Cloning, Expression, and Characterization of Two
β-Glucosidases from Isoflavone
Glycoside-Hydrolyzing Bacillus subtilis natto
L
UN-C
HENGK
UO ANDK
UNG-T
AL
EE*
Institute of Microbiology and Biochemistry, National Taiwan University, No. 1, Sec. 4, Roosevelt Road, Taipei, Taiwan, Republic of China
On the basis of the genomic sequence of Bacillus subtilis 168, two β-glucosidase genes (bglH and yckE) from B. subtilis natto, which has been reported to have high isoflavone glucoside-hydrolyzing activity, were cloned and overexpressed in E. coli M15. The temperature for the optimal p-nitrophenyl-β-D-glucoside hydrolyzing activity of both enzymes was between 37 and 45°C, but BglH had a higher thermal stability than YckE. Both showed high activity at pH 6.0, but YckE was stable over a wider pH range than BglH. Recombinant BglH was inhibited 73%, 63%, and 43% by 1.0 mM Cd2+, Fe2+, or Cu2+, respectively, while other divalent metal ions resulted in 0–23% inhibition, whereas YckE was inhibited by less than 20% by any of the divalent metal ions we tested. Among the substrate we used, BglH showed the highest affinity for genistin and YckE showed the highest affinity for p-nitrophenyl-β-D-fructopyranoside. Both BglH and YckE hydrolyzed genistin and daidzin into their isoflavone aglycones, genistein and daidzein, but BglH was more efficient than YckE in isoflavone glucoside hydrolysis (20-fold higher kcat). Our results suggest that recombinant BglH may be applicable
in the process of isoflavones deglycosylation.
KEYWORDS: Bacillus subtilis natto; isoflavone;β-glucosidases; BglH; YckE INTRODUCTION
Among the foods eaten by humans, soybeans contain the
highest concentration of isoflavones. These soy isoflavones (e.g.,
daidzein and genistein) are implicated in some health-enhancing
properties, such as prevention of certain cancers (1–3), lowering
the risk of cardiovascular diseases (4, 5), and improvement in
bone health (6, 7). In vitro studies of metabolites produced from
the total microflora of volunteers have shown that the
bioavail-ability of soybean isoflavonoids depends upon the bioavail-ability of the
gut microflora to metabolize these compounds (8, 9).
Numerous studies have shown that the biological effects of
isoflavones are not due to the glycoside form, but mainly to
their aglycones, such as daidzein and genistein (10, 11).
Aglycone isoflavones are highly bioactive due to their
unim-peded intestinal absorption, unlike their related glycosides, which
are not absorbed across enterocytes because of their higher
hydrophilicity and molecular weights (12–14). Isoflavone
ag-lycones are present in high amounts in soy products, such as
miso, natto, and tempeh, produced, respectively, by Aspergillus
oryzae (15), Bacillus subtilis natto (16, 17), and Rhizopus
oligosporus (18). Bifidobacteria
β-glucosidase can convert
isoflavone glucosides in soymilk to their aglycones (19), and
β-glucosidase from Saccharopolyspora erythraea can hydrolyze
genistin during fermentation of soy-based media (20). These
bacteria with
β-glucosidase activity are potentially important
in the production of compounds with higher estrogenicity and
better absorption, thus affecting their bioavailability and
phar-macokinetics.
β-Glucosidase (β-glucoside glucohydrolase, EC 3.2.1.21) is
the key enzyme for carbohydrate metabolism in bacteria that
are able to cleave the
β-glucosidic linkages of di- and/or
oligosaccharides or other glucose conjugates (21).
β-Glucosi-dases are widely distributed in living organisms and play pivotal
roles in many biological processes, such as the degradation of
cellulosic biomass (22), cyanogenesis (23), and the cleavage
of glucosylated flavonoids (24).
According to previous reports, the isoflavone
glycoside-hydrolyzing
β-glucosidases of B. subtilis natto (16, 25) or lactic
acid bacteria (26, 27) are cell-associated and difficult to purify;
as a result, there have been no reports of the biochemical
properties of these enzymes. In our previous study (25), a strain
of B. subtilis natto NTU-18 with high isoflavone
glycoside-hydrolyzing
β-glucosidase activity was selected from
com-mercial natto product (a traditional soy fermentation product
in Japan and eastern Asia), and a fermentation process for
deglucosylation of black soybean isoflavones was established.
In the present study, two
β-glucosidase genes (bglH and yckE)
were cloned from B. subtilis natto NTU-18 and the
character-istics of the recombinant enzymes expressed in E. coli were
investigated.
* Corresponding author. E-mail: [email protected]. Phone: +886-2-33664436. Fax: +886-2-23640961.
10.1021/jf072287q CCC: $40.75 2008 American Chemical Society Published on Web 12/11/2007
MATERIALS AND METHODS
Chemicals and Reagents. Genistein, daidzein,
genistein-7-O-glucoside (genistin), daidzein-7-O-genistein-7-O-glucoside (daidzin), p-nitrophenol,
p-nitrophenyl-β-D-glucoside (pNPG), o-nitrophenyl-β-D
-galactopyra-noside (pNPGal), p-nitrophenyl-β-D-fructopyranoside (pNPF), and
p-nitrophenyl-β-D-cellobioside were purchased from Sigma Chemical
Co. (St. Louis, MO). Liquid chromatography grade acetonitrile and reagent grade absolute alcohol were purchased from Merck (Darmstadt, Germany). Nutrient broth (NB) and Bacto-agar were purchased from Difco (Detroit, MI). Isopropyl-β-D-thiogalactopyranoside (IPTG) was
purchased from MDBio (Taipei, Taiwan).
Bacterial Strains, Plasmids, and Media. A strain of B. subtilis
natto NTU-18 with high β-glucosidase activity (with pNPG as the
substrate) was previously obtained from commercial natto product (25). The strain was maintained on nutrient broth (NB) (Difco, Detroit, MI)
slants at 4 °C. Escherichia coli JM109 (Yeastern Biotech, Taiwan) containing the pGEM-T Easy vector (Promega, USA) was used for plasmid preparation and gene cloning, and the E. coli M15 strain (QIAGEN, Hilden, Germany) containing the pQE-30Xa vector (QIAGEN, Hilden, Germany) was used to express 6× His-tagged recombinant proteins. All E. coli cells containing plasmids were grown aerobically in Luria–Bertani (LB) medium (DIFCO, USA) or on LB agar plates at 37°C, supplemented with ampicillin (100µg/mL) and (or) kanamycin
(25µg/mL) when necessary.
Construction of the Expression System. Genomic DNA of B. subtilis
natto NTU-18 was isolated according to Sambrook et al. (28) and used as the polymerase chain reaction (PCR) template. To amplify the DNA fragment encoding bglH, PCR was performed using the forward primer
bglH-F (5
′GCAGGATCCATGAGTTCAAATGAAAAACGATTTCC-AGAAGG) and reverse primer bglH-R (5′CTACTGCAGTCAGA-GACTCTCTCCGTTTGTGGCG), which introduce BamHI and PstI sites, respectively, at the 5′and 3′-termini of the bglH gene. To amplify the DNA fragment encoding yckE, PCR was performed using the forward primer yckE-F (5′CTGAGGCCTATGATCCACCAGCATCCAGAATC) and reverse primer yckE-R (5′CTAGGATCCTTATAAACTTTCTC-CGTTTCTCTTG), which introduce StuI and BamHI sites, respectively, at the 5′and 3′-termini of the yckE gene (the enzyme sites are shown in bold). Amplification was carried out in a thermo Px2 thermal cycler machine using the conditions of 5 min of denaturation at 95°C, 30 cycles for 1 min at 94°C and 1 min at 55°C, and a final extension phase at 72 °C for 1 min and 40 s. The PCR products were ligated into the pGEM-T Easy Vector System (Promega, USA), the ligation products were trans-formed into E. coli strain JM109, and the recovered plasmids were confirmed as correct by restriction analysis and DNA sequencing. The
Figure 1. Putative catalytic regions of BglH and YckE. The arrows indicate the conserved catalytic residues.
Table 1. Purification of Recombinant BglH and YckE Expressed in E. coli
M15a name purification step total protein (mg) total activity (U) specific activity (U/mg) purification (fold) yield (%) BglH crude extractb 358.0 623.0 1.74 1.0 100.0 His-binding columnc 170.8 450.8 2.64 1.5 72.4 YckE crude extractb 421.4 573.2 1.36 1.0 100.0
His-binding columnc
168.3 333.0 1.98 1.5 58.1
aOne unit of enzyme activity was defined as the amount of enzyme which released 1 µmol of p-nitrophenol per minute.bThe recombinant strain was grown in LB medium with 25 kanamycin µg/mL and 100 amplicillin µg/mL at 37°C to OD6001.0 and was incubated further with IPTG at a final concentration of 0.05 mM for 12 h at 25°C. The cells of 700 mL cultures were harvested by centrifugation at 10 000g for 10 min at 4°C, resuspended in 50 mL lysis buffer (50 mM NaH2PO4, 300 mM NaCl, 10 mM imidazole, pH 8.0), and then disrupted by sonication.cThe resulting supernatants were loaded onto an Ni-NTA affinity column equilibrated with the binding buffer (50 mM NaH2PO4, 300 mM NaCl, 20 mM imidazole, pH 8.0), and the bound proteins were eluted with elution buffer (50 mM NaH2PO4, 300 mM NaCl, 250 mM imidazole, pH 8.0). The fractions exhibiting enzyme activity were pooled and dialyzed against 100 mM sodium phosphate buffer, pH 6.0.
Figure 2. SDS-PAGE analysis of purified recombinant BglH and YckE: BglH (1 or 2 µg; lanes 2 and 3), YckE (1 or 2 µg; lanes 4 and 5), and molecular markers (lane 1) stained with Coomassie Blue.
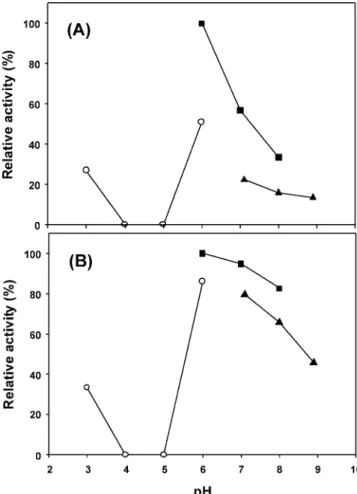
Figure 3. Effect of pH on the activity of recombinant BglH and YckE: (A) BglH, (B) YckE. The buffers used were the following: pH 3.0–6.0, 100 mM citric acid-citrate buffer (O); pH 6.0–8.0, 100 mM sodium phosphate buffer (9); pH 7.0–9.0, 50 mM Tris-HCl buffer (2). The optimal pH was determined at 37°C for 10 min in the 100 mM citric acid-citrate buffer at pH 3.0–6.0, 100 mM sodium phosphate buffer at pH 6.0–8.0, or Tris-HCl buffer at pH 7.0–9.0 with pNPG as a substrate.
nucleotide and amino acid sequences were compared to the sequences in the GeneBank at the National Center for Biotechnology Information (Bethesda, MD) using the BLAST network server. Plasmid DNA in the clones was extracted using a QIAprep Spin Miniprep Kit (Qiagen, Hilden, Germany), digested appropriately, and ligated into pQE30-Xa (Qiagen, Hilden, Germany) which provides high-level expression in E. coli of proteins containing a 6× His affinity tag at the N-terminus of the protein; then, the ligation products were transformed into E. coli strain M15, and the recovered plasmids were confirmed as correct by restriction analysis and DNA sequencing. The resulting plasmids were named pQE-30Xa-BglH and pQE-30Xa-YckE, respectively. DNA manipulations, including digestion with restriction enzymes, purification of DNA fragments, ligation with T4 ligase, and transformation, were performed as described by Sambrook et al. (28).
Expression and Purification of the Recombinantβ-Glucosidases. E. coli M15 harboring the recombinant plasmid pQE-30Xa-BglH or
pQE-30Xa-YckE were grown to an A600 of 1.0 in LB medium supplemented with 25µg/mL kanamycin and 100 µg/mL ampicillin at
37°C, then induced to produce a target protein by adding IPTG at a final concentration of 0.05 mM, followed by incubation for 12 h at 25 °C. All subsequent steps were at 4°C. The cells were harvested by centrifugation, washed twice with lysis buffer (50 mM NaH2PO4, 300 mM NaCl, 10 mM imidazole, pH 8.0), and the cell pellet resuspended in the same buffer and lysed by sonication. The cell lysate was centrifuged at 10 000g for 20 min to remove cell debris, and the supernatant was applied to an Ni-NTA affinity column (Qiagen, Hilden, Germany) equilibrated with 1× binding buffer (50 mM NaH2PO4, 300 mM NaCl, 20 mM imidazole, pH 8.0). The recombinant proteins were eluted with Elution buffer (50 mM NaH2PO4, 300 mM NaCl, 250 mM imidazole, pH 8.0). Fractions containing enzyme activity were pooled and dialyzed against 100 mM sodium phosphate buffer, pH 6.0, and the enzyme preparation was stored at -80°C. The protein concentration was determined using a Bio-Rad protein assay kit, with bovine serum albumin as the standard.
Electrophoresis Analyses and Protein Assay. The apparent
mo-lecular weights of the purified enzymes were determined using 12.5% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). Proteins were stained with Coomassie Brilliant Blue R-250.
Enzyme Assay. Theβ-glucosidase assay was modified from the
work of Choi et al. (26). The activity of the purifiedβ-glucosidases
was estimated by mixing 50µL of enzyme with 50 µL of 5 mM pNPG
in 100 mM sodium phosphate buffer, pH 6.0, incubating the mixture for 10 min at 37°C, adding 100µL of 2 M Na2CO3to stop the reaction, measuring the absorbance at 405 nm, and comparing it to a standard curve prepared by measuring the A405 of various concentrations of
p-nitrophenol. One unit of enzyme activity was defined as the amount
of enzyme that liberates 1µmol of p-nitrophenol per minute.
times and, then, assaying the remaining activity under the standard conditions.
Effect of pH on Enzyme Activity. To determine the optimum pH
for the enzyme, activities were measured over a pH range of 3.0–9.0 in increments of 1.0 pH unit using the standard assay conditions. The buffers used were a 100 mM citric acid-citrate buffer (pH 3.0–6.0), 100 mM sodium phosphate buffer (pH 6.0–8.0), and 50 mM Tris-HCl buffer (pH 7.0–9.0).
Substrate Specificity and Kinetic Studies. Substrate specificity was
determined by incubating the enzyme at 37°C for 10 min in 100 mM sodium phosphate buffer, pH 6.0, containing a 0–2.5 mM final concentration of different nitrophenyl substrates or isoflavone glucosides and the Michaelis–Menten kinetic parameters for the purified enzyme determined from substrate saturation assays. The substrates used were
pNPG, pNPGal, pNPF, p-nitrophenyl-β-D-cellobioside, daidzin, and genistin. The values for the maximum velocity and half-saturation coefficient (Km) were determined by plotting the substrate concentration vs the initial velocity for each reaction and subjecting the data to nonlinear regression analysis. Kinetic analyses by curve fitting were performed using SigmaPlot software. Values shown are the mean of duplicate experiments with each substrate, and the variation about the mean was below 5%.
HPLC Analyses. HPLC analysis of isoflavones was based on the
work of Wang and Murphy (29). The HPLC system consisted of a Shimadzu SCL-10Avp system controller, two Shimadzu LC-10ATvp liquid chromatograph pumps, and a Shimadzu SPD-M10Avp photo-diode array detector. A Develosil ODS-5 column (250 mm with 4.6 mm packing) was used. The solutions used to generate the gradient were 0.1% (v/v) glacial acetic acid in H2O (solvent A) and 80% (v/v) acetonitrile (Merck, Darmstadt, Germany) in solvent A (solvent B). Following injection of 20µL of sample, solvent B was increased linearly
from 15% to 70% over 30 min and, then, returned to 15% for 10 min The solvent flow rate was 1.0 mL/min. Elution was monitored by UV absorbance at 262 nm, and the spectral data from 190 to 800 nm of the peaks were recorded to confirm their identity. Peak areas were integrated for quantification. Daidzin, genistin, daidzein, and genistein were identified by comparison with the HPLC retention time and UV spectra of the authentic compounds. The isoflavone concentration of the samples was calculated using calibration curves prepared using various concentrations of the isoflavone standards.
Statistical Analysis. The data of the significantly altered expression
in each enzyme assay mean value (from triplicate experiments) with the corresponding mean value of the control and the test expression change using student’s t test. P values < 0.05 were considered statistically significant.
RESULTS
Gene Cloning of Two
β-Glucosidases from B. subtilis
natto. The B. subtilis natto NTU-18 strain was chosen for this
β-glucosidase gene cloning study because it efficiently
hydro-lyzes isoflavone glucosides into isoflavone aglycones (25), and
this reaction is known to be catalyzed by
β-glycosidase (16).
Two
β-glycosidase genes (bglH and yckE) in B. subtilis 168
were identified by searching the NCBI genome database and
were amplified from the genome of B. subtilis natto NTU-18
by PCR (30). The sizes of these two genes were 1410 base
pairs (bglH) and 1434 base pairs (yckE). The BglH contained
470 amino acids, with a calculated molecular weight of 53 kDa.
And, the YckE contained 478 amino acids, with a calculated
molecular weight of 54 kDa. Alignment of the two nucleotide
sequences with the NCBI database, the nucleotide sequence of
bglH showed 98% and 78% homology with the gene of B.
subtilis 168 and B. licheniformis, respectively, and the nucleotide
sequence of yckE showed 98% and 78% homology with the
Figure 4. Effect of temperature on the activity of recombinant BglH and YckE: BglH (b), YckE (O). The optimal temperature for the enzyme activity was determined by assay at 20-50°C in 100 mM sodium phosphate buffer, pH 6.0, with pNPG as a substrate.
gene of B. subtilis 168 and B. amyloliquefaciens, respectively.
The deduced amino acid sequence of BglH and YckE were used
to compare with other amino acid sequences deposed in the
NCBI database. The BglH exhibits the highest identity score
with B. subtilis 168 BglH (99% identity), followed by B.
licheniformis BglH (87%) and B. amyloliquefaciens BglH
(85%). In the case of YckE, the highest score was also obtained
with B. subtilis 168 YckE (98% identity), followed by B.
licheniformis YckE (86%) and B. amyloliquefaciens YckE
(84%). By comparison of bglH and yckE in B. subtilis natto
NTU-18, it showed 56.2% similarity in nucleotide sequence and
43.7% identical in amino acid sequences. For the found catalytic
domain of BglH and YckE, we using the SIM-Alignment tool
to compare the amino acid sequence of the putative catalytic
domain with other
β-glucosidases. BglH and YckE were found
to contain histidine and glutamate residues in the catalytic
domain (Figure 1).
Overexpression and Purification of the Two
Recombi-nant
β-Glucosidases. To investigate the biochemical
proper-ties of BglH and YckE, we expressed the six-histidine N
terminal-tagged proteins in E. coli M15. Bacteria were
transformed with the expression vector and induced with
IPTG which expressed the histidine-tagged protein. After
treatment of the His-binding column, recombinant BglH was
purified 1.5-fold with a 72.4% yield and YckE was purified
1.5-fold with a 58.1% yield (Table 1). The apparent
molecular weights, determined by SDS-PAGE (12.5% gel),
were about 57 and 58 kDa, respectively (Figure 2). Using
the pQE30-Xa expression system, the N-terminal of the
expression protein contains a six-histidine tag and a factor
Xa recognition site, resulting in a 4 kDa increase in the
molecular weight of the expressed protein. Thus, the actual
molecular weight of BglH was 53 kDa and that of YckE was
54 kDa, corresponding to the expected molecular weights.
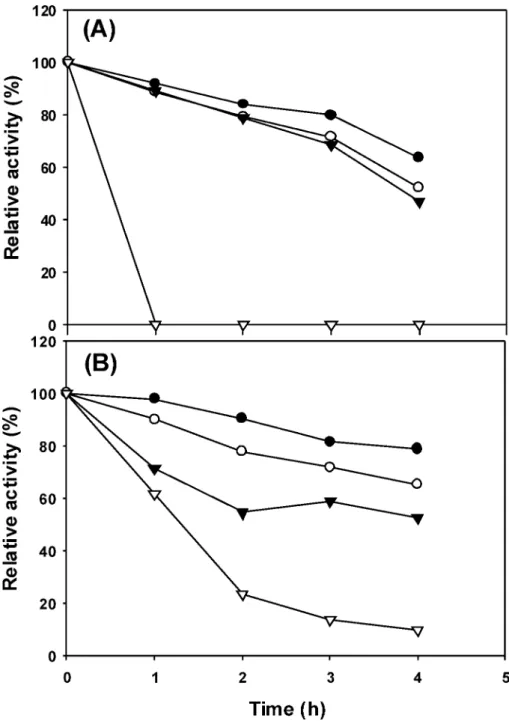
Figure 5. Thermal stability of recombinant BglH and YckE: (A) BglH, (B) YckE. The purified enzyme in 100 mM sodium phosphate buffer, pH 6.0, was preincubated for various times at 30 (b), 37 (O), 45 (1), or 50°C (3) in the absence of substrate and, then, was assayed under standard conditions with pNPG as a substrate. The activity of the enzyme without preincubation is taken as 100%.
Biochemical Characterization of the Two Recombinant
β-Glucosidases. To determine the optimal pH for recombinant
BglH and YckE, we measured the enzyme activity at 37
°
C
and various pH values (pH 3.0–9.0), using pNPG as substrate.
As shown in Figure 3, both enzymes showed the highest activity
at pH 6.0, but YckE was active over a wider pH range.
Meanwhile, precipitation was observed for both the recombinant
BglH and YckE treated at pH 4 and 5, and no enzyme activity
was detected in this pH range. To determine the optimal
temperature, enzyme activity was determined over the range of
20–50
°
C. As shown in Figure 4, in the temperature range of
35–45
°
C, recombinant BglH exhibited more than 95% of its
maximal activity, with a sharp decrease above 45
°
C, while
YckE exhibited about 70–100% of its maximal activity over
the same range. To examine the thermal stability of recombinant
BglH and YckE, we incubated the enzymes at different
temperatures (30, 37, 45, and 50
°
C) and measured the residual
activity under standard assay conditions. As shown in Figure
5, recombinant BglH and YckE were fairly stable up to 45
°
C,
but the thermal stability profiles of the two recombinant enzymes
were very different. Recombinant BglH retained 80% of its
activity when incubated at 45
°
C for 2 h, but was completely
inactivated by incubation at 50
°
C for 1 h, whereas recombinant
YckE retained 60% of its activity when incubated at 45
°
C for
2 h or at 50
°
C for 1 h.
The effects of different metal ions (Na
+, Li
+, Fe
2+, Mn
2+,
Mg
2+, Cu
2+, Cd
2+, Ca
2+, and Fe
3+) on enzyme activity were
investigated by addition of the test ions to the reaction mixture
at a final concentration of 1 mM. The activity was then measured
under standard assay conditions and expressed as a percentage
of the activity obtained in the absence of the added ion. As
shown in Table 2, recombinant BglH was inhibited 73%, 63%,
and 43% by 1.0 mM Cd
2+, Fe
2+, or Cu
2+, respectively, while
other divalent metal ions resulted in 0–23% inhibition, whereas
YckE was inhibited by less than 20% by any of the divalent
metal ions.
Kinetic Properties of the Two Recombinant
β-Glucosi-dases. To determine the kinetic properties, the enzyme activity
of the two recombinant
β-glucosidases was assayed by
monitor-ing the hydrolysis of
β-glucosides over different substrates and
a range of concentrations of these substrates. The substrate
specificities of the two recombinant
β-glucosidases are
sum-marized in Table 3. Recombinant BglH showed a lower
Michaelis constant (K
m) and higher k
catfor daidzin and genistin
than for the other substrates. Comparison of the K
mand k
catKm
) of the enzyme for different substrates was considered a
measurement of the enzyme’s specificity (specificity constant).
The catalytic efficiency (k
cat/K
m) values of recombinant BglH
for genistin, daidzin, p-NPG, p-NPGal, and p-NPF were 147.20,
49.19, 3.01, 1.76, and 0.90 1/(mM s), respectively. Among the
substrates we tested, recombinant BglH showed the highest
catalytic efficiency toward genistin. These results indicated that
genistin was clearly the preferred substrate. The catalytic
efficiency (k
cat/K
m) values of recombinant YckE for genistin,
daidzin, p-NPG, p-NPGal, and p-NPF were 19.45, 17.93, 2.44,
1.23 and 306.57 1/(mM s), respectively. The preferred substrate
for recombinant YckE was pNPF, the catalytic efficiency for
pNPF being about 150 times that for pNPG and 16 times that
for genistin. Recombinant YckE also had isoflavone glucoside
deglycosylation activity, but its main activity was against pNPF.
To compare these two recombinant
β-glucosidases, YckE had
lower K
mvalues for daidzin and genistin, but BglH’s k
catvalues
for daidzin and genistin were 20 times higher; the catalytic
efficiency of recombinant BglH for genistin was about 7.7 times
higher than that of YckE. BglH therefore hydrolyzed soybean
isoflavone glucosides more efficiently than YckE.
DISCUSSION
Two
β-glucosidase genes (bglH and yckE) were cloned from
B. subtilis natto NTU-18 and expressed in an E. coli system
for enzyme characterization.
β-Glucosidases BglH and YckE
of B. subtilis natto showed high nucleotide sequence similarity
and amino acid sequence identity with the relative genes of B.
subtilis 168.
β-Glucosidases belong to the cellulose family BG
and form two subfamilies, BGA and BGB (31). The BGA
subfamily contains mainly bacterial
β-glucosidases with
mo-lecular weights of about 50 kDa. BGA subfamily
β-glucosidases
have a putative catalytic domain containing histidine and
glutamate residues (32, 33) which has been suggested to be
involved in the cleavage of
β-glucosidic bonds. The sequence
homology with the
β-glucosidases of the BGA subfamily (31–34)
seen using the SIM-Alignment tool suggests that BglH and
YckE of B. subtilis natto should be classified as members of
the BGA subfamily. Activity characterization revealed that the
optimal pH values for recombinant BglH and YckE are similar
to BglA
β-glucosidase, an enzyme discovered in Bacillus sp.
GL1, which acts on the gellan degradation, was classified as
belonging to the BGA subfamily, and has its highest activity at
pH 6.0 (34). This pH range is not unexpected, as most
β-glucosidases from bacterial sources show a pH optimum in
slightly acidic or neutral pH ranges (35). The recombinant BglH
enzyme showed a significant difference in activity between citric
acid-citrate buffer and sodium phosphate buffer at pH 6.0 and
between sodium phosphate buffer and Tris-HCl buffer at pH
7.0. This phenomenon is similar to the result of the intracellular
β-glucosidase of B. circulan subsp. Alkalophilus which showed
that the buffers affected strongly the observed activity and the
optimal pH was 6.0 in phosphate buffer (36). Inhibition of
enzyme activity by Tris has been observed for several
glucosi-dases and has been though to be attributed to changes in
conformation or charge distribution (37). In addition, the
enzymes were completely inactive at pH 4 and 5, while, at pH
3, both enzymes retained 35–40% activity (Figure 3). This
means that the
β-glucosidase activity of the recombinant
control 100 100 FeSO4 36.8 90.5 NaF 76.8 93.4 MnCl2 99.5 85.6 LiCl 76.8 92.7 MgCl2 84.3 95.5 FeCl3 100 81.9 CuCl2 56.8 84.1 CdCl2 26.6 80.7 CaCl2 77.9 92.6
aThe compound was added into the reaction mixture at the final concentration of 1 mM. The purified enzymes were incubated with each reagent for 10 min before the addition of the pNPG substrate solution to initiate the enzyme reaction. The reaction was determined at 37°C for 10 min in 100 mM sodium phosphate buffer, pH 6.0. The activity without the added reagent was taken as 100%.
enzymes of BglH and YckE were lost between pH 4 and 5. We
used the ExPASy Proteomics tool to estimate the pI value of
the two enzymes and found that the theoretical pI values of the
two enzymes are both about pH 4.5. This suggests that this
phenomenon was caused by conformation change of the two
enzymes in this pH range. Both recombinant BglH and YckE
showed that the activities were inhibited by divalent metal ions.
Similarly, Hashimoto et al. (34) reported that divalent metal
ions were 15–20% inhibited on BglA
β-glucosidase from
Bacillus sp. GL1.
In B. subtilis, the real function of YckE is still unknown.
Meanwhile, the bglH gene has been tentatively identified as a
β-glucosidase, but the activity of the corresponding gene product
has not been directly demonstrated (38–40). In this study, the
recombinant BglH showed the higher specificity constant for
the isoflavone conjugates than that for the chromogenic substrate
p-NPG. Similarly, the
β-glucosidases purified from
Pseudomo-nas ZD-8 (41), soybean [Glycine max] roots (42), and Cicer
arietinum L (43) also showed a higher specificity constant for
isoflavone conjugates over the generic chromogenic substrate,
p-NPG. Setlow et al. (44) reported that, in B. subtilis, bglH is
induced by
β-glucosides and expressed during the
late-exponential or stationary phase, while yckE is expressed at a
low and constant level during growth, sporulation, and spore
germination and is not induced by
β-glucosides. In our previous
study (25), the isoflavone glycoside-hydrolyzing
β-glucosidases
of B. subtilis natto were induced by isoflavone glycosides and
expressed during the late-exponential phase in black soybean
medium. According to these results, BglH may play a more
important role than YckE in the deglycosylation of isoflavone
glucosides in B. subtilis natto during fermentation.
In addition to being applied in cellulose degradation,
β-glu-cosidases could also be used to hydrolyze the phenolic
compounds (e.g., phloridzin, arbutin, and salicin) and
phy-toestrogen glucosides to improve their biological activity, with
several uses in the field of medicine, in general biomedical
research, and in the food industry (45). For example, the
hydrolysis of phloridzin by
β-glucosidase could liberate the
aglycone moiety, which is a precursor of melanin, and the
melanin is known to have the functions of reducing the risk of
skin cancer and promoting a dark color of hair (46). Similarly,
the deglycosylation of oleuropein by
β-glucosidase could release
a pharmacologically active compound hydroxytyrosol, which
is used in the prevention of coronary heart disease and cancer
(47). In order to effectively prepare the valuable
drug-materials-free aglycones, it is necessary to isolate and screen new
β-glucosidase producing microorganisms (20). In this study, we
revealed the enzyme properties of recombinant BglH and YckE.
Our data show the possibility that these
β-glucosidases could
be applied in the formation of aglycone of bioactive compounds.
These aglycones then might alter the type of sugar attached to
them via enzymes such as glycosyltrasferases, to change their
bioactivity.
LITERATURE CITED
(1) Cappelletti, V.; Fioravanti, L.; Miodini, P.; Di Fronzo, G. Genistein blocks breast cancer cells in the G(2)M phase of the cell cycle.
J. Cell Biochem. 2000, 14, 594–600.
(2) Miura, T.; Yuan, L.; Sun, B.; Fujii, H.; Yoshida, M.; Wakame, K.; Kosuna, K. Isoflavone aglycon produced by culture of soybean extracts with basidiomycetes and its anti-angiogenic activity.
Biosci. Biotechnol. Biochem. 2002, 66, 2626–2631.
(3) Ravindranath, M. H.; Muthugounder, S.; Presser, N.; Viswanathan, S. Anticancer therapeutic potential of soy isoflavone, genistein.
AdV. Exp. Med. Biol. 2004, 546, 121–165.
(4) Anthony, M. S.; Clarkson, T. B.; Hughes, C. L.; Morgan, T. M.; Burke, G. L. Soybean isoflavones improve cardiovascular risk factors without affecting the reproductive system of peripubertal rhesus monkeys. J. Nutr. 1996, 126, 43–50.
(5) Goodman-Gruen, D.; Kritz-Silverstein, D. Usual dietary isoflavone intake is associated with cardiovascular disease risk factors in postmenopausal women. J. Nutr. 2001, 131, 1202–1206. (6) Cotter, A.; Cashman, K. D. Genistein appears to prevent early
postmenopausal bone loss as effectively as hormone replacement therapy. Nutr. ReV. 2003, 61, 346–351.
(7) Weaver, C. M.; Cheong, J. M. Soy isoflavones and bone health: the relationship is still unclear. J. Nutr. 2005, 135, 1243–1247. (8) Joannou, G. E.; Kelly, G. E.; Reeder, A. Y.; Waring, M.; Nelson, C. A urinary profile study of dietary phytoestrogens. The identification and mode of metabolism of new isoflavonoids. J.
Steroid Biochem. Mol. Biol. 1995, 54, 167–184.
(9) Kelly, G. E.; Joannou, G. E.; Reeder, A. Y.; Nelson, C.; Waring, M. A. The variable metabolic response to dietary isoflavones in humans. Proc. Soc. Exp. Biol. Med. 1995, 208, 40–43. (10) Kawakami, Y.; Tsurugasaki, W.; Nakamura, S.; Osada, K.
Comparison of regulative functions between dietary soy isofla-vones aglycone and glucoside on lipid metabolism in rats fed cholesterol. J. Nutr. Biochem. 2005, 16, 205–212.
(11) Hendrich, S. Bioavailability of isoflavones. J. Chromatogr., B
Analyt. Technol. Biomed. Life Sci. 2002, 777, 203–210.
(12) Setchell, K. D.; Brown, N. M.; Zimmer-Nechemias, L.; Brashear, W. T.; Wolfe, B. E.; Kirschner, A. S.; Heubi, J. E. Evidence for lack of absorption of soy isoflavone glycosides in humans, supporting the crucial role of intestinal metabolism for bioavail-ability. Am. J. Clin. Nutr. 2002, 76, 447–453.
(13) Brown, J. P. Hydrolysis of glycosides and esters. In Role of the
gut flora in toxicity and cancer; Rowland, I. R., Ed.; Academic:
San Diego, 1998; pp 109–144.
(14) Xu, X.; Harris, K. S.; Wang, H. J.; Murphy, P. A.; Hendrich, S. Bioavailability of soybean isoflavones depends upon gut micro-flora in women. J. Nutr. 1995, 125, 2307–2315.
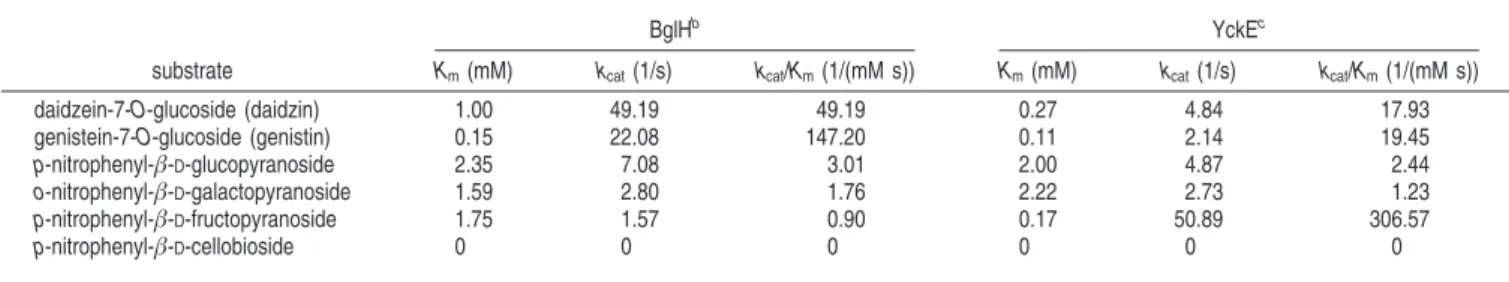
Table 3. Kinetic Properties of Recombinant BglH and YckEa
BglHb YckEc
substrate Km(mM) kcat(1/s) kcat/Km(1/(mM s)) Km(mM) kcat(1/s) kcat/Km(1/(mM s))
daidzein-7-O-glucoside (daidzin) 1.00 49.19 49.19 0.27 4.84 17.93 genistein-7-O-glucoside (genistin) 0.15 22.08 147.20 0.11 2.14 19.45 p-nitrophenyl-β-D-glucopyranoside 2.35 7.08 3.01 2.00 4.87 2.44 o-nitrophenyl-β-D-galactopyranoside 1.59 2.80 1.76 2.22 2.73 1.23 p-nitrophenyl-β-D-fructopyranoside 1.75 1.57 0.90 0.17 50.89 306.57 p-nitrophenyl-β-D-cellobioside 0 0 0 0 0 0
aValues shown are the mean of duplicate experiments with each substrate, and the variation about the mean was below 5%.bThe purified recombinant BglH (1 mg/mL; 50 µL) was incubated with 50 µL of different nitrophenyl substrates (0–5 mM) at 37°C for 10 min or isoflavone glucosides (0–5 mM) at 37°C for 30 min in 100 mM sodium phosphate buffer, pH 6.0. To stop the reaction, to the mixture of enzyme and nitrophenyl substrates was added 100 µL of 2 M Na2CO3buffer. Finally, the absorbance was measured at 405 nm and analyzed by HPLC after adding 100 µL EtOH.cThe reaction condition of purified recombinant YckE with the above substrates other than pNPF was the same as BglH, and it took only 2 min for YckE to react with it.
which contains a high level of isoflavone aglycons. Nippon
Shofuhin Kagaku Kogaku Kaishi 2001, 48, 27–34.
(17) Toda, T.; Uesugi, T.; Hirai, K.; Nukaya, H.; Tauji, K.; Ishida, H. New 6-O-acyl isoflavone glycosides from soybeans fermented with Bacillus subtilis (natto). I. 6-O-succinylated isoflavone glycosides and their preventive effects on bone loss in ovariectomized rats fed a calcium-deficient diet. Biol. Pharm. Bull. 1999, 22, 1193– 1201.
(18) Slavin, J. L.; Karr, S. C.; Hutchins, A. M.; Lampe, J. W. Influence of soybean processing, habitual diet, and soy dose on urinary isoflavonoid excretion. Am. J. Clin. Nutr. 1998, 68, 1492–1495. (19) Tsangalis, D.; Ashton, J. F.; McGill, A. E.; Shah, N. P. Enzymic transformation of isoflavone phytoestrogens in soymilk by
β-glu-cosidase-producing bifidobacteria. J. Food Sci. 2002, 67, 3104– 3113.
(20) Hessler, P. E.; Larsen, P. E.; Constantinou, A. I.; Schram, K. H.; Weber, J. M. Isolation of isoflavones from soy-based fermentations of the erythromycin-producing bacterium Saccharopolyspora erythraea. Appl. Microbiol. Biotechnol. 1997, 47, 398–404. (21) Esen, A.β-Glucosidase, overview. In β-Glucosidase: Biochemistry
and Molecular Biology. Esen A., Ed.; American Chemical Society:
Washinton, DC, 1993; pp 1–13.
(22) Ghosh, P.; Pamment, N. B.; Martin, W. R. B. Simultaneous saccharification and fermentation of cellulose: Effect of beta-D-glucosidase activity and ethanol inhibition of cellulases. Enzyme
Microb. Technol. 1982, 4, 425–430.
(23) Lei, V.; Amoa-Awua, W. K.; Brimer, L. Degradation of cyano-genic glycosides by Lactobacillus plantarum strains from spon-taneous cassava fermentation and other microorganisms. Int. J.
Food Microbiol. 1999, 53, 169–184.
(24) Marotti, I.; Bonetti, A.; Biavati, B.; Catizone, P.; Dinelli, G. Biotransformation of common bean (Phaseolus vulgaris L.) flavonoid glycosides by bifidobacterium species from human intestinal origin. J. Agric. Food Chem. 2007, 55, 3913–3919. (25) Kuo, L. C.; Cheng, W. Y.; Wu, R. Y.; Huang, C. J.; Lee, K. T.
Hydrolysis of black soybean isoflavone glycosides by Bacillus subtilis natto. Appl. Microbiol. Biotechnol. 2006, 73, 314–320. (26) Choi, Y. B.; Kim, K. S.; Rhee, J. S. Hydrolysis of soybean isoflavone glucosides by lactic acid bacteria. Biotechnol. Lett.
2002, 24, 2113–2116.
(27) Matsuda, S.; Norimoto, F.; Matsumoto, Y.; Ohba, R.; Teramoto, Y.; Ohta, N.; Ueda, S. Solubilization of a novel isoflavone glycosidehydrolyzing β-glucosidase from Lactobacillus casei
subsp. rhamnosus. J. Ferment. Bioeng. 1994, 77, 439–441. (28) Sambrook, J.; Fritsch, E. F.; Maniatis, T. Molecular cloning; a
laboratory manual, 2nd ed.; Cold Spring Harbor Laboratory Press:
Cold Spring Harbor, NY, 1989.
(29) Wang, H. J.; Murphy, P. A. Isoflavone content in commercial soybean foods. J. Agric. Food Chem. 1994, 42, 1666–1673. (30) Kunst, F.; Ogasawara, N.; Moszer, I.; Albertini, A. M.; Alloni,
G.; Azevedo, V.; Bertero, M. G.; Bessières, P.; Bolotin, A.; Borchert, S.; Borriss, R.; Boursier, L.; Brans, A.; Braun, M.; Brignell, S. C.; Bron, S.; Brouillet, S.; Bruschi, C. V.; Caldwell, B.; Capuano, V.; Carter, N. M.; Choi, S. K.; Codani, J. J.; Connerton, I. F.; Danchin, A.; et al. The complete genome sequence of the gram-positive bacterium Bacillus subtilis. Nature
1997, 390, 249–256.
beta-glucosidases. Mol. Gen. Genet. 1989, 217, 70–76. (33) Baird, S. D.; Hefford, M. A.; Johnson, D. A.; Sung, W. L.;
Yaguchi, M.; Seligy, V. L. The Glu residue in the conserved Asn-Glu-Pro sequence of two highly divergent endo-beta-1,4-gluca-nases is essential for enzymatic activity. Biochem. Biophys. Res.
Commun. 1990, 169, 1035–1039.
(34) Hashimoto, W.; Miki, H.; Nankai, H.; Sato, N.; Kawai, S.; Murata, K. Molecular cloning of two genes forβ-D-glucosidase in Bacillus
sp. GL1 and identification of one as a gellan-degrading enzyme.
Arch. Biochem. Biophys. 1998, 360, 1–9.
(35) Gekas, V.; Lopez-Levia, M. H. Hydrolysis of lactose. Process
Biochem. 1985, 20, 2–12.
(36) Paavilainen, S.; Hellman, J.; Korpela, T. Purification, characteriza-tion, gene cloning, and sequencing of a new beta-glucosidase from Bacillus circulans subsp. alkalophilus. Appl. EnViron. Microbiol.
1993, 59, 927–32.
(37) Patchett, M. L.; Daniel, R. M.; Morgan, H. W. Purification and properties of a stableβ-glucosidase from an extremely
thermo-philic anaerobic bacterium. Biochem. J. 1987, 243, 779–787. (38) Krüger, S.; Hecker, M. Regulation of the putative bglPH operon
for aryl-β-glucoside utilization in Bacillus subtilis. J. Bacteriol. 1995, 177, 5590–5597.
(39) Krüger, S.; Gertz, S.; Hecker, M. Transcriptional analysis of bglPH expression in Bacillus subtilis: evidence for two distinct pathways mediating catabolite repression. J. Bacteriol. 1996, 178, 2637– 2644.
(40) Minami, Y.; Kanafuji, T.; Miura, K. Purification and characteriza-tion of a β-glucosidase from Polygonum tinctorium, which
catalyzes preferentially the hydrolysis of indican. Biosci.
Bio-technol. Biochem. 1996, 60, 147–149.
(41) Yang, L.; Ning, Z. S.; Shi, C. Z.; Chang, Z. Y.; Huan, L. Y. Purification and characterization of an isoflavone-conjugates-hydrolyzingβ-glucosidase from endophytic bacterium. J. Agric. Food Chem. 2004, 52, 1940–1944.
(42) Hsieh, M. C.; Graham, T. L. Partial purification and characteriza-tion of a soybeanβ-glucosidase with high specific activity towards
isoflavone conjugates. Phytochemistry 2001, 58, 995–1005. (43) Hósel, W.; Barz, W. β-glucosidase from Cicer Arietinum L.:
purification and properties of isoflavone 7-O-glucoside-specific
β-glucosidases. Eur. J. Biochem. 1975, 57, 607–616.
(44) Setlow, B.; Cabrera-Hernandez, A.; Cabrera-Martinez, R. M.; Setlow, P. Identification of aryl-phospho-β-D-glucosidases in Bacillus subtilis. Arch. Microbiol. 2004, 181, 60–67.
(45) Bhatia, Y.; Mishra, S.; Bisaria, V. S. Micorbialβ-glucosidasees:
cloning, properties and applications. Crit. ReV. Biotechnol. 2002,
22, 375–407.
(46) Ridgway, T. J.; Tucker, G. A. Prospects for the production and use of new improved dietary oestrogens for cardioprotection.
Biochem. Soc. Trans. 1997, 25, 59–63.
(47) Briante, R.; Patumi, M.; Febbraio, F.; Nucci, R. Production of highly purified hydroxytyrosol from Olea europaea leaf extract biotransformed by hyperthermophilic beta-glycosidase. J.
Bio-technol. 2004, 111, 67–77.
Received for review July 30, 2007. Revised manuscript received October 19, 2007. Accepted October 26, 2007.