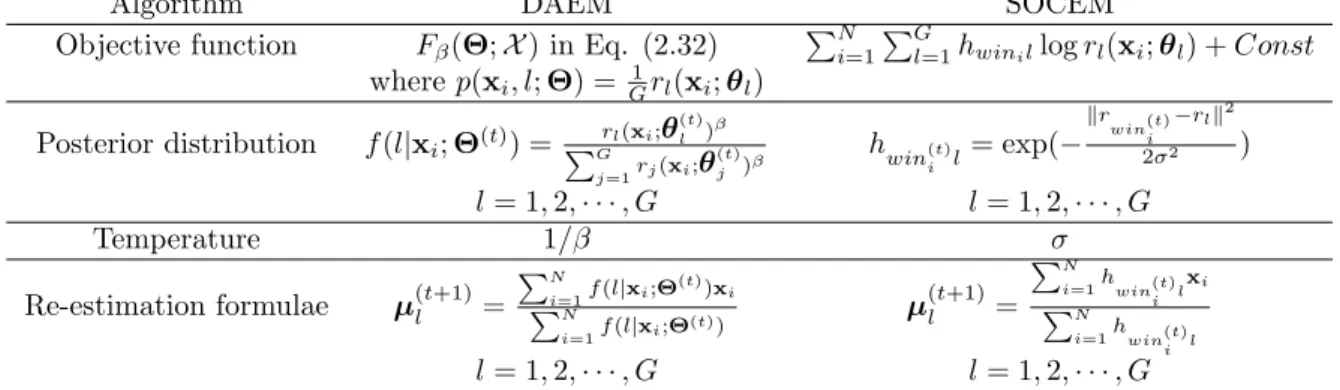國
立
交
通
大
學
資訊科學與工程研究所
博
士
論
文
機率式模型分群法之研究與其應用
Probabilistic Model-based Clustering and Its Applications
研 究 生:鄭士賢
指導教授:傅心家 教授
王新民 博士
機率式模型分群法之研究與其應用
Probabilistic Model-based Clustering and Its Applications
研 究 生:鄭士賢 Student:Shih-Sian Cheng
指導教授:傅心家 教授 Advisor:Prof. Hsin-Chia Fu
王新民 博士
Dr. Hsin-Min Wang
國 立 交 通 大 學
資 訊 科 學 與 工 程 研 究 所
博 士 論 文
A DissertationSubmitted to Department of Computer Science College of Computer Science
National Chiao Tung University in partial Fulfillment of the Requirements
for the Degree of Doctor of Philosophy
in
Computer Science
May 2009
Hsinchu, Taiwan, Republic of China
機率式模型分群法之研究與其應用
學生:鄭士賢
指導教授
:傅心家 教授
王新民 博士
國立交通大學資訊科學與工程研究所
摘
要
機率式模型分群法藉由學習一個有限混合模型(finite mixture model)而達成資料分群的 目的,其已經有很多成功的應用; 而最常用的混合模型乃是高斯混合模型(Gaussian mixture model, i.e., GMM)。當其目的函數的最佳化無法用分析方法來達成時,我們經 常用迭代式、局部最佳(local-optimal)的演算法來學習混合模型。此外,因為來自於混 合模型的資料樣本有非完全(incompleteness)的特性,因此我們通常採用 EM 形式 (EM-type)的演算法,如 EM 演算法與分類 EM 演算法(classification EM, i.e., CEM)。然 而,用傳統 EM 形式演算法來做模型分群有幾個缺點: 一、它的效能與模型初始值有 高度相關性; 二、混合元件(mixture component)的個數需要事先給定; 三、在完成分群 之後,資料樣本與群聚之間的拓樸關係(topological relationships)無法被保留。在本論 文中,針對上述前兩樣缺點,我們提出一個自我分裂之高斯混合模型學習演算法 (self-splitting Gaussian mixture learning, i.e., SGML)。此法起始於單一元件,然後迭代 式地,用貝氏資訊法則(Bayesian information criterion, i.e., BIC)來確認分裂的有效性, 分裂某一個元件而成為兩個新元件直到最佳的元件個數被找到為止,最佳的元件個數 亦是用 BIC 來決定。基於此分裂程序,SGML 可為學習不同模型複雜度的 GMM 提供 不錯的模型初始值。關於拓樸保留(topology-preserving)的議題,我們將一個機率式自 我組織圖(probabilistic self-organizing map, i.e., PbSOM)的學習視為一種可保留資料樣 本與群聚之間拓樸關係於一個神經網路(neural network)的模型分群法。基於此概念, 我們為 PbSOM 發展了一個耦合概似混合模型(coupling-likelihood mixture model),其延 伸 Kohonen SOM 的參考向量(reference vectors)至多變量高斯模型。基於最大概似法則
(maximum likelihood criterion),我們亦發展了三個學習 PbSOM 的 EM 形式演算法,亦 即 SOCEM、 SOEM、以及 SODAEM 演算法。SODAEM 是 SOCEM 與 SOEM 的決定 性 退 火 (deterministic annealing, i.e., DA) 變 形 ; 此 外 , 藉 由 逐 漸 縮 小 鄰 域 大 小 (neighborhood size),SOCEM 與 SOEM 可分別被解釋成: 高斯模型分群的 CEM 與 EM 演算法的 DA 變形。實驗結果顯示,我們所提出的 PbSOM 演算法與決定性 EM(DAEM) 演算法有相近的分群效能,並能維持拓樸保留之特性。關於應用方面,我們用 SGML 來訓練用於語者識別(speaker identification)的語者 GMMs;實驗結果顯示,SGML 可以 自動地為個別語者 GMM 決定適當的模型複雜度,而此在文獻裡通常是用經驗法則來 決定的。我們將所提出的 PbSOM 學習演算法應用於資料視覺化與分析;實驗結果顯示 它們可用於在一個二維的網路上探索資料樣本與群聚之間的拓樸關係。此外,我們提 出幾種分割且克服(divide-and-conquer)的方法來做音訊分段(audio segmentation),其用 BIC 來評定兩個音段之間的不相似度(dissimilarity);在廣播新聞的實驗結果顯示,相較 於公認最佳的視窗成長分段法(window-growing-based segmentation),我們的方法不僅 有比較低的計算量,亦有較高的分段正確性。
Probabilistic
Model-based Clustering and Its Applications
Student:Shih-Sian Cheng
Advisors:Prof. Hsin-Chia Fu
Dr. Hsin-Min Wang
Department of Computer Science
National Chiao Tung University
ABSTRACT
Probabilistic model-based clustering is achieved by learning a finite mixture model (usually a Gaussian mixture model, i.e., GMM) and has been successfully applied to many tasks. When the objective likelihood function cannot be optimized analytically, iterative, locally-optimal learning algorithms are usually applied to learn the model. Moreover, because the data samples drawn from a mixture model have the property of incompleteness, EM-type algorithms, such as EM and classification EM (CEM), are usually employed. However, model-based clustering based on conventional EM-type algorithms suffer from the following shortcomings: 1) the performance is sensitive to the model initialization; 2) the number of mixture components (data clusters) needs to be pre-defined; and 3) the topological relationships between data samples and clusters are not preserved after the learning process. In this thesis, we propose a self-splitting Gaussian mixture learning algorithm (SGML) to address the first two issues. The algorithm starts with one single component and iteratively splits a component into two new components using Bayesian information criterion (BIC) as the validity measure for the splitting until the most appropriate component number is found, which is also achieved by BIC. Based on the splitting process, SGML provides a decent model initialization for learning Gaussian mixture models with different model complexities. For the topology-preserving issue, we consider the learning process of a probabilistic self-organizing map (PbSOM) as a model-based clustering procedure that preserves the topological relationships between data samples and clusters in a neural network. Based on this concept, we develop a
coupling-likelihood mixture model for the PbSOM that extends the reference vectors in Kohonen's SOM to multivariate Gaussians distributions. We also derive three EM-type algorithms, called the SOCEM, SOEM, and SODAEM algorithms, for learning the model (PbSOM) based on the maximum likelihood criterion. SODAEM is a deterministic annealing (DA) variant of SOCEM and SOEM; moreover, by shrinking the neighborhood size, SOCEM and SOEM can be interpreted, respectively, as DA variants of the CEM and EM algorithms for Gaussian model-based clustering. The experiment results show that the proposed PbSOM learning algorithms achieve comparable data clustering performance to that of the deterministic annealing EM (DAEM) approach, while maintaining the topology-preserving property. For applications, we apply SGML to the training of speaker GMMs for the speaker identification task; the experiment results show that SGML can automatically determine an appropriate model complexity for a speaker GMM, which is usually determined empirically in the literature. We apply the proposed PbSOM algorithms to data visualization and analysis; the experiment results show that they can be used to explore the topological relationships between data samples and clusters on a two-dimensional network. In addition, we propose divide-and-conquer approaches for the audio segmentation task using BIC to evaluate the dissimilarity between two audio segments; the experiment results on the broad-cast news data demonstrate that our approaches not only have a lower computation cost but also achieve higher segmentation accuracy than the leading window-growing-based segmentation approach.
ACKNOWLEDGEMENTS
I am grateful to my advisor, Prof. Hsin-Chia Fu and Dr. Hsin-Min Wang, for their intensive suggestions, patient guidance, and enthusiasm of research. Moreover, I would like to thank all the members of the Neural Network Multimedia Laboratory at Department of Computer Science, National Chiao Tung University, and the Spoken Language Group, Chinese Information Processing Laboratory at Institute of Information Science, Academia Sinica, for their valuable discussions. Finally, I would like to express my appreciation to my family for their supporting and encouragement. I dedicate this dissertation to my parents.
Contents
1 Introduction 1
1.1 Background . . . 1
1.2 Contributions of this dissertation . . . 5
1.2.1 On model initialization and complexity . . . 5
1.2.2 On topology-preserving ability . . . 5
1.2.3 On applications . . . 6
1.3 Organization of this dissertation . . . 6
2 Preliminaries and related works 7 2.1 K-means clustering . . . 7
2.2 Hierarchical agglomerative clustering . . . 8
2.3 Self-organizing map . . . 9
2.4 Probabilistic model-based clustering . . . 10
2.4.1 The mixture likelihood approach . . . 10
2.4.1.1 The EM algorithm for mixture models . . . 11
2.4.1.2 The EM algorithm for Gaussian mixture models . . . 11
2.4.1.3 The Bayesian approach for model selection and BIC . . . 12
2.4.2 The classification likelihood approach . . . 13
2.4.2.1 The CEM algorithm for mixture models . . . 13
2.4.2.2 The CEM algorithm for Gaussian mixture models . . . 14
2.4.2.3 AWE and ICL for model selection . . . 14
2.4.3 The DAEM algorithm . . . 15
2.4.3.1 The DAEM algorithm for Gaussian mixture models . . . . 16
3 BIC-based self-splitting Gaussian mixture learning 17 3.1 The SGML algorithm . . . 17
3.1.1 Computational complexity of SGML . . . 18
3.2 The fastSGML algorithm . . . 19
3.3 Experiments on Gaussian mixture learning . . . 19
3.3.1 Results on the synthetic data . . . 20
3.4 Application of SGML to GMM-based speaker identification . . . 24
3.4.1 The GMM-based speaker identification . . . 24
3.4.2 Experiments . . . 26
3.4.2.1 Database description and feature extraction . . . 26
3.4.2.2 Configuration of model training and test utterance . . . . 27
3.4.2.3 Results . . . 27
4 Model-based clustering by probabilistic self-organizing maps 34 4.1 Formulation of the coupling-likelihood mixture model for PbSOMs . . . 34
4.2 The SOCEM algorithm for learning PbSOMs . . . 36
4.2.1 SOCEM - a DA variant of CEM for GMM . . . 37
4.2.2 Relation to Kohonen’s batch algorithm . . . 39
4.2.3 Computational cost . . . 40
4.3 The SOEM algorithm for learning PbSOMs . . . 41
4.3.1 SOEM - a DA variant of EM for GMM . . . 42
4.3.2 Computational cost . . . 42
4.4 The SODAEM algorithm for learning PbSOMs . . . 43
4.4.1 Computational cost . . . 44
4.5 Relation to other algorithms . . . 44
4.5.1 For SOCEM . . . 45
4.5.2 For SOEM and SODAEM . . . 46
4.6 Experiments on organizing property and data clustering . . . 46
4.6.1 Experiments on organizing property . . . 46
4.6.1.1 Results on the synthetic data . . . 47
4.6.1.2 Results on PenRecDigits C0 . . . 49
4.6.2 Experiments to evaluate the performance of data clustering . . . 50
4.6.2.1 Results on ImgSeg by using SOCEM and SODAEM C . . 51
4.6.2.2 Results on ImgSeg by using SOEM and SODAEM M . . . 51
4.6.2.3 Results on Ecoli . . . 52
4.7 Application of SOCEM, SOEM, and SODAEM to data visualization and analysis . . . 52
4.7.1 Experiment results on ImgSeg by using SOCEM and SODAEM C . 53 4.7.2 Experiment results on ImgSeg by using SOEM and SODAEM M . . 54
4.7.3 Experiment results on Ecoli . . . 54
5 BIC-based audio segmentation using divide-and-conquer 69 5.1 Window-growing-based segmentation . . . 71
5.1.1 ∆BIC as the distance measure of two audio segments . . . 71
5.1.2 One-change-point detection . . . 72
5.2 Divide-and-Conquer-based segmentation . . . 73
5.2.1 The DACDec1 approach . . . 73
5.2.2 The DACDec2 approach . . . 75
5.2.3 Sequential segmentation by DACDec1 and DACDec2 . . . 77
5.3 Computational cost analysis . . . 78
5.3.1 For DACDec1 . . . 79 5.3.2 For DACDec2 . . . 80 5.3.3 For FixSlid . . . 81 5.3.4 For WinGrow . . . 81 5.3.5 Discussion . . . 82 5.4 Experiments . . . 83
5.4.1 Experiments on the synthetic data . . . 83
5.4.2 Experiments on broadcast news data . . . 84
6 Conclusion and future work 89 6.1 Conclusion . . . 89
6.2 Future work . . . 90
A 92 A.1 The SOCEM, SOEM, and SODAEM algorithms where mixture weights are learned . . . 92
B 95 B.1 Compute T1(k) . . . 95
B.2 Compute T0(m) . . . 96
List of Figures
2.1 (a) {A, B, · · ·, G} are data samples in <2. (b) An illustrative dendrogram of the data samples in (a) obtained by hierarchical agglomerative clustering. 8 3.1 The learning process of SGML on the synthetic data, where full covariance
GMMs are used. . . 22 3.2 The learning curves of SGML and the baseline approaches on the synthetic
data, where full covariance GMMs are used. . . 23 3.3 The learning curve of SGML on the synthetic data, where diagonal
covari-ance GMMs are used. . . 23 3.4 The learning curves of applying the various GMM learning methods to the
15-second speech data of #5007 in NIST01SpkEval. (a) and (b) depicts the full covariance case, while (c) and (d) depicts the diagonal covariance case. . . 25 3.5 The learning curves of applying fastSGML, K-means-BinSplitting and SGML
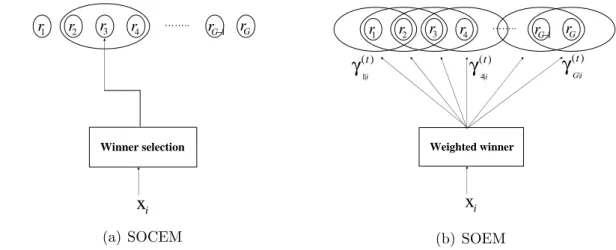
to learn diagonal covariance GMMs with 60-second speech data of #5007 in NIST01SpkEval. The splitting confidence of the fastSGML was set at 150, 100, and 50, respectively, and K-means-BinSplitting was forced to stop at GMM43. . . 26 4.1 (a) The network structure of a Gaussian mixture model, and (b) the
pro-posed coupling-likelihood mixture model. Here, rl(xi; θl) denotes the
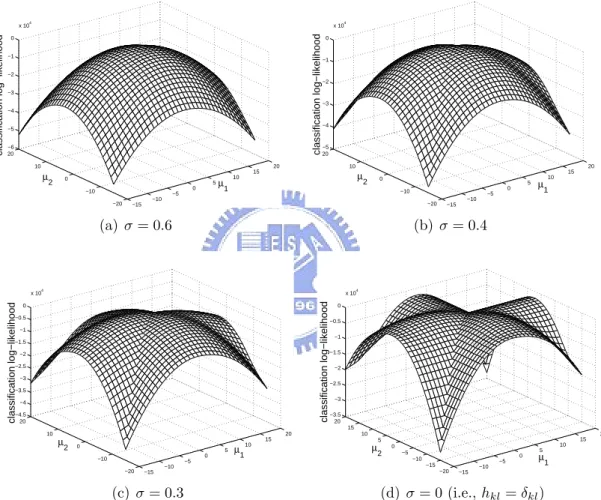
mul-tivariate Gaussian distribution described in Eq. (2.12). . . 55 4.2 SOCEM’s objective function becomes more complex with the reduction of
neighborhood size (σ in hkl). . . 56
4.3 For each data sample xi, the adaptation of the reference models in SOCEM
is restricted to the winning reference model and its neighborhood. However, in SOEM, the winner is relaxed to the weighted winners by the posterior probabilities γk|i(t), for k = 1, 2, · · · , G. Each data sample xi contributes
pro-portionally to the adaptation of each reference model and its neighborhood according to the posterior probabilities. . . 57
4.4 The family of Gaussian model-based clustering algorithms derived from the
SODAEM, SOEM and SOCEM algorithms. δkl = 1 if k = l; otherwise,
δkl = 0. . . 57
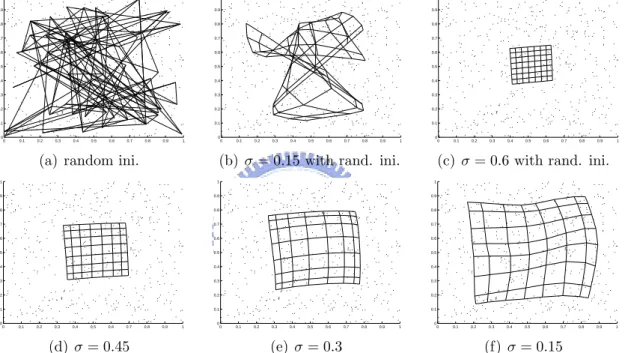
4.5 The map-learning process obtained by running the SOCEM algorithm on the synthetic data. Simulation 1 ((a)-(b)): When SOCEM is run with the random initialization in (a) and σ = 0.15, it converges to the unordered map in (b). Simulation 2 ((a) and (c)-(f)): SOCEM starts with σ = 0.6 and the random initialization in (a). Then, the value of σ is reduced to 0.15 in 0.15 decrements. . . 58 4.6 The map-learning process obtained by running the SOEM algorithm on
the synthetic data. Simulation 1 ((a)-(b)): When SOEM is run with the random initialization in (a) and σ = 0.15, it converges to the unordered map in (b). Simulation 2 ((a) and (c)-(f)): SOEM starts with σ = 0.6 and the random initialization in (a). Then, the value of σ is reduced to 0.15 in 0.15 decrements. . . 59 4.7 The map-learning process obtained by running the SODAEM algorithm
on the synthetic data. The value of σ is fixed at 0.15, while value of β is initialized at 0.16 and increased in multiples of 1.6 up to 17.592. . . 60 4.8 The map-learning process obtained by running the SOCEM algorithm on
PenRecDigits C0. Simulation 1 ((a)-(b)): When SOCEM is run with the random initialization in (a) and σ = 0.15, it converges to the unordered map in (b). Simulation 2 ((a) and (c)-(f)): SOCEM starts with σ = 0.6 and the random initialization in (a). Then, the value of σ is reduced to 0.15 in 0.15 decrements. . . 61 4.9 The map-learning process obtained by running the SOEM algorithm on
PenRecDigits C0. Simulation 1 ((a)-(b)): When SOEM is run with the random initialization in (a) and σ = 0.15, it converges to the unordered map in (b). Simulation 2 ((a) and (c)-(f)): SOEM starts with σ = 0.6 and the random initialization in (a). Then, the value of σ is reduced to 0.15 in 0.15 decrements. . . 62 4.10 The map-learning process obtained by running the SODAEM algorithm
on PenRecDigits C0. The value of σ is fixed at 0.15, while value of β is initialized at 0.16 and increased in multiples of 1.6 up to 17.592. . . 63 4.11 The data clustering performance of CEM, DAEM C, SOCEM, SODAEM C,
and KohonenGaussian on ImgSeg in terms of the classification log-likelihood. 64 4.12 Learning a Gaussian mixture model by applying EM, DAEM M, SOEM,
and SODAEM M to ImgSeg. . . 64 4.13 The data clustering performance on Ecoli in terms of (a) the classification
4.14 Data visualization for ImgSeg by running KohonenGaussian ((b)), SOCEM ((c), (d)), and SODAEM C ((e), (f)) with the random initialization in (a). The network structure is a 7 × 7 equally spaced square lattice in a unit square. . . 66 4.15 Data visualization for ImgSeg by running SOEM ((a), (b)) and SODAEM M
((c), (d)) with the random initialization in Figure 4.14 (a). The network
structure is a 7 × 7 equally spaced square lattice in a unit square. . . . . 67
4.16 Data visualization for Ecoli by running (b) KohonenGaussian, (c) SOCEM, (d) SOEM, (e) SODAEM C, and (f) SODAEM M with the random initial-ization in (a). The network structure is a 7 × 7 equally spaced square lattice in a unit square. . . 68 5.1 The fixed-size sliding window detection approach. . . 71 5.2 Diagram of the multiple-change-point detection in window-growing-based
segmentation (WinGrow). The audio stream contains three segments, namely Seg1, Seg2, and Seg3; P and Q denote the change points. . . 73 5.3 (a) An audio stream comprised of three speech segments, each derived from
a distinct speaker. C1 and C2 are the change points. (b) The ∆BIC curve
obtained by applying OCD-Chen to the audio stream in (a). . . 74 5.4 An illustration that data samples distribute as three Gaussian clusters.
For this case, generally, two Gaussians (H1) fit the distribution of the
data better than one Gaussian (H0) if the samples belonging to the same
Gaussian cluster are used together to estimate the parameters. . . 76 5.5 (a) An audio stream comprised of three speech segments; the first and third
segments are derived from the same speaker (Speaker1), while the second is derived from another speaker (Speaker2). (b) The ∆BIC curve obtained by applying OCD-Chen to the audio stream in (a). (c) The diagram of
the hypothesis test at the change point C2 in (b). (d) The diagram of the
hypothesis test at the non-change point R in (b). . . 77 5.6 A recursive tree that simulates the recursive process of DACDec2 on the
audio stream in Figure 5.5 (a). . . 79 5.7 Diagram of the detection process of SeqDACDec1 and SeqDACDec2. If a
change point is detected in the fixed-size analysis window by DACDec1 or DACDec2, the window is moved to the change point with the largest time index. Otherwise, it is moved forward by ηL samples, where L denotes the window size, and η > 0. . . 80 5.8 An audio stream comprised of k+1 homogeneous segments, each containing
5.9 ROC curves obtained by running SeqDACDec1 and SeqDACDec2 on the synthetic data using 10-second, 20-second, and 30-second analysis windows.
L denotes the size of the analysis window. . . 84
5.10 The empirical cumulative distributions of the size of homogeneous segments in MATBN3hr and HUB4-98. . . 85 5.11 A significant local maximum on the distance curve. . . 86
5.12 The ROC curves for MATBN3hr obtained by (a) SeqDACDec1 with Nmin =
2 seconds and analysis windows of different size; (b) SeqDACDec2 with
Nmin = 2 seconds and analysis windows of different size; and (c)
Seq-DACDec1 with Nmin = 2 seconds and L = 20 seconds, SeqDACDec2 with
Nmin = 2 seconds and L = 20 seconds, WinGrow with Nmin = 3 seconds
and Nmax = 20 seconds, and FixSlid with a 2-second window. . . 87
5.13 The ROC curves for HUB4-98. . . 88 A.1 The map-learning process obtained by running the SOCEM algorithm on
the synthetic data with an ordered initialization in (a). Simulation 1 ((a)-(e)): The mixture weights are initialized at 1
16, and updated in the learning process; the algorithm starts with the initialization in (a) and converges to the unordered map in (e). Simulation 2 ((a) and (f)): SOCEM is performed with equal mixture weights throughout the learning process; the algorithm starts with the initialization in (a) and converges to the map in (f). The network structure is a 4 × 4 square lattice; the value of σ is set at 0.4. . . 94
List of Tables
2.1 Common inter-cluster distance measures for hierarchical agglomerative clus-tering. d(xi, xj) denotes the distance measure for xi and xj. . . 9
3.1 The mean and standard deviation of the component number of the diago-nal covariance male speaker GMMs obtained by SGML and fastSGML on different amounts of training data. The first number in parentheses is the mean value, while the second number after ’/’ is the standard deviation. . 29 3.2 The mean and standard deviation of the component number of the diagonal
covariance female speaker GMMs obtained by SGML and fastSGML on different amounts of training data. The first number in parentheses is the mean value, while the second number after ’/’ is the standard deviation. . 29 3.3 The average CPU time (in second) of the diagonal covariance male speaker
GMMs obtained by SGML and fastSGML on different amounts of training data. . . 30 3.4 The average CPU time (in second) of the diagonal covariance female speaker
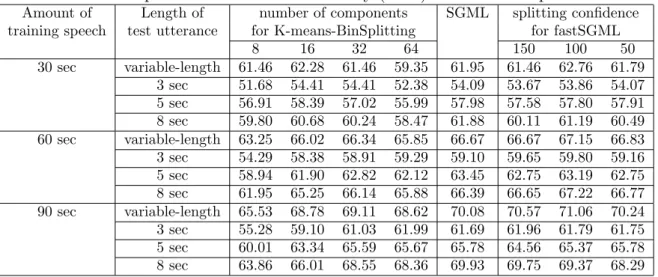
GMMs obtained by SGML and fastSGML on different amounts of training data. . . 30 3.5 Speaker identification accuracy (in %) for the male speakers. . . 30 3.6 Speaker identification accuracy (in %) for the female speakers. . . 31 4.1 The DAEM algorithm for learning GMMs with equal mixture weights and
the SOCEM algorithm. . . 40 4.2 Results of simulations using KohonenGaussian, SOCEM, SOEM, and
SO-DAEM in 20 independent random initialization trials on the synthetic data.
The algorithms were run with two setups for σ in hkl. When σ = 0.15,
Ko-honenGaussian succeeded in converging to an ordered map in one random initialization case (S:1), but failed in the remaining cases (F:19). . . 49 4.3 Results of simulations using KohonenGaussian, SOCEM, SOEM, and
SO-DAEM in 20 independent random initialization trials on PenRecDigits C0.
The algorithms were run with two setups for σ in hkl. When σ = 0.15,
Ko-honenGaussian succeeded in converging to an ordered map in one random initialization case (S:1), but failed in the remaining cases (F:19). . . 50
5.1 The CPU time of different audio segmentation approaches evaluated on MATBN3hr in the EER case and the associated EERs, where M and F denote the miss detection rate and the false alarm rate, respectively. . . 86 5.2 The CPU time of different audio segmentation approaches evaluated on
HUB4-98 in the EER case and the associated EERs. . . 87 A.1 The mixture weights learned by SOCEM with the initialization in Figure
A.1 (a). The mixture weights are initialized at 1
Chapter 1
Introduction
1.1
Background
The goal of data clustering is to group data samples into clusters such that samples from the same cluster are more similar to each other than samples from different clusters. It is also known as unsupervised learning, numerical taxonomy, typological analysis and vector quantization [1, 2]. Among the various clustering approaches, probabilistic model-based clustering is the one derived from statistical learning; and it has been successfully applied to many tasks, for example, speaker recognition [3, 4], speech recognition [5], handwritten recognition [6], image segmentation [7], and clustering of microarray expression data [8, 9]. In model-based clustering, data samples are grouped by learning a finite mixture model (usually a Gaussian mixture model, i.e., GMM), in which each mixture component represents a cluster. There are two major learning methods for model-based clustering: the mixture likelihood approach, where the likelihood of each data sample is a mixture of all the component likelihoods of the data sample; and the classification likelihood approach, where the likelihood of each data sample is generated by its winning component only [10, 11, 12, 13, 14, 15, 16, 17, 18]. In both approaches, when the globally optimal estimation of the model parameters cannot be obtained analytically, iterative learning algorithms that only guarantee obtaining locally optimal solutions are usually employed. The expectation-maximization (EM) algorithm for mixture likelihood learning [19, 20, 21, 22] and the classification EM (CEM) algorithm for classification likelihood learning [17] are two such algorithms and have been the dominant approaches in this task. The conventional EM or CEM-based model-based clustering has three critical aspects, namely the initialization
of model parameters, the model complexity1, and the topology-preserving ability. These
aspects are discussed as follows.
• For model initialization: The learning performance of EM and CEM are very
sen-sitive to the initial conditions of the model’s parameters. To address this issue, the
authors in [17] proposed a simulated annealing implementation for CEM, which re-duces the initial-position dependence based on random perturbations. Rather than applying the randomization power of simulated annealing, the authors in [23] pro-posed a deterministic annealing EM (DAEM) algorithm that tackles the initializa-tion issue via a deterministic annealing process. DAEM was originally derived as a DA variant of EM; however, as shown in Section 2.4.3, it is also a DA variant of CEM. Although DAEM has been reported achieving decent performance, there is still no guarantee that it finds the globally optimal model parameters because, like EM, it is a iterative, single-token search scheme. In order to search the parameter space in multiple pathes, the authors in [24, 25] and [26, 27] proposed multi-thread search strategies when employing EM and DAEM, respectively, where the search pathes are adapted in the search process according to the eigen-decomposition of the Hessian matrix of the target log-likelihood function. As another kind of multiple-path search, a GA-EM algorithm was proposed in [28], where EM learning is integrated into a Genetic search procedure. It is less sensitive to the initialization because of the sto-chastic search nature of the Genetic Algorithms (GA). Some heuristic-like learning algorithms have also been proposed. For example, an initialization approach for EM which is based on subsampling was presented in [29]. The authors in [30] proposed an SMEM algorithm that finds the appropriate initial conditions for EM learning by using split and merge operations. Similarly, Young and Woodland [31] proposed a component-splitting approach to learn a GMM, which iteratively splits the mean vector of the Gaussian component with the largest weight into two new ones, and then performs EM to update all the Gaussian components. Another method based on component splitting is presented in [32]. In [33], the authors suggested a simple way that one can perform several short runs of EM (by early stopping) first, then select the best model from the results and use them as the initial model for the long run (standard) EM learning. As a common and simple way, one can apply K-means clustering or hierarchical agglomerative clustering (HAC) to locate the initial mean vectors of Gaussian components for the EM learning [1, 34, 35].
Note that all the approaches mentioned above are performed with a given target number of mixture components.
• For model complexity: Assessment of the number of mixture components (data
clusters) is an important issue in model-based clustering. The mixture model would over-fit the data if it contains too many mixture components; in contrast, it would not be flexible enough to describe the structure of the data if the number of com-ponents is too small. Various approaches have been proposed to address this issue. In [36], Furman and Lindsay developed two hypothesis test procedures based on the moment estimators to assess the number of components for a GMM. As
an-other hypothesis test-based approach, McLachlan and Khan estimated the number of components by likelihood ratio test, where the re-sampling process is applied to assess its null hypothesis [37]. In [38], the authors estimated the mixture complex-ity by comparing an information theoretic-derived nonparametric estimator with the best parametric fit of a given complexity. As another information theoretic-based approach, a maximum entropy-based approach with a modified EM algorithm was proposed to assess the model complexity of GMM in [39]. Moreover, model se-lection criteria, also known as penalized-likelihood criteria, have been proposed to assess the model complexity; for example, Akaike’s Information Criterion (AIC) [40], Bayesian Information Criterion (BIC) [41, 10, 42], Integrated Completed Like-lihood (ICL) [43], Approximate Weight Evidence (AWE) [18], Minimum Description Length (MDL) [44] (which is formally identical to BIC), and Minimum Message Length (MML) [45]. BIC is derived on the basis of mixture likelihood, ICL and AWE are derived on the basis of classification likelihood, and AIC, MDL and MML are information-theoretic-derived criteria. A common way to applying these criteria
to assess model complexity is that defining the upper bound, Gmax, of the
compo-nent number of candidate models first, and then choosing the one with the best score calculated using the employed criterion as the best model. For example, when using BIC, the best component number is
ˆ
G = arg max
G {2 log p(X ; ˆΘG) − P enalty(G)|G = 1, 2, . . . , Gmax}, (1.1)
where ˆΘG is the maximum likelihood estimate of parameters of the mixture model
with G components, P enalty(G) is a monotonically increasing function of G that penalizes more for a more complex model [10]. However, there are two potential drawbacks with this model-selection-based approach. First, it needs to define the
upper bound of the component number, Gmax, beforehand. On the one hand, if
Gmax is too large (much larger than the best component number determined by
the model selection criterion), the learning process will waste a lot of computation time. On the other hand, if Gmax is too small, the selected model may be not flexible
enough to describe the structure of the data. Second, ˆΘG is usually obtained by
EM, whose performance is highly dependent on the model initialization.
Rather than assessing the model complexity by incrementally adding mixture com-ponents during the learning process, as discussed above, the variational Bayesian framework in [46, 47] automatically determines the number of components by setting a larger component number initially and then suppressing unwanted components.
• For topology-preserving ability: Conventional model-based clustering cannot
clustering procedure. To overcome this shortcoming, the clustering task can be per-formed by using Kohonen’s self-organizing map (SOM) [48, 49]. The SOM, rather than being a supervised neural network model for pattern recognition, is an unsuper-vised model for data clustering and visualization. After SOM’s clustering procedure, the topological relationships among data samples and clusters can be preserved (or visualized) on the network, which is usually a two dimensional lattice. Kohonen’s sequential and batch SOM learning algorithms have proved successful in many prac-tical applications [48, 49]. However, they also suffer from some shortcomings, such as the lack of an objective (cost) function, a general proof of convergence, and a prob-abilistic framework [50]. Some related works that have addressed these issues are as follows. In [51, 52], the behavior of Kohonen’s sequential learning algorithm was studied in terms of energy functions, based on which, Cheng [53] proposed an energy function for SOM whose parameters can be learned by a K-means type algorithm. Luttrell [54, 55] proposed a noisy vector quantization model called the topographic vector quantizer (TVQ), whose training process coincides with the learning process of SOM. The cost function of TVQ represents the topographic distortion between the input data and the output code vectors in terms of Euclidean distance. Graepel
et al. [56, 57] derived a soft topographic vector quantization (STVQ) algorithm
by applying a deterministic annealing process to the optimization of TVQ’s cost function. Based on the topographic distortion concept, Heskes [58] applied a dif-ferent DA implementation from that of STVQ, and obtained an algorithm identical to STVQ when the quantization error is expressed in terms of Euclidean distance. In [59], Chow and Wu proposed an on-line algorithm for STVQ; later, motivated by STVQ, they proposed a data visualization method that integrates SOM and multi-dimensional scaling [60]. Based on the Bayesian analysis of SOMs in [61], Anouar et al. [62] proposed a probabilistic formalism for SOM, where the para-meters are learned by a K-means type algorithm. To help users select the correct model complexity for SOM by probabilistic assessment, Lampinen and Kostiainen [63] developed a generative model in which the SOM is trained by Kohonen’s algo-rithm. Meanwhile, Van Hulle [64] developed a kernel-based topographic formation in which the parameters are adjusted to maximize the joint entropy of the kernel outputs. He subsequently developed a new algorithm with heteroscedastic Gaussian mixtures that allows for a unified account of vector quantization, log-likelihood, and Kullback-Leibler divergence [65]. Another probabilistic formulation is proposed in [66], whereby a normalized neighborhood function of SOM is used as the posterior distribution in the E-step of the EM algorithm for a mixture model to enforce the self-organizing of the mixture components. Sum et al. [67] interpreted Kohonen’s sequential learning algorithm in terms of maximizing the local correlations (cou-pling energies) between neurons and their neighborhoods for the given input data.
They then proposed an energy function for SOM that reveals the correlations, and a gradient ascent learning algorithm for the energy function.
1.2
Contributions of this dissertation
1.2.1
On model initialization and complexity
This thesis proposes a BIC-based self-splitting Gaussian mixture learning (SGML) algo-rithm that starts with a single component and iteratively splits a component into two new components until the most appropriate component number is found. The proposed algorithm has several advantages as follows. 1) SGML provides a better initialization for EM. For the Gaussian mixture learning process whose initialization is based on a data clustering process like K-mens or HAC, the clustering phase may results in an poor initial mixture model with too many components in one part of the space and too few in another widely-separated part of the space; in this case, the following EM phase may fail to escape from this configuration and fall into an poor local maximum. In the splitting process of SGML, however, BIC is used to determine which part of the space should be divided into two parts. In this way, the ill initialization situation can be avoided to some extent and, thus, a better estimation for model parameters can be obtained. 2) SGML automati-cally determines the appropriate component number without the need to define the upper bound of the component number beforehand. It stops when a “significant maximum” in the learning curve (i.e., the BIC plot) is found, and then outputs the model yielding the maximum BIC valve. 3) SGML is deterministic; its output is always the same in different runs on the same data set since it does not contain any randomization procedure in its learning rules.
1.2.2
On topology-preserving ability
In Kohonen’s SOM architecture, neurons in the network associate with reference vectors in the data space. This contrasts with a SOM whose neurons associate with reference models that present probability distributions, such as the isotropic Gaussians used in [66] and the heteroscedastic Gaussians used in [62, 65]. In this thesis, the latter is called a probabilistic SOM (PbSOM). Motivated by the coupling energy concept in Sum et al.’s work [67], a coupling-likelihood mixture model for the PbSOM that uses multivariate Gaussian distributions as the reference models is developed. In the proposed model, local coupling energies between neurons and their neighborhoods are expressed in terms of probabilistic likelihoods; and each mixture component expresses the local coupling-likelihood between one neuron and its neighborhood. Based on this model, we develop CEM, EM, and DAEM algorithms for learning PbSOMs, namely the SOCEM, SOEM, and SODAEM algorithms, respectively. SODAEM is a DA variant of SOCEM and SOEM.
Moreover, we show that SOCEM and SOEM can be interpreted, respectively, as DA variants of the CEM and EM algorithms for Gaussian model-based clustering, where the neighborhood shrinking is interpreted as an annealing process. The experiment results show that the proposed PbSOM learning algorithms achieve comparable data clustering performance to the DAEM algorithm, while maintaining the topology-preserving property.
1.2.3
On applications
In this thesis, we apply SGML to the training of speaker GMMs for the speaker identi-fication task. The experiment results on NIST 2001 speaker recognition evaluation [68] show that SGML can automatically determine the appropriate model complexity for a speaker GMM, which is usually empirically determined in the literature [4, 69]. The proposed PbSOM algorithms are applied to data visualization and analysis. The exper-iment results on UCI data sets show that we can explore the topological relationships between data samples and clusters on a two-dimensional network. Moreover, we propose divide-and-conquer approaches for the audio segmentation task using BIC to evaluate the dissimilarity between two audio segments. The experiment results on the broadcast news data demonstrate that our approaches not only have a lower computation cost but also achieve a higher segmentation accuracy than the leading window-growing-based segmen-tation approach [70, 71].
1.3
Organization of this dissertation
To help readers understand the content of this thesis, we briefly review K-means clustering, hierarchical agglomerative clustering, Kohonen’s SOM, and model-based clustering in Chapter 2. In Chapter 3, we describe the SGML algorithm for learning a Gaussian mixture model and its application to speaker identification (based on [72]). In Chapter 4, we describe the SOCEM, SOEM, and SODAEM algorithms for model-based clustering where the topological relationships between data samples and clusters can be preserved on a network and their application to data visualization and analysis (based on [73]). We present the application of BIC to audio segmentation (based on [74]) in Chapter 5. Finally, we give the conclusion and discuss our future works in Chapter 6.
Chapter 2
Preliminaries and related works
2.1
K-means clustering
Given a data set X = {x1, x2, · · · , xN} ⊂ <d, data clustering can be achieved based on
learning the vector prototypes {m1, m2, · · · , mG} ⊂ <d; after the learning process, X is
partitioned into clusters P = {P1, P2, · · · , PG} by classifying each data sample to the
cluster associated with its nearest prototype. For the learning, the goal is to find the partition and the prototypes that minimize the distance function
D(P, {m1, m2, · · · , mG}; X ) = G X k=1 X xi∈Pk kxi− mkk2, (2.1)
which can not be minimized analytically.
K-means clustering is a popular approach to learning the prototypes in order to
min-imize Eq. (2.1). Given the initial prototypes, it iteratively and alternatively applies a classification step and a distance minimization step on the data samples and prototypes as follows.
• Nearest-neighbor classification: Given the current prototypes, {m(t)1 , m(t)2 , · · · , m(t)G},
assign each data sample to the cluster associated with its nearest prototype, i.e., xi ∈ ˆPj(t) if j = arg minkkxi− m(t)k k2.
• Distance minimization: Suppose we have the clusters { ˆP1(t), ˆP2(t), · · · , ˆPG(t)} after the
classification step; then, by minimizing Px
i∈ ˆPk(t)kxi − mkk
2 with respect to m
k,
we obtain the update rules for the prototypes as: m(t+1)k = 1
| ˆPk(t)|
P
xi∈ ˆPk(t)xi, for
k = 1, 2, · · · , G.
Although the K-means clustering algorithm is simple and efficient, it only guaran-tees converging to a local minimum of the distance function in Eq. (2.1). Besides, it has two more shortcomings that the performance highly depends on the initialization of
1
x
2x
(a) A B C D E F G {B ,C } {A ,B ,C } {D ,E } {F,G } {D ,E ,F,G } {A ,B ,C ,D ,E ,F,G } P ro xi m ity (b)Figure 2.1: (a) {A, B, · · ·, G} are data samples in <2. (b) An illustrative dendrogram of the data samples in (a) obtained by hierarchical agglomerative clustering.
prototypes and the number of prototypes need to be defined beforehand. To overcome the initialization issue, the authors in [75] proposed to iteratively split each mean vec-tor into two new ones until the desired number of clusters is reached. Similarly, in [76], the authors proposed iteratively splitting the mean vector of the cluster with the largest accumulated distance between data samples and its prototype until the desired number of clusters is reached. As a simple way, one may conduct the K-means algorithm with random initialization for many runs, say 20, and then select the best one.
2.2
Hierarchical agglomerative clustering
Different from K-means clustering that performs data clustering based on prototype learn-ing, hierarchical agglomerative clustering (HAC) performs the clustering according to the proximity between clusters. When performing HAC, each data sample is considered as a cluster initially; then, the algorithm iteratively merges the two clusters with the smallest distance into a new cluster. For example, if HAC is applied to clustering {A, B,· · ·,G} in Figure 2.1 (a) with Euclidean distance, we may obtain a dendrogram (tree) that rep-resents the hierarchy of the clusters in Figure 2.1 (b). As shown in the figure, B and C are merged into a cluster first, then D and E are merged, and so on. And we can obtain different configurations of clusters by cutting the dendrogram according to the proximity. As for the inter-cluster distance measure, the most commonly used ones are the so-called single-linkage, average-linkage, and complete-linkage approaches defined in Table 2.1. For more discussions of these distance measure, reader can refer to [77].
Table 2.1: Common inter-cluster distance measures for hierarchical agglomerative clus-tering. d(xi, xj) denotes the distance measure for xi and xj.
Approach distance between clusters Pm and Pn
Single-linkage minxi∈Pm,xj∈Pnd(xi, xj) Average-linkage 1 |Pm||Pn| P xi∈Pm P xj∈Pnd(xi, xj) Complete-linkage maxxi∈Pm,xj∈Pnd(xi, xj)
2.3
Self-organizing map
Self-organizing map (SOM) is a neural network model for data clustering that preserves the topological relationships between data samples and clusters in a network [48, 49]. Like K-means clustering, the training of SOM is a prototype-learning process. How-ever, in addition to the interconnections between the data samples and the prototypes as in K-means clustering, SOM involves lateral interactions between prototypes via a neighborhood definition on the network to learn the topological relationships between the prototypes. Kohonen’s sequential (incremental) and batch learning algorithms have been well recognized and successfully applied to many tasks, such as data visualization [48], document processing [78, 79], image processing [80], and speech precessing [81, 82, 83].
Kohonen’s SOM model consists of G neurons R={r1, r2, · · · , rG} on a network with
a neighborhood function hkl that defines the strength of lateral interaction between two
neurons, rk and rl, for k, l ∈ {1, 2, · · · , G}; and each neuron rk associates with a vector
prototype mk(a reference vector) in the input data space [48, 49]. In Kohonen’s sequential
learning algorithm, the training data samples x1, x2, · · · , xN are applied sequentially. For
each data sample, say xi, its winning prototype mwin(t)
i such that
win(t)i = arg min
j kxi− m
(t)
j k, (2.2)
is retrieved first, then the prototypes are adapted by the following rules: m(t+1)k = m(t)k + α(t)hwin(t)
i k(xi− m (t)
k ), (2.3)
for k = 1, 2, · · · , G. α(t) is the learning-rate factor such that 0 < α(t) < 1; it decreases monotonically with the increasing of the learning iterations. The neighborhood function
hkl is usually a rectangular function, i.e., hkl = 1 for rl that is in the neighborhood of rk
and hkl = 0 for rl that is not in the neighborhood of rk; or a monotonically decreasing
function of the Euclidean distance krk− rlk between rk and rl on the SOM network, for
example, the widely applied unnormalized Gaussian neighborhood function
hkl = exp(−
krk− rlk2
Kohonen’s batch learning algorithm applies all the data samples as a whole to up-date the prototypes, rather than one by one as in the sequential learning algorithm. In each learning iteration, the winning prototype for each sample is found first; then, the prototypes are updated by
m(t+1)k = PN i=1hwin(t)i kxi PN i=1hwin(t)i k , (2.5) for k = 1, 2, · · · , G.
Kohonen’s learning algorithms need to be applied with two stages. First, a large neighborhood is applied to the algorithm to learn an ordered map near the center of the data samples. Then, the prototypes are adapted to fit the distribution of the data samples by gradually shrinking the neighborhood. When the neighborhood is reduce to zero-neighborhood, i.e., hkl = δkl where δkl = 1 if k = l and otherwise δkl = 0, Kohonen’s
batch learning algorithm becomes the K-means clustering algorithm.
2.4
Probabilistic model-based clustering
2.4.1
The mixture likelihood approach
In the mixture likelihood approach for model-based clustering, it is assumed that the given data set X = {x1, x2, · · · , xN} ⊂ <d is generated by a set of independently and
identically distributed (i.i.d.) random vectors from a mixture model:
p(xi; Θ) = G
X
k=1
w(k)p(xi; θk), (2.6)
where w(k) is the mixing weight of the mixture component p(xi; θk), subject to 0 ≤
w(k) ≤ 1 for k = 1, 2, · · · , G; PG
k=1w(k) = 1; and θk denotes the parameter set of
p(xi; θk).
The maximum likelihood estimate of the parameter set of the mixture model ˆΘ =
{ ˆw(1), ˆw(2) , · · · , ˆw(G), ˆθ1, ˆθ2, · · · , ˆθG} can be obtained by maximizing the following
log-likelihood function: L(Θ; X ) = log N Y i=1 p(xi; Θ) = N X i=1 log( G X k=1 w(k)p(xi; θk)). (2.7)
This is usually achieved by using the expectation-maximization (EM) algorithm [20, 22]. After learning the mixture model, we derive a partition of X , ˆP = { ˆP1, ˆP2, · · · , ˆPG}, by
for xi, i.e., xi ∈ ˆPj if j = arg maxkp(k | xi; ˆΘ).
2.4.1.1 The EM algorithm for mixture models
If the maximum likelihood estimation of the parameters cannot be accomplished analyti-cally, the EM algorithm is normally used as an alternative approach when the given data is incomplete or contains hidden information.
In the case of the mixture model, suppose that Θ(t) denotes the current estimate of
the parameter set, and k is the hidden variable that indicates the mixture component from which the data sample is generated. The E-step of EM algorithm then computes the following so-called auxiliary function:
Q(Θ; Θ(t)) = XN i=1 G X k=1 p(k | xi; Θ(t)) log p(xi, k; Θ), (2.8) where p(xi, k; Θ) = w(k)p(xi; θk), (2.9) and p(k | xi; Θ(t)) = w(k)(t)p(x i; θ(t)k ) PG j=1w(j)(t)p(xi; θ(t)j ) (2.10) denotes the posterior probability of the kth mixture component for xi with the given Θ(t).
Then, in the following M-step, the Θ(t+1) that satisfies
Q(Θ(t+1); Θ(t)) = max
Θ Q(Θ; Θ
(t)) (2.11)
is chosen as the new estimate of the parameter set. By iteratively creating the auxiliary function in Eq. (2.8) and performing the subsequent maximization step, the EM algorithm guarantees to converge to a local maximum of the log-likelihood function in Eq. (2.7).
When Q(Θ; Θ(t)) can not be maximized analytically, the M-step is modified to find
some Θ(t+1) such that Q(Θ(t+1); Θ(t)) > Q(Θ(t); Θ(t)). This type of algorithm, called Generalized EM (GEM), is also guaranteed to converge to a local maximum [20, 22].
2.4.1.2 The EM algorithm for Gaussian mixture models
Suppose Eq. (2.6) is a Gaussian mixture model where
p(xi; θk) = 1 (2π)d/2|Σ k|1/2 exp(−1 2(xi− µk) TΣ−1 k (xi− µk)), (2.12)
is the multivariate Gaussian distribution and θk = {µk,Σk} are its mean vector and
covariance matrix. Then, each iteration of the EM algorithm for learning GMMs is summarized as follows.
• E-step: Given the current model, Θ(t), compute the posterior probability of each mixture component of p(xi; Θ(t)) for each xi using Eq. (2.10).
• M-step: Substituting Eq. (2.12) into Eq. (2.8) and then setting its derivative to
zero, we obtain the re-estimation formulae of the parameters as follows.
w(k)(t+1)= 1 N N X i=1 p(k | xi; Θ(t)), (2.13) µ(t+1)k = PN i=1p(k | xi; Θ(t))xi PN i=1p(k | xi; Θ(t)) , (2.14) Σ(t+1)k = PN i=1p(k | xi; Θ(t))(xi− µ(t+1)k )(xi− µ(t+1)k )T PN i=1p(k | xi; Θ(t)) . (2.15)
2.4.1.3 The Bayesian approach for model selection and BIC
Given the data set X and a set of candidate probability models M = {MG | G =
1, . . . , Gmax} where model MG associates with a parameter set ΘG, the posterior
proba-bility p(MG | X ) can be used to select the appropriate model from M to represent the
distribution of X . According to Bayes’ theorem, p(MG | X ) can be expressed as
p(MG | X ) =
p(MG)p(X ; MG)
p(X ) , (2.16)
where p(MG) is the prior probability of model MG. p(X ) can be ignored because it is
identical for all models and does not affect the selection. Moreover, it is assumed that each model is equally likely (i.e., p(MG) = 1/Gmax); then, p(MG | X ) is proportional to the
probability that the data conforms to the model MG, p(X ; MG), which can be computed
by p(X ; MG) = Z p(X ; ΘG, MG)p(ΘG; MG)dΘG, = Z p(X ; ΘG)p(ΘG)dΘG. (2.17)
The calculation of log p(X ; MG) can be achieved by the Laplace approximation [42, 47, 84],
which gives
log p(X ; MG) ≈ log p(X ; ˆΘG) −
1
2d(ΘG) log N, (2.18)
where ˆΘG is the maximum likelihood estimate of ΘG, d(ΘG) is the number of free
model MG over the data set X , BIC(MG, X ), is defined as1
BIC(MG, X ) ≡ 2 log p(X ; ˆΘG) − d(ΘG) log N. (2.19)
Accordingly, the larger the BIC value, the stronger the evidence for the model is. In other words, the model with the maximum BIC value will be selected. The BIC can be used to compare models with different parameterizations, different numbers of components, or both.
2.4.2
The classification likelihood approach
In the classification likelihood approach for model-based clustering [15, 16, 17], instead of maximizing the log-likelihood function of the mixture model in Eq. (2.7), the objective is to find the partition ˆP = { ˆP1, ˆP2, · · · , ˆPG} of X and the model parameters that maximize
C1(P, {θ1, θ2, · · · , θG}; X ) = G X k=1 X xi∈Pk log p(xi; θk), (2.20) or C2(P, Θ; X ) = G X k=1 X xi∈Pk log(w(k)p(xi; θk)). (2.21)
The relation between C1 and C2 is
C2(P, Θ; X ) = C1(P, {θ1, θ2, · · · , θG}; X ) + G
X
k=1
|Pk| log w(k), (2.22)
If all the mixture components are equally weighted,PG
k=1|Pk| log w(k) becomes a constant,
such that C1 and C2 are equivalent.
2.4.2.1 The CEM algorithm for mixture models
Celeux and Govaert [17] proposed the Classification EM (CEM) algorithm for estimating
the partition ˆP and model parameters. Like the EM algorithm, CEM is also an iterative
learning approach. In each iteration, it inserts a classification step (C-step) between the
E-step and M-step of the EM algorithm. In the E-step, the posterior probability of each
mixture component is calculated for each data sample. In the C-step, to obtain the parti-tion ˆP of the data samples, each sample is assigned to the mixture component that yields
the largest posterior probability for that sample. In the M-step, the maximization process is applied to ˆPk individually for k = 1, 2, · · · , G. From a practical point of view, CEM
1Kass and Raftery [85] defined BIC as minus the value given in Eq. (2.19); Fraley and Raftery [10]
used the BIC defined in Eq. (2.19) to make it easier to interpret the plot of BIC values. Here, we follow Fraley and Raftery’s BIC definition.
is a K-means type algorithm where a cluster is associated with a probability distribution rather than a vector prototype [17].
2.4.2.2 The CEM algorithm for Gaussian mixture models
Suppose the multivariate Gaussian is used as the mixture component, each iteration of the CEM algorithm for learning GMMs is summarized as follows.
• E-step: Given the current model, Θ(t), compute the posterior probability of each mixture component of p(xi; Θ(t)) for each xi using Eq. (2.10).
• C-step: Assign each xi to the cluster whose corresponding mixture component has
the largest posterior probability for xi, i.e., xi ∈ ˆPj(t) if j = arg maxkp(k|xi; Θ(t)).
• M-step: After the C-step, the partition of X (i.e., ˆP(t)) is formed, and the objective
function C2 defined in Eq. (2.21) becomes
C2(Θ; ˆP(t), X ) = G X k=1 X xi∈ ˆPk(t) log(w(k)p(xi; θk)). (2.23)
Substituting Eq. (2.12) into Eq. (2.23) and then setting the derivative of C2 with
respect to individual parameters to zero, we obtain the re-estimation formulae of the parameters as follows.
w(k)(t+1) = | ˆP (t) k | N , (2.24) µ(t+1)k = P xi∈ ˆPk(t)xi | ˆPk(t)| , (2.25) Σ(t+1)k = P xi∈ ˆPk(t)(xi− µ (t+1) k )(xi− µ(t+1)k )T | ˆPk(t)| . (2.26)
2.4.2.3 AWE and ICL for model selection
Based on classification likelihood, Banfield and Raftery proposed the AWE criterion [18, 86] to assess the number of Gaussian components (clusters) for a given data set. The AWE criterion is
AW E(G clusters; X ) = C1( ˆP, { ˆθ1, ˆθ2, · · · , ˆθG}; X ) − d({θ1, θ2, · · · , θG})(
3
2+ log N),
(2.27) In addition, Biernacki et al. proposed the ICL criterion [43],
ICL(G clusters; X ) = C2( ˆP, ˆΘG; X ) −
1
2.4.3
The DAEM algorithm
In the DAEM algorithm for learning a mixture model [23], the objective is to minimize the following system energy function during the annealing process:
Fβ(Θ; X ) = − 1 β N X i=1 log( G X k=1 (w(k)p(xi; θk))β), (2.29)
where 1/β corresponds to the temperature that controls the annealing process. The aux-iliary function in this case is
Uβ(Θ; Θ(t)) = − N X i=1 G X k=1 f (k | xi; Θ(t)) log p(xi, k; Θ), (2.30) where f (k | xi; Θ(t)) = (w(k)(t)p(x i; θ(t)k ))β PG j=1(w(j)(t)p(xi; θ(t)j ))β (2.31) is the posterior probability derived by using the maximum entropy principle.
Ueda and Nakano [23] showed that Fβ(Θ; X ) can be iteratively minimized by
it-eratively minimizing Uβ(Θ; Θ(t)). When using DAEM to learn a mixture model, β is
initialized with a small value (less then 1) such that the energy function itself is simple enough to be optimized. Then, the value of β is gradually increased to 1. During the learning process, the parameters learned in the current learning phase are used as the initial parameters of the next phase2. In the case of β = 1, F
β(Θ; X ) and Uβ(Θ; Θ(t)) are
the negatives of the log-likelihood function in Eq. (2.7) and the Q-function in Eq. (2.8),
respectively; thus, minimizing Fβ(Θ; X ) is equivalent to maximizing the log-likelihood
function.
According to [23], Eq. (2.29) can be rewritten as
Fβ(Θ; X ) = Uβ(Θ) − 1 βSβ(Θ), (2.32) where Uβ(Θ) = − N X i=1 G X k=1 f (k | xi; Θ) log p(xi, k; Θ), (2.33) and Sβ(Θ) = − N X i=1 G X k=1 f (k | xi; Θ) log f (k | xi; Θ) (2.34)
is the entropy of the posterior distribution. When β→∞, the rational function f (k | xi; Θ)
approximates to a zero-one function; thus, the entropy term Sβ(Θ)→0. In this case,
Fβ(Θ; X ) is equivalent to the negative of the objective function for CEM in Eq. (2.21).
2Each β value corresponds to a learning phase. The algorithm proceeds to the next phase after it
Therefore, DAEM can be viewed as a DA variant of CEM.
2.4.3.1 The DAEM algorithm for Gaussian mixture models
For the given β value, each iteration of the DAEM algorithm for learning GMMs is summarized as follows.
• E-step: Given the current model, Θ(t), compute the posterior probability of each mixture component of p(xi; Θ(t)) for each xi using Eq. (2.31).
• M-step: Substituting Eq. (2.12) into Eq. (2.30) and then setting its derivative to
zero, we obtain the re-estimation formulae of the parameters as follows.
w(k)(t+1)= 1 N N X i=1 f (k | xi; Θ(t)), (2.35) µ(t+1)k = PN i=1f (k | xi; Θ(t))xi PN i=1f (k | xi; Θ(t)) , (2.36) Σ(t+1)k = PN i=1f (k | xi; Θ(t))(xi− µ(t+1)k )(xi− µ(t+1)k )T PN i=1f (k | xi; Θ(t)) . (2.37)
Chapter 3
BIC-based self-splitting Gaussian
mixture learning
3.1
The SGML algorithm
Given the data set X = {x1, x2, · · · , xN}, we can partition it into G clusters after applying
the EM algorithm to learn a G-component Gaussian mixture model (GMMG). Here, to
help explain the scenario of the proposed approach, we denote a cluster obtained by EM learning a EM cluster. The self-splitting Gaussian mixture learning algorithm (SGML) starts with a single component (i.e., a single EM cluster) in the data space (i.e., X ) and splits the selected component adaptively during the learning process until the most appropriate number of components (EM clusters) is found. We describe the details of SGML in Algorithm 1. There are three main aspects with respect to the self-splitting learning rules:
I1: How to group the data samples into clusters?
I2: Which component should be split into two new components?
I3: How many components are enough for fitting the distribution of the data samples?
On issue I1: After EM, each xi ∈ X is assigned to the EM cluster whose Gaussian
component has the largest posterior probability for xi, i.e., assign xi to EM clusterj for
j = arg maxkp(k | xi; ˆΘ).
On issue I2: For each EM cluster, we calculate the value of ∆BIC21(EM cluster):
∆BIC21(EM cluster) = BIC(GMM2, EM cluster) − BIC(GMM1, EM cluster),
(3.1)
where GMM1 fits the EM cluster with a single Gaussian component while GMM2 fits
the EM cluster with a mixture of two Gaussian components. The component
because, according to the BIC theory, the larger the ∆BIC21 is, the higher the confidence
that GMM2 fits the corresponding EM cluster better than GMM1 does. The mean
vec-tor and covariance matrix of GMM1can be optimally estimated in an analytic way (which
are respectively the sample mean vector and sample covariance matrix of EM cluster),
whereas those of GMM2 need to be estimated by the EM algorithm. For the learning
of GMM2, we use K-means clustering (here, K=2) to find two initial mean vectors of
GMM2, and then apply EM to fine-tune the parameters. Suppose µ is the mean vector of
GMM1, we use µ − ε and µ + ε as the initial centroids of the K-means clustering, where
ε is a constant vector.
On issue I3: After the jth split operation, the number of components grows to j + 1
and these Gaussian components are used as the initialization of EM to learn the GMMj+1
for X ; and the value of BIC(GMMj+1, X ) is computed after the EM learning. During the
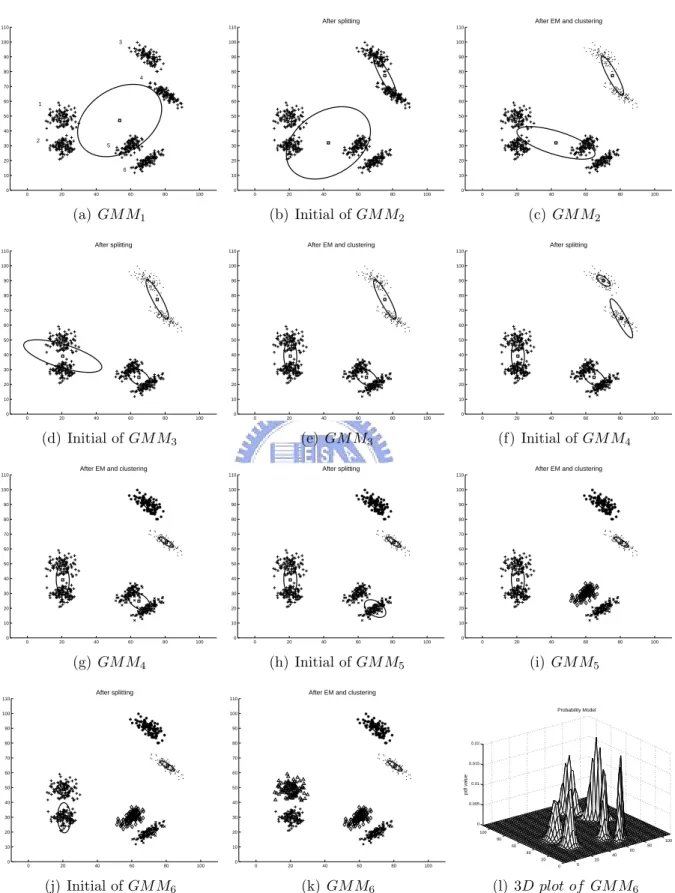
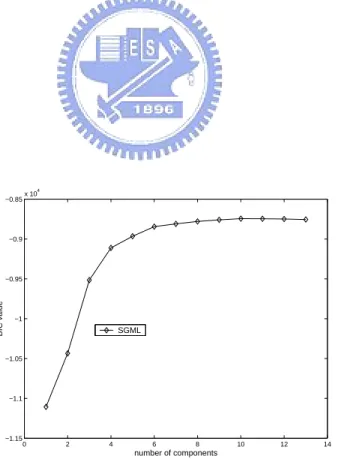
learning process the BIC values of the learned GMMs with different numbers of mixture components are collected to form a learning curve (i.e., the BIC plot); and the learning process stops when a “significant maximum” in the learning curve is found. Given that
SGML has generated {GMM1,GMM2,· · ·,GMMcN um}, if BIC(GMMcN um−SRange, X ) is
the maximum among {BIC(GMM1, X ),BIC(GMM2, X ),· · ·,BIC(GMMcN um, X )}, it is
called the “significant maximum” of the learning curve and SGML stops and returns
the model GMMcN um−SRange; otherwise, SGML continues to learn GMMcN um+1. Here,
SRange is a positive integer; it is appropriate to set SRange around 5 according to our
experience.
3.1.1
Computational complexity of SGML
Without loss of generality, we assume the splitting process of SGML stops at GMMGmax.
With the data set X , we use TKM(k, X ) to denote the computational cost of applying
K-means clustering to initialize GMMkfor EM and use TEM(k, X ) to denote the cost of using
EM for learning GMMk. For SGML, the computational cost of the Splitting step that
splits GMMM to GMMM +1 consists of the cost of computing the sample mean vectors
and sample covariance matrices for all clusters, i.e.,PM
j=1TEM(1, EM clusterj), plus the
cost of applying K-means (K=2 here) followed by EM to learn GMM2 for all clusters, i.e.,
PM
j=1[TKM(2, EM clusterj) +TEM(2, EM clusterj)].
PM
j=1TEM(1, EM clusterj) is
al-most equal to TEM(1, X ). Moreover, from Eqs. (2.13)-(2.15), it is obvious that TEM(k, X )
is proportional to the component number k and the size of data set X . In fact, TKM(k, X )
also has this property. Therefore, PM
j=1TEM(2, EM clusterj) can be approximated by
TEM(2, X ), while
PM
j=1TKM(2, EM clusterj) can be approximated by TKM(2, X ). In
of SGML is about
GXmax
k=1
[TEM(1, X ) + TKM(2, X ) + TEM(2, X ) + TEM(k, X )]. (3.2)
For the common approach that uses K-means (with random initialization) followed by EM to learn the candidate models for the model selection criterion, the computational cost is
GXmax
k=1
[TKM(k, X ) + TEM(k, X )]. (3.3)
Comparing Eq. (3.2) to Eq. (3.3), it is clear that SGML is more efficient when k À 2 because for which case, TKM(k, X ) is larger than TEM(1, X )+ TKM(2, X )+TEM(2, X ).
Comparing to the method proposed by Young and Woodland [31], SGML has an over-head of TEM(1, X ) + TKM(2, X ) + TEM(2, X ) in each splitting step. However, TEM(1, X )
is much small than 1
2TEM(2, X ) because it does not involve likelihood calculations when
estimating GMM1. Moreover, TKM(2, X ) is much smaller than TEM(2, X ). Thus, the
overhead is just slightly higher than TEM(2, X ). When M À 2, the overhead for splitting
GMMM to GMMM +1is negligible since TEM(2, X ) is much smaller than TEM(M + 1, X ).
3.2
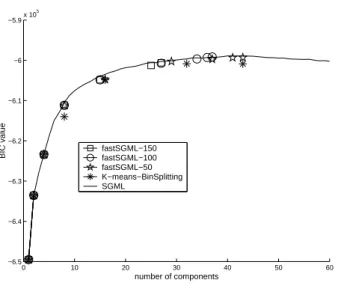
The fastSGML algorithm
The computation cost becomes an important issue when the data set is very large; a fast
learning procedure is therefore desirable. Since we can validate if GMM2 fits a cluster
better than GMM1 using BIC, a straightforward idea to speedup SGML is that split all
the clusters whose ∆BIC21values are larger than a threshold in each splitting step. Here, we call the threshold the splitting confidence. Based on the concept of multiple splitting, we proposed a fastSGML approach in Algorithm 2. To avoid the over-fitting condition caused when using a too small splitting confidence, the fastSGML stops not only when the ∆BIC21 value for each cluster is smaller than the splitting confidence (in Step 3) but also when the learning curve starts to go down (in Step 4). Clearly, fastSGML needs a much lower computation requirement than SGML because it allows multiple splits in the splitting step. fasSGML resembles the LBG algorithm [75], but the difference is that the later iteratively splits each of the clusters into two clusters until a given cluster number is reached and no validation measure is applied to the splitting.
3.3
Experiments on Gaussian mixture learning
We evaluated the proposed SGML and fastSGML algorithms on one synthetic data set which consists of six Gaussian clusters in <2 and one real-world data set which consists of speech feature vectors from speaker #5007 of the NIST 2001 cellular speaker
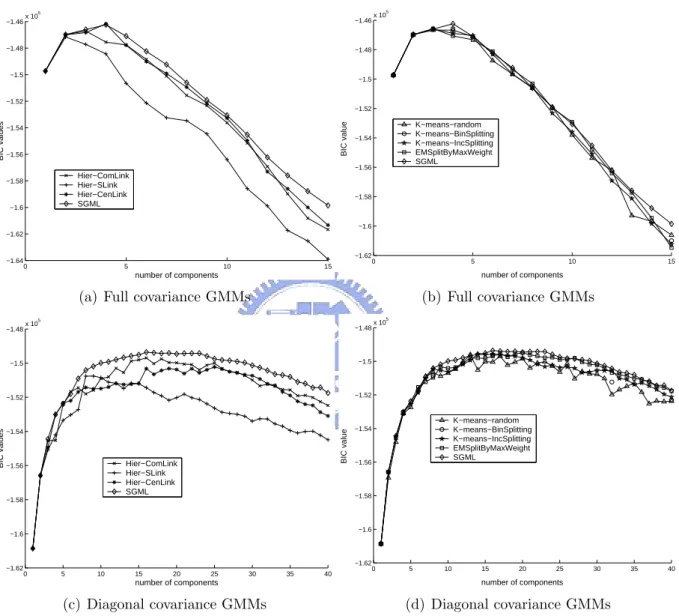
recog-nition evaluation data (NIST01SpkEval) [68]. We evaluated the algorithms using the performance of speaker GMM training because we shall apply SGML to the GMM-based speaker identification task, as shown in Section 3.4. In each task, the performance of the proposed algorithms were compared to that of other EM-based learning approaches whose initial mean vectors of GMMs are located by hierarchical agglomerative clustering (HAC) or K-means clustering. The baseline approaches are as follows.
1. Hier-ComLink method: The initial Gaussian mean vectors in EM were obtained by the complete-linkage HAC [35].
2. Hier-SLink method: The initial Gaussian mean vectors in EM were obtained by the single-linkage HAC [35].
3. Hier-CenLink method: The initial Gaussian mean vectors in EM were obtained by the centroid-linkage HAC [35].
4. K-means-random method: The initial Gaussian mean vectors in EM were obtained by K-means clustering, in which the initial centroids are randomly selected from the data set.
5. K-means-BinSplitting method [75]: The initial Gaussian mean vectors in EM were determined by the LBG algorithm, in which each mean vector was split into two new ones in each splitting step until the desired number of clusters was reached. 6. K-means-IncSplitting method [76]: The initial Gaussian mean vectors in EM were
obtained by the incremental splitting K-means algorithm, in which only the mean vector of the cluster with the largest accumulated distance was split into two new ones in each splitting step until the desired number of clusters was reached.
7. EMSplitByMaxWeight method [31]: This method splits the mean vector of the Gaussian component with the largest mixture weight into two new ones in each splitting step, and then performs EM to update all Gaussian components. The number of mixture component is incremented from one to the pre-defined number.
For each baseline approach, the initial covariance matrix of GMMk was set as ρkI,
where ρk=minj6=k{kµ(0)j − µ(0)k k} and µ(0)j denotes the initial mean vector of the jth
Gaussian component.
3.3.1
Results on the synthetic data
Figure 3.1 (a) shows the synthetic data that contains six Gaussian clusters, each with 100 samples. First, we conducted experiments using full covariance Gaussian components. Figure 3.1 schematically depicts the learning process of the SGML algorithm. For this