Abstract.
Antrodia cinnamomea is an expensive and highly
valued folk medicinal fungus that grows only inside the
rotten trunk of Cinnamomum kanehirae, an evergreen
broad-leaved tree. This fungus has recently been used commercially
in the formulation of nutraceuticals and functional foods in
Taiwan. It has been used for centuries as a detoxificant in
cases of food poisoning, diarrhea, vomiting, hepatic disease
and various kinds of cancers. The present study investigated
the effects of Antrodia cinnamomea on mutagenicity using a
bacterial reverse mutation assay employing the Salmonella
typhimurium strains TA97, TA98, TA100, TA102, and
TA1535. The effects of Antrodia cinnamomea on chromosome
structure were tested in Chinese hamster ovary (CHO) cells.
Antrodia cinnamomea was not mutagenic in all bacterial
strains and it was not genotoxic in CHO cells.
Antrodia cinnamomea is a rare and precious medicinal
fungus, the so-called ‘national treasure of Taiwan’, that grows
naturally inside the trunk of aged (over 100 years)
Cinnamonum kanehirae, which is a coniferous tree and
endemic to Taiwan, where it grows in broad-leaved and
coniferous forests at altitudes between 1,300 and 2,800 min
the central and northern parts of the island (1). Many studies
have shown that several chemical components of A.
cinnamomea have anti-oxidant, anti-cancer, anti-virus, and
antibiotic properties (2-4). Demand for A. cinnamomea has
far exceeded the supply and it is now considered an expensive
herbal medicine. Cinnamomum comphora trees have been
illegally harvested resulting in protection by the Taiwan
government which has increased the price of the compound.
According to Enforcement Rules of the Health Food
Control Act established by the Taiwan Department of Health,
health food products should be evaluated for their
pharmacological effects and safety by the Ames test and the
in vitro mammalian chromosome aberration test. In this
study the Ames Salmonella/microsome mutagenicity assay
(Salmonella test; Ames test) was used, which is a short-term
bacterial reverse mutation assay specifically designed to
detect a wide range of chemical substances that can produce
genetic damage that leads to gene mutations (5). A positive
*Both Authors contributed equaly to this study.Correspondence to: Dr. J.-G. Chung, Department of Biological Science and Technology, China Medical University, No 91, Hsueh-Shih Road, Taichung City 404, Taiwan, R.O.C. Tel: +886 4220533662500, Fax: +886 422053764, e-mail: jgchung@mail.cmu.edu.tw. Hsu-Feng Lu, Department of Clinical Pathology, Cheng-Hsin General Hospital, Taipei; Taiwan, R.O.C. No.45,Cheng Hsin St., Pai-Tou,Taipei, Taiwan, R.O.C. Tel: +886 228264400 ext 5850; Fax: +886 228264517, e-mail: ch1835@chgh.org.tw
Key Words: Genotoxicity, Antrodia cinnamomea, Ames test, chromosomal aberration test.
Evaluation of Genotoxicity of Antrodia cinnamomea in the
Ames Test and the In Vitro Chromosomal Aberration Test
MING-FANG WU
1*, FU-CHUO PENG
1*, YUNG-LIANG CHEN
2, CHING-SUNG LEE
3,
YI-YUAN YANG
4, MING-YANG YEH
5, CHI-MING LIU
6, JIN-BIOU CHANG
7,
RICK SAI-CHUEN WU
8, CHIEH-CHIH YU
9, HSU-FENG LU
2,4,10and JING-GUNG CHUNG
11,12 1National Taiwan University College of Medicine Animal Medicine Center, Taipei, Taiwan, R.O.C;
2Department of Medical Laboratory Science and Biotechnology, Yuanpei University, Hsinchu, Taiwan, R.O.C;
3
Department of Restaurant, Hotel and Institutional Management,
Fu-Jen Catholic University, Taipei, Taiwan, R.O.C;
4
School of Medical Laboratory Science and Biotechnology, Taipei Medical University, Taipei, Taiwan, R.O.C;
5Office of Director,
6Department of Research and Education,
10
Department of Clinical Pathology, Cheng Hsin General Hospital, Taipei, Taiwan, R.O.C;
7Department of Pathology, National Defense Medical Center, Division of Clinical Pathology,
Tri-Service General Hospital, Taipei, Taiwan, R.O.C;
8
Departments of Anesthesiology, China Medical University Hospital, Taichung 404, Taiwan, R.O.C;
11Department of Biological Science and Technology, China Medical University, Taichung, Taiwan, R.O.C;
9
Schools of Pharmacy, China Medical University, Taichung, Taiwan, R.O.C;
12Department of Biotechnology, Asia University, Taichung, Taiwan, R.O.C.
test indicates that the chemical might act as a carcinogen
(although a number of false-positives and false-negatives are
known). As cancer is often linked to DNA damage, the test
also serves as a quick assay to estimate the carcinogenic
potential of a compound since it is difficult to ascertain
whether standard carcinogen assays on rodents are
successful. The procedure is described in a series of papers
from the early 1970s by Bruce Ames and his group at the
University of California, Berkeley (6-9).
Materials and Methods
Preparation of Antrodia cinnamomea test solution. Antrodia cinnamomea (500 mg) and 10 ml DMSO were mixed thoroughly and filtered (0.22 μm pore size) to provide a solution with a concentration of 50 mg/ml.
Bacterial strains. Bacterial strains were provided by the Food Science Institute, Hsinchu, Taiwan. The strains used were Salmonella typhimurium TA97 (LLuvrB/rfa/ pKM101), TA98 (LLuvrB/rfa/ pKM101), TA100 (LLuvrB/rfa/ pKM101), TA102 (rfa/ pKM101), and TA1535 (LLuvrB/rfa). Strains were prepared by preculturing for 8 hr at 37˚C in a nutrient broth. Strain properties, including their susceptibility to mutagens, were confirmed prior to use in the assays by the National Taiwan University College of Medicine Animal Medicine Center, Taipei, Taiwan.
Preparation of liver S9 fractions. Rats treated with enzyme-inducing agent β-naphthoflavone were sacrificed by spinal dislocation. Briefly, rat livers were removed, placed in beakers on ice, rinsed with ice-cold homogenization KCl (1.15%) buffer, minced with scissors and then placed in 4 vol. of ice-cold KCl buffer. They were then homogenized with a tissue grinder. The homogenate was transferred to a close-fitting (0.045 mm clearance) perspex [poly(methyl methacrylate)]/glass homogenizer and homogenized. After diluting the homogenate to 10% with the homogenization buffer and centrifuged at 9000× g, the microsomal pellets were suspended in KH2PO4buffer PH 7.4 and stored at –80˚C. Mutagenicity assay. The Ames test was used to examine the mutagenicity of Antrodia cinnamomea. For the plate incorporation method, without metabolic activation (S9), 0.1 ml of the test solutions (a series of various concentrations of Antrodia cinnamomea), 0.1 ml of fresh bacterial broth and 0.5 ml of sterile buffer were mixed with 2.0 ml of overlay agar. For the assay with metabolic activation (S9), usually 0.5 ml of metabolic activation mixtures containing an adequate amount of post-mitochondrial fraction were mixed with the overlay agar (2.0 ml), together with the bacteria and sample. The contents of each tube were mixed and poured over the surface of a minimal glucose agar plate. The overlay agar was allowed to solidify before incubation. The plate was incubated for 48 h at 37˚C and the number of revertant colonies was counted. For a proper estimate of variation, triplicate plating was used at each dose level. All plates in a given assay were incubated at 37˚C for 48-72 hr. After the incubation period, the number of reverting colonies per plate was counted. As positive controls, with S9 mixtures, 1 μg/plate of benzo[a]pyrene for both TA98 and TA102, and 4μg/plate of 2-aminoanthracene for TA97, TA100 and TA1535 were used. As positive controls but without S9 mixtures, 0.5 μg/plate of 4-nitroquinoline-N-oxide for both TA97 and TA98, and 0.5 μg/plate of
mitomycin C for TA102, and 4 μg/plate of sodium azide for both TA100 and TA1535 were used. Control solvent was used as the negative control. Mutagenicity was evaluated based on the rule reported previously by Claxton et al. (10). The value of the positive control should be significantly higher than that of the negative control. To confirm that the experiment was successful, negative control values for TA97, TA98, TA100, TA102 and TA1535 should be 90-180, 30-60, 150-240, 240-320 and 15-35 CFU, respectively. Mutagenicity was judged to be positive when the revertants in the test solution increased more than 2-fold compared with those in the negative control. All the tests of this experiment were performed in triplicate.
In vitro chromosomal aberration test. Antrodia cinnamomea was diluted in dimethylsulfoxide (DMSO; Sigma) before treatment and used at concentrations of 50, 25, 12.5, 6.25 and 3.125 mg/ml. The S9 solution from the rat livers was prepared as described above. The CHO cell line (Food Industry Research and Development Institute, Hsinchu, Taiwan) was grown in McCoy’s 5A medium (Sigma, USA), supplemented with 10% fetal bovine serum, sodium bicarbonate (0.22%), L-glutamine (2 mM), streptomycin (100 μg/ml) and penicillin-G (100 units/ml). Cells were cultured in T-75 plastic cell culture flasks, with 10 ml of culture medium at 37˚C in a humidified atmosphere with 5% CO2 in air. Different concentrations of Antrodia cinnamomea were used in the following conditions: (i) treatment by metabolic activation S9 for 20 h; (ii) treatment by no metabolic activation S9 for 3 h; and (iii) treatment by no metabolic activation S9 for 20 h. After all the above treatments, the cells were harvested for 20 h after exposed to Colcemid (0.1 μg/ml final concentration) for the last 2 h of the incubation period. The cells were harvested and fixed and slides prepared and air-dried as previously described (11). The frequencies of chromosomal aberrations were determined in the first metaphase after treatment. Chromosomal aberrations were classified following the criteria recommended by Archer et al. (12) and by the World Health Organization (13). A total at 100 metaphases per treatment were scored. Data were recorded independently by two groups of observers.
Results
Table I shows the results of the mutagenicity of Antrodia
cinnamomea treatment using the Ames test. Compared with
the negative control, the Antrodia cinnamomea solutions with
S9 or without S9 did not affect bacterial growth. The values
with S9 were lower than without S9 for the TA97 strain
treated with 3.125 and 12.5 mg/ml. Generally, mutagenicity
was negative in all strains with or without the S9 mix, as
shown in Table I.
Chromosome structural alterations, were observed in
well-spread metaphase cells. Among 50 cells, there were 36, 4
and 10 cells with 20, 21 and 19 chromosomes, respectively.
Chromosomal damages induced by Antrodia cinnamomea
indicating frequency of cells with chromosome and
chromatid-type aberrations as well as cells with both types
of aberrations (are shown in Table II). All three frequencies
were increased in cells treated with Antrodia cinnamomea.
Antrodia cinnamomea with S9 mix increased the percentage
of aberrant cells and those effects were dose-dependent. The
percentage of aberrant cells treated with 50 mg/ml was less
compared with the positive control. For the chromosome
structure alteration, 12 out of 21 aberrant cells among 500
cells observed in the S9 mix experiments displayed a
chromatid gap. No chromosome gap, chromosome break,
dicentric, exchange or multiple alterations were observed in
this S9 mix experiments.
Table II also shows that Antrodia cinnamomea without S9
mix at the 3 h raised the percentage of aberrant cells and
these effects were dose-dependent. The percentage of
aberrant cells at the 50 mg/ml treatment was 7%, which was
less than that of the seven positive controls.
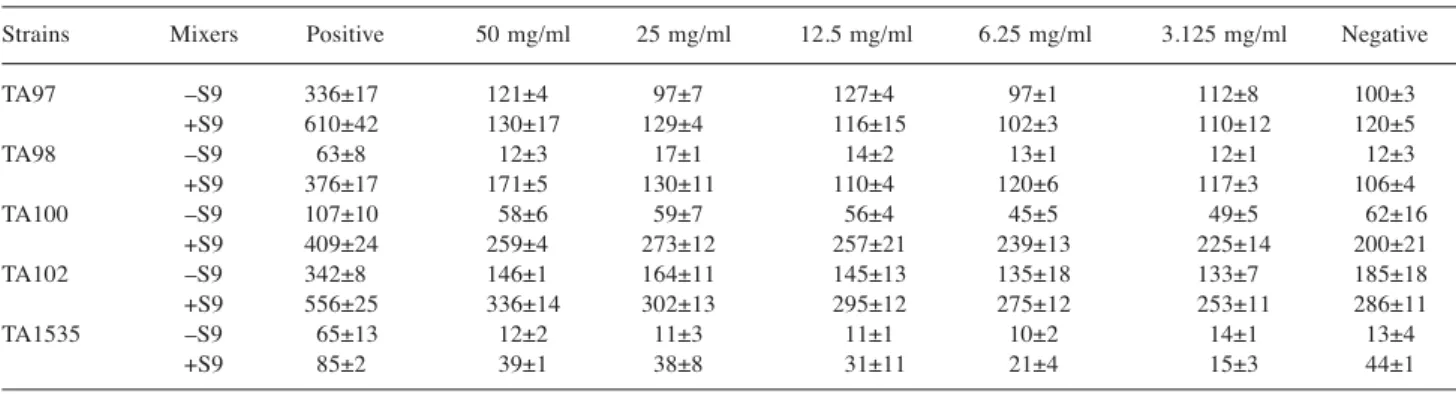
Table I. The numbers of total colonies counts (CFU) including spontaneous revertant colonies that appeared on a plate were measured by the Ames test in various concentrations of Antrodia cinnamomea.
Strains Mixers Positive 50 mg/ml 25 mg/ml 12.5 mg/ml 6.25 mg/ml 3.125 mg/ml Negative
TA97 –S9 336±17 121±4 97±7 127±4 97±1 112±8 100±3 +S9 610±42 130±17 129±4 116±15 102±3 110±12 120±5 TA98 –S9 63±8 12±3 17±1 14±2 13±1 12±1 12±3 +S9 376±17 171±5 130±11 110±4 120±6 117±3 106±4 TA100 –S9 107±10 58±6 59±7 56±4 45±5 49±5 62±16 +S9 409±24 259±4 273±12 257±21 239±13 225±14 200±21 TA102 –S9 342±8 146±1 164±11 145±13 135±18 133±7 185±18 +S9 556±25 336±14 302±13 295±12 275±12 253±11 286±11 TA1535 –S9 65±13 12±2 11±3 11±1 10±2 14±1 13±4 +S9 85±2 39±1 38±8 31±11 21±4 15±3 44±1
The negative control was solvent. The positive control during –S9 mix was 4-nitroquinoline-N-oxide for TA97 and TA98 strains, mitomycin c for TA102 strain and sodium azide for TA100 and TA1535 strains; during +S9 mix, Benzo[a]pyrene was used for TA98 and TA102 strains, and 2-aminoanthracene for TA97, TA100 and TA1535 strains.
Table II. Chromosome analysis of Chinese hamster ovary cells treated with different concentrations of Antrodia cinnamomea and conditions of exposure to S9 or not.
Treatment Abnormal Chromosome aberrations per 100 cells (mg/ml) metaphases (%)a
G B D R g b e MA
S9(–)
3h incubation Positive controlb 13 0 2 0 6 4 1 0 0
50 6 0 0 0 2 1 3 0 0 25 5 0 0 0 1 3 1 0 0 12.5 5 0 0 0 0 2 3 0 0 6.25 3 0 0 0 0 1 2 0 0 3.125 2 0 0 0 0 2 0 0 0 Negative control 1 0 0 0 0 1 0 0 0 S9(+)
20h incubation Positive controlc 15 0 3 0 3 5 4 0 0
50 7 0 0 0 1 4 2 0 0 25 5 0 0 0 1 2 2 0 0 12.5 4 0 0 0 0 3 1 0 0 6.25 4 0 0 0 1 2 1 0 0 3.125 1 0 0 0 0 1 0 0 0 Negative control 1 0 0 0 0 1 0 0 0 S9(–)
20h incubation Positive controld 16 0 4 0 4 6 2 0 0
50 6 0 0 0 0 4 2 0 0 25 6 0 0 0 1 3 2 0 0 12.5 6 0 0 0 1 3 2 0 0 6.25 3 0 0 0 0 2 1 0 0 3.125 1 0 0 0 0 1 0 0 0 Negative control 3 0 0 0 0 2 1 0 0
G: chromosome gap; B: chromosome break; D: dicentric; R: ring; g: chromatid gap; b: chromatid break; e: exchange. MA:multiple aberrations.
aAberrant cells were calculated excluding cells with gaps; bpositive control was 1μM mitomycin C for 3 h incubation; cpositive control was 40 μM
Antrodia cinnamomea with S9 mix and at 20 h incubation
elevated the percentage of aberrant cells and these effects
were dose-dependent. The percentage of aberrant cells in the
50 g/ml treatment was less than that of the positive control.
For the chromosome structure alterations, 13 of 22 cells
among 500 cells showed a chromatid gap. Chromosome
gaps, chromosome breaks, dicentrics, exchange and multiple
alterations were not observed in any S9 mixture at 3 or 20 h
incubation.
Discussion
Three new steroids, zhankuic acids A, B, and C were isolated
from the fruit bodies of Antrodia cinnamomea by
bioassay-guided fractionation. The structures of these compounds were
elucidated by detailed analysis of their 1H- and 13C-NMR
spectra. Biological studies revealed that one exhibited cytotoxic
activity against P-388 murine leukemia cells and two showed
weak anticholinergic and antiserotonergic activities (14).
Well-established in vitro methods for testing the genotoxic
potency of chemicals such as the Ames/Salmonella test, the
mouse lymphoma assay, the micronucleus test and the
chromosomal aberration test, show a high false-positive rate
for predicting in vivo genotoxicity and carcinogenicity (15).
Drugs that contain the nitrate moiety sometimes are shown
positive for Ames when they are indeed safe. Also nitrate
compounds that can potentially generate nitric oxide will
give false positives (16). Nitroglycerin is an example that
gives a positive Ames yet is still used in treatment today.
Thus, there is a need for more reliable in vitro assays. For
example, gene expression profiling in metabolically
competent primary mouse hepatocytes is capable of
discriminating true genotoxic (GTX) compounds from
false-positive genotoxic (FP-GTX) compounds (15). Long-term
toxicology and outcome studies are needed with such
compounds to disprove a positive Ames test.
A positive result in an Ames test does not by itself indicate
that a particular chemical is capable of causing cancer. It
does however suggest that a chemical can produce mutations
and that more extensive testing is needed to determine
whether the chemical is likely to produce cancer in humans.
The test is useful as a screening tool for setting priorities
because it is an inexpensive and quick way to help single out
chemicals that should be subjects of further testing. It is also
used in industry as a primary preventive approach to
eliminate potential carcinogens early in the process of
developing new commercial chemicals.
The chromosome aberration test is also crucial in the
evaluation of products prior to market release. Structural
aberrations may be of two types, chromosome or chromatid.
With the majority of chemical mutagens, induced aberrations
are of the chromatid type, but chromosome type aberrations
also occur. An increase in polyploidy may indicate that a
chemical has the potential to inhibit mitosis. However, this
method is not designed to measure numerical aberrations and
is not routinely used for that purpose. Chromosome
mutations are the cause of many human genetic diseases and
there is substantial evidence that chromosome mutations and
related events causing alterations in oncogenes and tumor
suppressor genes of somatic cells are involved in cancer
induction in humans and experimental animals. This test is
used to screen for possible mammalian mutagens and
carcinogens. Many compounds that are positive in this test
are mammalian carcinogens; however, there is not a perfect
correlation between this test and carcinogenicity. Correlation
is dependent on chemical class and there is increasing
evidence that there are carcinogens that are not detected by
this test because they appear to act through mechanisms
other than direct DNA damage (17-19).
The results presented show that Antrodia cinnamomea is not
mutagenic in all the Salmonella strains used and is also not
genotoxic in the CHO cell in vitro chromosomal aberration test.
References
1 Chang TT and Chou WN: Antrodia cinnamomea reconsidered and A. salmonea sp. nov. on Cunninghamia konishii in Taiwan. Bot Bull Acad Sin 45: 347-352, 2004.
2 Song TY and Yen GC: Antioxidant properties of Antrodia camphorata in submerged culture. J Agric Food Chem 50: 3322-3327, 2002.
3 Chen CH and Yang SW: New steroid acids from Antrodia cinnamomea, a fungal parasite of Cinnamomum micranthum. J Nat Prod 58: 1655-1661, 1995.
4 Lee IH, Huang RL, Chen CT, Chen HC, Hsu WC and Lu MK: Antrodia camphorata polysaccharides exhibit anti-hepatitis B virus effects. FEMS Microbiol Lett 209: 63-67, 2002.
5 Mortelmans K and Zeiger E: The Ames Salmonella/microsome mutagenicity assay. Mutat Res 455: 29-60, 2000.
6 Ames BN, Gurney EG, Miller JA and Bartsch H: Carcinogens as frameshift mutagens: metabolites and derivatives of 2-acetylaminofluorene and other aromatic amine carcinogens. Proc Natl Acad Sci USA 69: 3128-3132, 1972.
7 Ames BN, Lee FD and Durston WE: An improved bacterial test system for the detection and classification of mutagens and carcinogens. Proc Natl Acad Sci USA 70: 782-786, 1973. 8 McCann J, Spingarn NE, Kobori J and Ames BN: Detection of
carcinogens as mutagens: bacterial tester strains with R factor plasmids. Proc Natl Acad Sci USA 72: 979-983, 1975. 9 Ames BN, Durston WE, Yamasaki E and Lee FD: Carcinogens
are mutagens: a simple test system combining liver homogenates for activation and bacteria for detection. Proc Natl Acad Sci USA 70: 2281-2285, 1973.
10 Claxton LD, Allen J, Auletta A, Mortelmans K, Nestmann E and Zeiger E: Guide for the Salmonella typhimurium/mammalian microsome tests for bacterial mutagenicity. Mutat Res 189: 83-91, 1987.
11 Tjio JH and Puck TT: Genetics of somatic mammalian cells II. Chromosomal constitution of cells in tissue culture. J Exp Med 108: 259-271, 1958.
12 Archer PG, Bender M, Bloom A, Brewen J, Carano A and Preston R: Guidelines for cytogenetic studies in mutagen-exposed human populations. In: Bloom AD (ed.). Guidelines For Cytogenetic Studies on Human Populations Exposed to Mutagenic and Reproductive Hazards. March of dimes birth defects foundation, pp. 1-35, 1981.
13 WHO: Guide to short-term tests for detecting mutagenic carcinogenic chemicals. Environmental Health Criteria 51. Geneva: World Health Organization, pp. 57-67, 1985.
14 Chen CH, Yang SW and Shen YC: New steroid acids from Antrodia cinnamomea, a fungal parasite of Cinnamomum micranthum. J Nat Prod 58: 1655-1661, 1995.
15 Mathijs K, Brauers KJ, Jennen DG, Lizarraga D, Kleinjans JC and van Delft JH: Gene expression profiling in primary mouse hepatocytes discriminates true from false-positive genotoxic compounds. Mutagenesis 25: 561-568, 2010.
16 Raisfeld-Danse IH and Chen J: Drug interactions. III. Formation of nitrosamines from therapeutic drugs. Formation, mutagenic properties and safety assessment of propranolol hydrochloride with respect to the intragastric formation of N-nitrosopropranolol under conditions found in patients. J Pharmacol Exp Ther 225: 713-719, 1983.
17 Evans HJ: Cytological Methods for Detecting Chemical Mutagens. In: Chemical mutagens, Principles and Methods for their Detection, Vol. 4, Hollaender A (ed.). Plenum Press, New York and London, pp. 1-29, 1976.
18 Ishidate, M.Jr. and Sofuni, T: The in Vitro Chromosomal Aberration Test Using Chinese Hamster Lung (CHL) Fibroblast Cells in Culture. In: Progress in Mutation Research, Vol. 5, Ashby J et al. (eds.). Elsevier Science Publishers, Amsterdam-New York-Oxford, pp. 427-432, 1985.
19 Galloway, SM, Armstrong MJ, Reuben C, Colman, S, Brown B, Cannon C, Bloom AD, Nakamura F, Ahmed M, Duk S, Rimpo J, Margolin GH, Resnick MA, Anderson G and Zeiger E: Chromosome aberration and sister chromatic exchanges in Chinese hamster ovary cells: Evaluation of 108 chemicals. Environs. Molec. Mutagen 10(suppl 10): 1-175, 1978.