase chain reaction was carried out in a total volume of 50 μL, containing genomic DNA 50ng, 2–6 pmol of each primer, 1X Taq polymerase buffer (1.5mM MgCl2) and 0.5 units of AmpliTaq DNA polymerase (Perkin
Elmer; Foster City,CA,USA). In the study of the EGFR rs17337023 SNP, the primers used were upstream 5'-ATATATGCCAAAGAAGTAG-3' and downstream 5'-TGATCAGGACAGAGGACAG-3'. For the EGFR rs11543848 SNP, the primers used were upstream 5'-TGCTGTGACCCACTCTGTCT-3' and downstream 5’-CCAGAAGGTTGCACTTGTCC-3'. Polymerase chain reaction amplification was performed in a programmable PCR thermal cycler (GeneAmp PCR System 2400, Perkin Elmer). The PCR cycling conditions for EGFR rs17337023 SNP examination were as follows: one cycle at 95°C for 5 min ,35 cycles at 95 °C for 30 seconds, 56 °C for 30 seconds, 72 °C for 45 seconds. One final cycle of extension at 72°C for 7 min, then holding at 25°C. The EGFR rs17337023 SNP was analyzed by PCR amplification followed by restriction enzyme analysis with BsrI. Two fragments of 136 bp and 64 bp were present if the product was excised (TT homozygote). The uncut band showed up as a 200 bp length on the gel. The reaction was then incubated for overnight at 65°C, and then 10μl of the products were loaded into a 3% agarose gel containing ethidium bromide for electrophoresis. The EGFR rs17337023 SNP was categorized as excisable (TT
homozygote), non-excisable (AA homozygote), and (AT heterozygote). The PCR cycling conditions for EGFR rs11543848 SNP examination were as follows: one cycle at 95°C for 5 min ,35 cycles at 95 °C for 30 seconds,59 °C for 30 seconds, 72 °C for 45 seconds. One final cycle of extension at 72°C for 7 min, then holding at 25°C. The EGFR rs11543848 SNP was analyzed by PCR amplification followed by restriction enzyme analysis with BstNI. Three fragments of 67 bp ,50bp and 38 bp were present if the product was excised(GG homozygote). The uncut band showed up as two fragments of 117bp and 38 bp length on the gel. The reaction was then incubated for overnight at 60°C, and then 10μl of the products were loaded into a 3% agarose gel containing ethidium bromide for electrophoresis. The EGFR rs11543848 SNP was categorized as excisable (GG homozygote), non-excisable (AA homozygote), and (AG heterozygote). Statistical analysis
The genotypic and allelic frequencies of EGFR SNPs (rs17337023 and rs11543848) for the RA patients and controls were compared using the chi-square test. Among the RA patients, genotype groups with different clinical variables were also compared using chi-square test. When one cell had an expected count of <1 or >20% of the cells had an expected count of <5, Fisher’s exact test was used. Results were considered statistically significant when p values less than 0.05.The odds ratios (OR) were calculated from the genotypic frequency and allelic frequency with a 95%
confidence interval (95% CI) for the EGFR SNPs (rs17337023 and rs11543848). The statistical analysis was performed by using the SPSS version 11.
Results:
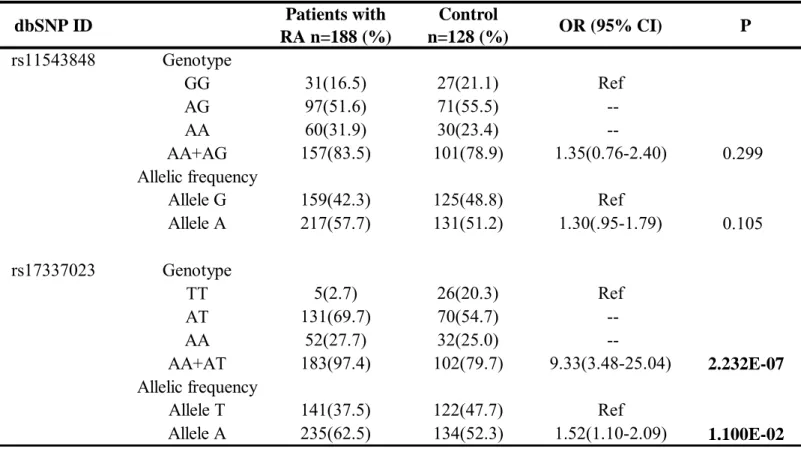
Table 1 depicts genotypic and allelic frequencies of rs11543848 and rs17337023. Genotype distributions were in Hardy-Weinberg equilibrium. We observed A allele as the major one at rs11543848 polymorphism, both in RA patients (57.7%; 217/376) and controls (51.2%; 131/256), A allele as the major allele at rs17337023 polymorphism in patients (62.5%; 235/376) and controls (52.3%; 134/256) was also found. Comparison of allelic and genotypic distributions between RA patients and controls yielded significant differences for the rs17337023 SNP (p=1.1E-02 and p=2.232E-07, respectively). Data indicated that individuals with A allele or A carrier (AA + AT) genotype at rs17337023 SNP are at higher risk for RA.
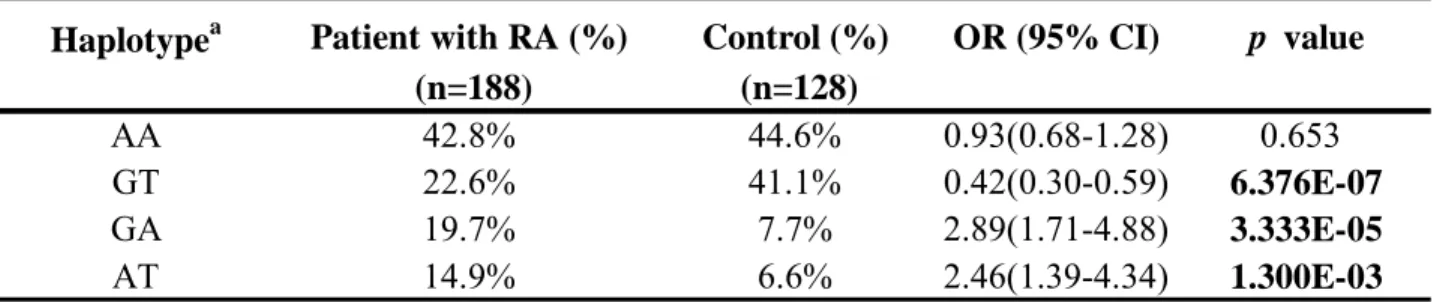
Haplotype frequencies were estimated for rs11543848 and rs17337023 SNPs. Four haplotypes EGFR emerged in the study population: AA is the most common both in RA patients (42.8%) and healthy controls (44.6%). When we compared the overall distribution of haplotype frequencies between RA patients and health controls, significant differences were observed as shown Table 2. Our data showed that the GT haplotype appeared to be a significant “protective” haplotype compared with other
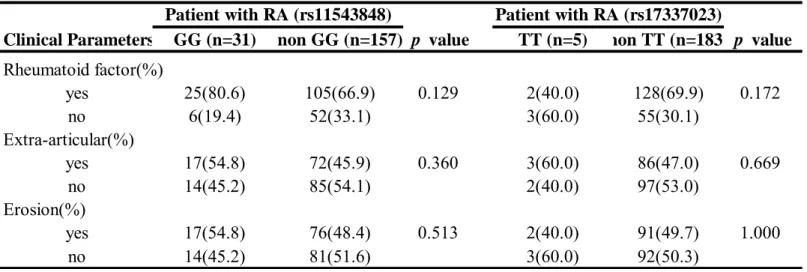
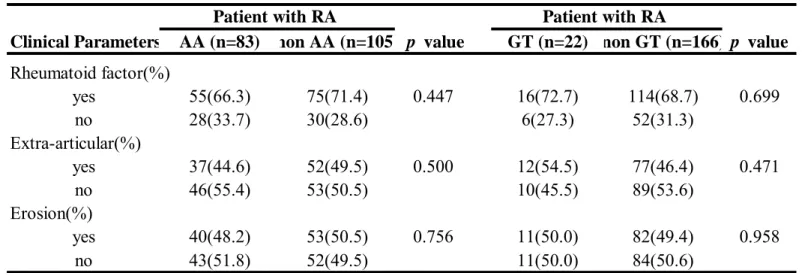
haplotypes (OR: 0.42, 95% CI: 0.3-0.59; p=6.376E-07) in RA development. On the other hand the GA or AT haplotype appeared to be the significant “at-risk” haplotype for RA development (p = 3.333E-05 and 1.3E-03, respectively). In addition, comparisons of the clinical features of RA patients according to different genotypes and haplotypes were performed and revealed no significant difference ( Table 3 and 4).
Discussion:
Epidermal growth factor receptor (EGFR) is a trans-membrane receptor with a molecular weight of approximately 170 kDa encoded by the c-erbB1 proto-oncogene (20-22). It belongs to a family of four closely related receptor tyrosine kinases which has been recognized as a convergence and switch point for diverse signal transduction pathways involved in cellular communication (23,24). The biological responses to the EGFR signal transductions are pleiotrophic including mitogenesis, protein secretion, enhanced cell motility, proliferation, invasion, cell adhesion, and angiogenesis (21,25). Many studies have shown enough evidence supporting the role of EGFR activation and signaling in the pathogenesis of malignancies. Meanwhile, numerous reports have suggested a link between rheumatic diseases, autoimmune phenomena, and cancer (26). Increased risk of certain cancers, particularly hematologic cancers, has been found in
RA and SLE patients compared with the general population, (26-28). Autoantibodies to EGFR in systemic lupus erythematosis, systemic sclerosis, and autoimmune mice also have been found (29), further suggesting the possible link between autoimmune disease and tumor immunology in relation to EGFR. Therefore, it is possible that EGFR gene polymorphism is associated with the risk of RA development. Our results in the study of the EGFR rs17337023 SNP support this hypothesis (Table1). When compared with the controls, RA patients were found to have significantly higher frequencies of the A carrier (AA + AT). These result suggests that individuals with A carrier genotype at rs17337023 SNP are at higher risk for RA. On the other hand, our analyses of allelic, genotypic and halplotype frequencies suggest that the T allele has a protective effect at rs17337023. Although the study of rs11543848 SNP showed no such risk association (Table 1), the risk association of the rs17337023 SNP in autoimmune diseases was further supported by previous study of our group in identifying the association of rs17337023 SNP with SLE (13). However, we found that the EGFR polymorphisms have no association with clinical features of RA, which is similar to our previous findings in SLE (13). Although the exact pathogenic mechanism relating these polymorphisms to these chronic inflammatory diseases still remain unknown, the extremely complex signaling network should be considered in the disease progression. We observed the rs17337023 SNP could be in linkage disequilibrium with other
polymorphisms, something like rs11543848 SNP, leading to disease development in RA and SLE patients. In addition, we also observed a residue change ( R [Arg] to K [Lys]) in the protein level at rs11543848 SNP. Therefore, further studies are required to explore if rs17337023 and rs11543848 SNPs are independent risk factor or an indirect marker of different genetic factors in RA development. Moreover, due to the limited subject number from one medical center in our study, larger scale studies for further confirmation of our findings are also needed in the future. In conclusion, our present study is the first report to identify that the EGFR rs17337023 polymorphism is associated with the risk of RA development.
References:
1. Firestein G (2005) Etiology and pathogenesis of rheumatoid arthritis. In: Harris ED, Budd RC, Firestein GS, et al (Eds) Kelley’s textbook of rheumatology, 7th edn.
Philadelphia: Elsevier/Saunders company, pp 996-1042.
2. Asahara H, Hasumuna T, Kobata T, Yagita H, Okumura K, Inoue H, Gay S, Sumida T, Nishioka K (1996) Expression of Fas antigen and Fas ligand in the rheumatoid synovial tissue. Clin Immuno Immunopathol 81: 27-34
3. Baier A, Meineckel I, Gay S, Pap T (2003) Apoptosis in rheumatoid arthritis. Curr Opin Rheumatol 15: 274-279
4. Williams WV, VonFeldt JM, Ramanujam T, Weiner DB (1992) Tyrosine kinase signal transduction in rheumatoid synovitis. Semin Arthritis Rheum 21:317-329 5. Migita K, Eguchi K, Tsukada T, Kawabe Y, Aoyagi T, Nagataki S (1995) The role
of protein kinase in human synovial fibroblast growth. Biochem Biophys Res Commun 210:1066–1075.
6. Takeuchi T, Abe T (1998) Tyrosine phosphorylated proteins in synovial cells of rheumatoid arthritis. Intl Rev Immunol 17:365–381.
7. Wieduwilt MJ, Moasser MM (2008) The epidermal growth factor receptor family: Biology driving targeted therapeutics. Cell Mol Life Sci 65:1566-1584.
8.Goddard DH, Grossman SL, Newton R (1990) Polypeptide growth factors augment interleukin 1-induced release of prostaglandin E2 by rheumatoid arthritis synovial
cells in vitro. Cytokine 2: 294–299.
9. Yamane S, Ishida S, Hanamoto Y, et al (2008) Proinflammatory role of amphiregulin, an epidermal growth factor family member whose expression is augmented in rheumatoid arthritis patients. J Inflamm (Lond) 27:5:5.
10. Hallbeck AL, Walz TM, Briheim K, Wasteson A (2005) TGF-α and ErB2 production in synovial joint tissue: increased expression in arthritic joints. Scand J Rheumatol 34: 204-211
of the 110 kb locus and identification of sequences regulating its transcription. Oncogene Res 1:375-396.
12. Shintani S, Matsuo K, Crohin CC, et al (1999) Intragenic mutation analysis of the human epidermal growth factor receptor (EGFR) gene in malignant human oral keratinocytes. Cancer Res 59: 4142–4147
13.Huang CM, Tsai CH, Chen CL, Chang CP, Lai CC, Tsai FJ (2004) Epidermal growth factor receptor (EGFR) gene Bsr I polymorphism is associated with systemic lupus erythematosus. Lupus 13:773-776
14.Hsieh YY, Chang CC, Tsai FJ, Lin CC, Tsai CH (2005) T homozygote and allele of epidermal growth factor receptor 2073 gene polymorphism are associated with
higher susceptibility to endometriosis and leiomyomas. Fertil Steril 83:796-799 15.Gao LB, Zhou B, Zhang L et al (2008) R497K polymorphism in epidermal growth
factor receptor gene is associated with the risk of acute coronary syndrome BMC Med Genet 9:74
16. Wang WS, Chen PM, Chiou TJ, et al (2007) Epidermal growth factor receptor R497K polymorphism is a favorable prognostic factor for patients with colorectal carcinoma. Clin Cancer Res 13:3597-3604
favorable prognostic factor for advanced lung cancer. J Cancer Res Clin Oncol 135:313-318
18.Arnett FC, Edworthy SM, Bloch DA, et al (1988) The American Rheumatism Association 1987 revised criteria for the classification of rheumatoid arthritis. Arthritis Rheum 31:315-324
19.Yen JH, Chen JR, Tsai WJ, Tsai JJ, Liu HW (1995) HLA-DRB1 genotyping in patients with rheumatoid arthritis in Taiwan. J Rheumatol 22:1450-1454
20. Ullrich A, Coussens L, Hayflick JS, et al (1984) Human pidermal growth factor receptor cDNA sequence and aberrant expression of the amplified gene in A431 epidermoid carcinoma cells. Nature 309:418-425.
21.Wells A (1999) EGF receptor. Int J Biochem Cell Biol 31:637-643
22.Yarden Y (2001) The EGFR family and its ligands in human cancer signalling mechanisms and therapeutic opportunities. Eur J Cancer 37 Suppl 4:S3-8 23. Hackel PO, Zwick E, Prenzel N, Ullrich A (1999) Epidermal growth factor
receptors: critical mediators of multiple receptor pathways. Curr Opin Cell Biol 11:184-189
24. Moghal N, Sternberg PW (1999) Multiple positive and negative regulators of signaling by the EGF-receptor. Curr Opin Cell Biol 11:190-196
ErbB/epidermal growth factor (EGF) receptor family and its ligands in ovarian cancer. Biochim Biophys Acta 1785:232-265
26.Chakravarty EF, Genovese MC (2005) Musculoskeletal syndromes in malignancy. In: Harris ED, Budd RC, Firestein GS, et al (Eds) Kelley's textbook of
rheumatology, 7th edn. Philadelphia: Elsevier/Saunders company, pp1754-1772 27. Smitten AL, Simon TA, Hochberg MC, Suissa S (2008) A meta-analysis of the
incidence of malignancy in adult patients with rheumatoid arthritis. Arthritis Res Ther 10:R45.
28. Bernatsky S, Boivin JF, Joseph L, et al (2005) An international cohort study of cancer in systemic lupus erythematosus. Arthritis Rheum 52:1481-1490
29.Planque S, Zhou YX, Nishiyama Y, et al (2003) Auto antibodies to the epidermal growth factor receptor in systemic sclerosis,lupus, and autoimmune mice. FASEB J 17:136-143
Table 1. Genotypic and allelic frequencies of EGFR genetic polymorphisms in the patients with RA and controls. dbSNP ID Patients with RA n=188 (%) Control n=128 (%) OR (95% CI) P rs11543848 Genotype GG 31(16.5) 27(21.1) Ref AG 97(51.6) 71(55.5) --AA 60(31.9) 30(23.4) --AA+AG 157(83.5) 101(78.9) 1.35(0.76-2.40) 0.299 Allelic frequency Allele G 159(42.3) 125(48.8) Ref Allele A 217(57.7) 131(51.2) 1.30(.95-1.79) 0.105 rs17337023 Genotype TT 5(2.7) 26(20.3) Ref AT 131(69.7) 70(54.7) --AA 52(27.7) 32(25.0) --AA+AT 183(97.4) 102(79.7) 9.33(3.48-25.04) 2.232E-07 Allelic frequency Allele T 141(37.5) 122(47.7) Ref Allele A 235(62.5) 134(52.3) 1.52(1.10-2.09) 1.100E-02
Table 2. Distribution of EGFR haplotype frequencies in the patients with RA and controls. Haplotypea Patient with RA (%) Control (%) OR (95% CI) p value
(n=188) (n=128)
AA 42.8% 44.6% 0.93(0.68-1.28) 0.653
GT 22.6% 41.1% 0.42(0.30-0.59) 6.376E-07
GA 19.7% 7.7% 2.89(1.71-4.88) 3.333E-05
AT 14.9% 6.6% 2.46(1.39-4.34) 1.300E-03
CI, confidence interval; OR, odds ratio.
Table 3. Comparison of clinical and biochemical manifestations of RA patients with EGFR genetype distribution.
Clinical Parameters GG (n=31) non GG (n=157) p value TT (n=5) non TT (n=183 p value Rheumatoid factor(%) yes 25(80.6) 105(66.9) 0.129 2(40.0) 128(69.9) 0.172 no 6(19.4) 52(33.1) 3(60.0) 55(30.1) Extra-articular(%) yes 17(54.8) 72(45.9) 0.360 3(60.0) 86(47.0) 0.669 no 14(45.2) 85(54.1) 2(40.0) 97(53.0) Erosion(%) yes 17(54.8) 76(48.4) 0.513 2(40.0) 91(49.7) 1.000 no 14(45.2) 81(51.6) 3(60.0) 92(50.3)
Table 4. Comparison of clinical and biochemical manifestations of RA patients with EGFR haplotype distribution.
Clinical Parameters AA (n=83) non AA (n=105) p value GT (n=22) non GT (n=166) p value Rheumatoid factor(%) yes 55(66.3) 75(71.4) 0.447 16(72.7) 114(68.7) 0.699 no 28(33.7) 30(28.6) 6(27.3) 52(31.3) Extra-articular(%) yes 37(44.6) 52(49.5) 0.500 12(54.5) 77(46.4) 0.471 no 46(55.4) 53(50.5) 10(45.5) 89(53.6) Erosion(%) yes 40(48.2) 53(50.5) 0.756 11(50.0) 82(49.4) 0.958 no 43(51.8) 52(49.5) 11(50.0) 84(50.6)