行政院國家科學委員會專題研究計畫 期中進度報告
臺灣 methicillin 抗藥性金黃色葡萄球菌抗藥性基因之分型
研究(1/2)
計畫類別: 個別型計畫 計畫編號: NSC92-2314-B-002-160- 執行期間: 92 年 08 月 01 日至 93 年 07 月 31 日 執行單位: 國立臺灣大學醫學院內科 計畫主持人: 張上淳 共同主持人: 王振泰 計畫參與人員: 吳佳玲 報告類型: 精簡報告 處理方式: 本計畫可公開查詢中 華 民 國 93 年 5 月 26 日
行政院國家科學委員會補助專題研究計畫期中報告
計畫名稱:臺灣 methicillin 抗藥性金黃色葡萄球菌抗藥性基因之分型研究(1/2)
計畫類別:; 個別型計畫 □ 整合型計畫
計畫編號:NSC-92-2314-B-002-160
執行期間:2003 年 8 月 1 日至 2004 年 7 月 31 日
計畫主持人:張上淳醫師
共同主持人:王振泰醫師
計畫參與人員: 吳佳玲
成果報告類型(依經費核定清單規定繳交):;精簡報告 □完整報告
本成果報告包括以下應繳交之附件:
□赴國外出差或研習心得報告一份
□赴大陸地區出差或研習心得報告一份
□出席國際學術會議心得報告及發表之論文各一份
□國際合作研究計畫國外研究報告書一份
處理方式:除產學合作研究計畫、提升產業技術及人才培育研究計畫、列
管計畫及下列情形者外,得立即公開查詢
□涉及專利或其他智慧財產權,□一年□二年後可公開查詢
執行單位:國立台灣大學醫學院內科
中 華 民 國 九十三 年 五 月 三十一 日
研究計畫中文摘要及關鍵詞
methicillin 抗藥性金黃色葡萄球(methicillin-resistant Staphylococcus aureus)菌自 1981 年於台灣出現後,在 1990 年代迅速增加,目前已經成為引起台灣各醫院院內感染 的重要致病菌之一。再加上可資用以治療 methicillin 抗藥性金黃色葡萄球菌感染的用藥 有限,methicillin 抗藥性金黃色葡萄球菌感染對臨床醫學造成了莫大的衝擊。因之,對 methicillin 抗藥性金黃色葡萄球菌的 methicillin 抗藥性機轉的研究,一直是臨床上十分 重要的課題。 在過去 20 多年來對 methicillin 抗藥性金黃色葡萄球菌之抗藥性機轉的研究中,發 現其 methicillin 抗藥性主要源自於 mecA gene 的存在。除了 mecA gene 之外,尚有 mecI 及 mecR1 等調節基因,以此共同組成 mecA gene complex。研究中更發現 mecA gene complex 廣泛的存在 methicillin 抗藥性金黃色葡萄球菌及凝 固 酶 陰性的葡萄球菌 (coagulase-negative Staphylococci)中;由此,抗藥性基因──mecA gene complex 可在 不同種的葡萄球菌中轉移的可能性,應該是存在的。再進一步針對 methicillin 抗藥性金 黃色葡萄球菌的 mecA gene complex 鄰近的染色體做研究,日本學者 Hiramatsu 等人發 現,methicillin 抗藥性金黃色葡萄球菌的染色體基因上帶有一段 methicillin 感受性金黃 色 葡 萄 球 菌 所 沒 有 的 染 色 體 序 列 , 命 名 為 SCCmec 元 件 (Staphylococcal cassette chromosome mec element)。SCCmec 元件中,除了 mecA gene complex 外,另有兩個命名 為 ccrA (cassette chromosome recombinase A)和 ccrB(cassette chromosome recombinase B) 的基因,負責將整個 SCCmec 元件精準的插入及移出金黃色葡萄球菌的染色體。藉由對
mecA gene complex 及 ccr gene 的分型,目前已知世界上的 methicillin 抗藥性金黃色葡萄
球菌的 SCCmec 元件,可分作四個主要的型態。
台灣過去對 methicillin 抗藥性金黃色葡萄球菌的研究,鮮有針對抗藥性基因的部 份。本研究希望藉由對台灣 methicillin 抗藥性金黃色葡萄球菌 mecA gene complex 及 ccr genes 的研究,能釐清台灣本土 methicillin 抗藥性金黃色葡萄球菌其 SCCmec 元件的組 態為何;除了能用以和國外報告相比對以明白台灣 methicillin 抗藥性金黃色葡萄球菌 methicillin 抗藥性之種源外,對本土 MRSA 菌株的抗藥性機轉,也將有更清楚的認識。
II
研究計畫英文摘要及關鍵詞
The first clinical isolate of methicillin-resistant Staphylococcus aureus (MRSA) in Taiwan was found in 1981. During the past decades, the nosocomial MRSA infection rate increased rapidly and MRSA became one of the most important nosocomial pathogens in Taiwan. Given the fact that only few antibiotics can be used to treat MRSA infections, MRSA had become a major problem for current medicine. Thus the mechanism of methicillin
resistance carried by MRSA is also an important clinical issue.
In the previous studies on methicillin resistance of MRSA, it has been found that the methicillin resistance of MRSA is carried by a chromosome gene, mecA gene. The mecA gene and its regulator genes, mecI and mecR1, consists of the mecA gene complex. The mecA gene complex is widely distributed among S. aureus species as well as among other staphylococcal species collectively called coagulase-negative staphylococci. Therefore, it has been
speculated that mec may be freely transmissible among staphylococcal species. In the 1980s, direct chromosome analysis of MRSA strains revealed that a substantial length of the
chromosomal DNA segment (greater than 30 kb) carrying mec has no allelic equivalence in methicillin-susceptible S. aureus (MSSA) strains. According to the studies of Hiramatsu et al, the “mec DNA” is now formally renamed as “SCCmec (Staphylococcal cassette chromosome
mec) element” and the SCCmec element is almost universally found in all MRSA isolates.
The most important components in SCCmec element are the ccr (cassette chromosome recombinase) genes, ccrA and ccrB, and mecA gene complex. The mecA gene complex can be classified into four types, type A, B, C, D. The ccr genes can be classified into to three types. The function of ccr genes is to precisely excise and insert the SCCmec element in way of both site and orientation specifically. Based on the structures and combinations of ccr genes and
mecA gene complex, the SCCmec gene element in MRSA can be classified into four types
In Taiwan, previous studies rarely deal with the genetics of methicillin resistance of MRSA. Our study will focus on the genetic classification of mecA gene complex and ccr complex of MRSA isolated in Taiwan. Despite further determining the types of SCCmec element of MRSA in Taiwan to compare with those in other country and then clarify the its phylogenic source, the mechanism of methicillin resistance of MRSA in Taiwan will be also more illuminated at the same time.
Key Words: methicillin-resistant Staphylococcus aureus, MRSA, mecA gene complex, ccr complex, SCCmec element
研究計畫之前言、目的、及文獻探討
The first report of methicillin-resistant Staphylococcus aureus (MRSA) in the world is by Dr. Jevons in 1961 (1). And the first isolate of MRSA in Taiwan is found in 1981 (2). Thereafter, the rates of nosocomial MRSA infections increased rapidly in Taiwan and most hospitals in Taiwan now have a high incidence of nosocomial MRSA infections (3, 4). The rates of MRSA over all nosocomial isolates of S. aureus in some Taiwan hospitals has already exceeded 80% in 1998 (5), which was much higher than that reported by the National Nosocomial Infection Surveillance System (NNIS) (6). Our previous studies have proved that the reasons leading to rapid increased of MRSA infection in Taiwan include
overuse of antibiotics, poor adherence to isolation precaution of health care worker, and introduction of endemic strain (7-10). MRSA thus has become a major
pathogen in Taiwan. In addition, because of its resistance, limited choice of drug to treat MRSA infection is another important clinical problem. However, there is still no detailed study on the genetic mechanism of methicillin resistance of MRSA in Taiwan. Understanding the detailed genetics of methicillin resistance of MRSA may be helpful to overcome this resistance in the future.
The genetic coding for methicillin resistance in S. aureus has been proven to be mecA gene (11). The expression of mecA gene results in a specific
penicillin-binding protein, PBP2’, that has a decreased binding affinity to β-lactam antibiotics and thus leading to methicillin resistance. The expression of mecA gene is regulated by two adjacent regulatory gene, mecI and mecR1. The mecA, mecI, and mecR1 genes consist of the mecA gene complex (12). The mecA gene complex is widely distributed among S. aureus species as well as among other
staphylococcal species collectively called coagulase-negative staphylococci (13-15). Therefore, it has been speculated that mec may be freely transmissible among staphylococcal species. In the 1980s, direct chromosome analysis of MRSA strains revealed that a substantial length of the chromosomal DNA segment
(greater than 30 kb) carrying mec has no allelic equivalence in
methicillin-susceptible S. aureus (MSSA) strains; the segment was called “additional DNA” or “mec DNA” (16, 17). The size, structure, and biological properties of mec DNA had long remained unclear (18).
The most striking findings in recent studies on MRSA are the existence of SCCmec gene element (19). According to the studies of Hiramatsu et al, the “mec DNA” is now formally renamed as “SCCmec (Staphylococcal cassette
chromosome mec) element” and the SCCmec element is almost universally found in all MRSA isolates (20). The most important components in SCCmec element are the ccr (cassette chromosome recombinase), ccrA and ccrB, genes and mecA gene
2
complex (19, 20). The mecA gene complex can be classified into four types, type A, B, C, D. The ccr genes can be classified into to three types. The function of ccr genes is to precisely excise and insert the SCCmec element in way of both site and orientation specifically (19). Based on the structures and combinations of ccr genes and mecA gene complex, the SCCmec gene element in MRSA can be classified into four types (20, 21). Type III SCCmec gene carries more drug-resistant determinants than any other types. According to the study, conducted by Ito et al, on the analysis of SCCmec element of 38 major hospital-acquired MRSA strains isolated
worldwide, SCCmec elements of MRSA isolates from Europe belong to type I and III, those from northern America belong to type II, most of those from Japan
belong to type II, those from Australia and southeastern Asia belong to type III, and those from south Africa belong to type I (20). Type IV SCCmec element is so far only found in community-acquired MRSA isolates (21).
There is still no published report about the detailed genetics of methicillin resistance of MRSA isolates in Taiwan. The type of SCCmec elements of MRSA isolated in Taiwan is also obscure. Our study is designed to illuminate the mecA complex (in the first year), and ccr genes (in the second year) of MRSA isolates in Taiwan as well as to determine the types of SCCmec elements in Taiwan and compare these results with MRSA isolates in other country. In addition, there are some sporadic cases of community-acquired MRSA infections. By typing the SCCmec gene element of MRSA isolates isolated from patients with nosocomial MRSA infections and those with community-acquired MRSA infections, whether the methicillin resistance comes from the same source between these isolates or not can also be determined.
研究方法
Definitions and MRSA isolates:
Patients with community-acquired MRSA septicemia is defined as that patients without history of hospitalization within prior 30 days develop signs and symptoms of sepsis before admission and their blood cultures taken within 48 hours after hospitalization yield MRSA. Patients with nosocomial MRSA
septicemia is defined as that patients develop signs and symptoms of sepsis three more days after admission and their blood cultures yield MRSA. Based on our previous study, there are seven major types of nosocomial MRSA isolates determined by pulsed-field gel electrophoresis (PFGE) in Taiwan (10). Seven type-specific nosocomial MRSA strains, determined in our previous study, and at least 30 community-acquired MRSA isolates collected from our hospital,
National Taiwan University Hospital, during January 2003 to December 2003 (estimated according to the database of National Taiwan University Hospital in year 2002), are preserved for the following studies.
Study in the first year: Genetic organization of mecA gene complex
Determination of minimum inhibitory concentration (MIC):
All isolates will be tested for their MIC levels of oxacillin, gentamicin, clindamycin, erythromycin, ofloxacin, levofloxacin, chloramphenicol, tetracycline, rifampin, trimethoprim/sulfamethoxazole, vancomycin,
teicoplanin, and linezolide using agar dilution method proposed by NCCLS (22).
PFGE:
All isolates will be typed first by PFGE to determine whether those community-acquired isolates belonged to the same molecular types of nosocomial isolates or not. The methods used for undergoing PFGE will be as those described in our previous study (9). The interpretation of PFGE patterns will be according to the principals proposed previously (23, 24). Once a community-acquired isolate is proved to belong to the same type as a nosocomial isolate, it will not be used for further molecular study. All results will be double checked.
PCR and nuleotide sequencing for the analysis of mecA complex:
The chromosomal DNA will be prepared by means of the method described by Hiramatsu et al and Matsuhashi et al (25, 26). PCR
4
Foster City, Calif.) in 50 µl of reaction mixture (10mM Tris-HCl [pH 8.3], 50 mM KCl, 0.001% [wt/vol] gelatin, 50% [vol/vol] glycerol, 1.5 mM MgCl2,
200 mM each deoxynucleoside triphosphatge, 1.0 mM each primer, and template DNA). The reaction will be carried out by using a Gene Amp PCR system 9600 (Perkin-Elmer). Thermal cycling will be set at 30 cycles (30 s for denaturation at 94℃, 1 min for annealing at 50℃, and 2 min for elongation at 72℃)
Long-range PCR amplification will be performed using 2.6 U of Expand high-fidelity PCR system enzyme mix as recommended by the manufacturer (Boehringer Mannheim Biochemica, Mannheim, Germany). A 5-µl portion of the reaction volume was subjected to electrophoresis in a 0.8% agarose gel containing 1 µl of ethidium bromide per ml to detect the amplified DNA fragment. All PCR products will be further sequenced using a 377 automated fluorescent DNA sequencing system (Perkin-Elmer, Foster City, Calif.) to compare the nucleotide homology with the published sequence in GenBank. The PCR and sequencing results will also be double-checked.
PCR primers and detection of mecA gene complex:
The primers will be used for the detection of mecA gene complex include: mA2 (5’-AACGTTGTAACCACCCCAAGA-3’), mA4 (5’-AGTGTATGATGAGCTATGAGA-3’), mA5 (5’- CGCTCAGGAAATTTGTTGTGC-3’), mA6 (5’-TATACCAAACCCGACAAC-3’), iS1 (5’-ACATTAGATATTTGGTTGCGT-3’), iS3 (5’-TCGGATGCTATCATTAAGCAT-3’), iS4 (5’-ACAATCTGTATTCTCAGGTCGT-3’), mI-1 (5’-AATGGCGAAAAAGCACAACA-3’), mI-2 (5’-GACTTGATTGTTTCCTCTGTT-3’), mcR2 (5’-CGCTCAGAAATTTGTTGTGC-3’), and mcR3 (5’-ATACTCCACGTTAATTCCATT-3’) (27).
Technical detection of the class A mecA gene complex will be based on the positive PCR test results for both sets of primers, mI-1 plus mI-2 and mcR2 plus mcR3. The class B mecA gene complex will be detected by
long-range PCR using two sets of primers, iS-3 plus mA5 and iS-4 plus mA5. Detection of the Class C mecA gene complex will be based on the negative PCR test results for two sets of primer, mI-1 plus mI-2 and mcR2 plus mcR3,
as well as a positive long-range PCR test result for one set of primers, iS-1 plus mA6. Detection of the Class D mecA gene complex will be based on the negative PCR test results for two sets of primer, mI-1 plus mI-2 and mcR2 plus mcR3, as well as a negative long-range PCR test result for one set of primers, iS-1 plus mA6 (27, 28).
6
結果
MRSA isolates
Based on our previous study using pulsed-field gel electrophoresis (PFGE) for molecular typing, there are seven major PFGE types (type C, G, J, K, R, U, and X, Figure 1) of nosocomial MRSA allover Taiwan. We randomly selected one MRSA isolate of each major type (seven MRSA isolates in total) to undergo further molecular studies. In addition, there are 55 MRSA isolates causing community-acquired MRSA infection from January 2003 to December 2003. These 55 MRSA isolates are also enrolled for PFGE typing.
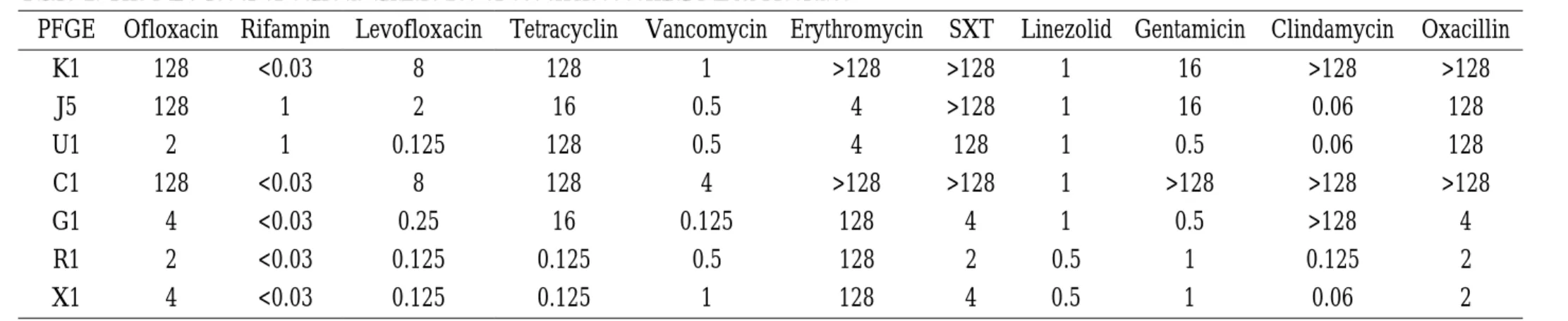
Determination of minimum inhibitory concentration (MICs) of various antibiotics: For the seven nosocomial MRSA isolates, the MICs level of various antibiotics are described in details in Table 1. For the 55 community-acquired MRSA isolates, the MIC50s and MIC90s of various antibiotics are described in
details in Table 2.
PFGE molecular typing:
Using PFGE molecular typing, we find the 55 community-acquired MRSA isolates belong to the 11 PFGE types (type C, G, J, K, R, U, X, D, H, S, and A, Figure 2), which all have been already found in nosocomial MRSA isolates in our previous study. However, 34 of the 55 community-acquired MRSA isolates belong to other four minor PFGE types (type D, H, S, and A) and not to the seven major PFGE types. We also randomly select one MRSA isolate of each minor types (four isolates in total) to undergo analysis of their mecA gene complex. Therefore, 11 isolates in total are handled to determine types of their
mecA gene complex.
Analysis of mecA complex:
Class A mecA gene complex, determined by positive polymerase chain reaction (PCR) test results for both sets of primers, mI-1 plus mI-2 and mcR2 plus mcR3, is found in MRSA isolates of PFGE type C, J, K, and U. Class B mecA gene complex, determined by long-range PCR using two sets primers, iS-3 plus mA5 and iS-4 plus mA5, is found in MRSA isolates of type G, R, X, D, H, S, and A. No class C or class D mecA gene complex is found in our MRSA isolates.
Table 1. The MIC level of various antibiotics of seven nosocomial MRSA isolates
PFGE Ofloxacin Rifampin Levofloxacin Tetracyclin Vancomycin Erythromycin SXT Linezolid Gentamicin Clindamycin Oxacillin
K1 128 <0.03 8 128 1 >128 >128 1 16 >128 >128 J5 128 1 2 16 0.5 4 >128 1 16 0.06 128 U1 2 1 0.125 128 0.5 4 128 1 0.5 0.06 128 C1 128 <0.03 8 128 4 >128 >128 1 >128 >128 >128 G1 4 <0.03 0.25 16 0.125 128 4 1 0.5 >128 4 R1 2 <0.03 0.125 0.125 0.5 128 2 0.5 1 0.125 2 X1 4 <0.03 0.125 0.125 1 128 4 0.5 1 0.06 2 SXT, trimethoprim/sulfamethoxazole
8
Table 2. The MIC50s and MIC90s of various antibiotics of 55 community-acquired MRSA isolates
Ofloxacin Rifampin Levofloxacin Tetracyclin Vancomycin Erythromycin SXT Linezolid Gentamicin Clindamycin Oxacillin MIC90 128 <0.03 8 128 1 >128 >128 1 16 >128 >128
Figure 1. PFGE types of seven nosocomial MRSA isolate K1 C1 G1 m m m m J5 X1 U1 R1 100 90 80 70 60 50
10
Figure 2. The 11 PFGE types of community-acquired MRSA isolates
X1 R1 m J5 K1 C1 U1 D3 H3 S G1 A1 100 90 80 70 60 50 40
討論
MRSA isolates of PFGE type C comprise about half of the all nosocomial MRSA isolates allover Taiwan. PFGE type C is the predominant type in Taiwan. Our study finds that class A mecA gene complex is the major mecA gene complex in Taiwan. As previous studies, no class C or D
mecA gene complex is found in our MRSA isolates. All MRSA isolates tested in our study are
susceptible to rifampin, vancomycin, and linezolid. In addition, almost During the following year, we will determine the ccr gene types of the 11 MRSA isolates. After the availability of ccr gene and
mecA gene complex typing, we can further determine the types of SCCmec element of the 11
12
計劃成果自評:
All the objects listed in our planning have been achieved. Our study demonstrates the distribution of mecA gene complex in MRSA in Taiwan. This will be helpful for further discovering the whole picture of resistant mechanism of MRSA isolates in Taiwan. And after the results of typing of ccr gene complex become available, we think it is suitable to publish our findings in literatures cited by SCI.
Reference:
1. Jevons MP. ‘Celbenin’-resistant staphylococci. Br Med J 1961;1:124-5.
2. Chang SC, Hsu LY, Luh KT, Hsieh WC. Methicillin-resistant Staphylococcus aureus infections. J Formos Med Assoc 1988;87:157-63
3. Chang SC, Sun CC, Yang LS, et al. Increasing nosocomial infections of
methicillin-resistant Staphylococcus aureus at a teaching hospital in Taiwan. Int J
Antimicrob Agents. 1997;8:109-114.
4. Lee SC, Wang CH, Lee N. Infections caused by methicillin-resistant Staphylococcus
aureus: distinct risk factors and prognosis. Nosocomial Infect Control J (Taiwan).
1996;6:131-138.
5. Ho M, McDonald L, Lauderdale T, et al. Surveillance of antibiotic resistance in Taiwan, 1998. J Microbiol Immunol Infect. 1999;32:239-249.
6. National Nosocomial Infections Surveillance (NNIS) System. (2001). National
Nosocomial Infections Surveillance (NNIS) System report: data summary from January 1992-June 2001, issued August 2001. Am J Infect Control. 2001;29:404-421.
7. Chang SC, Chang HJ, Hsiao ML. Antibiotic usage in public hospitals in Taiwan. J
Microbiol Immunol Infect 1998;31:125-32
8. Wang JT, Chang SC, Ko WJ, Chang YY, Chen ML, Pan HJ, Luh KT. A hospital-acquired outbreak of methicillin-resistant Staphylococcus aureus infection initiated by a surgeon carrier. J Hosp Infect 2000;47:104-9
9. Chen ML, Chang SC, Pan HJ, Hsueh PR, Yang LS, Ho SW, Luh KT. Longitudinal analysis of methicillin-resistant Staphylococcus aureus isolates at a teaching hospital in Taiwan. J Formos Med Assoc 1999;98;426-32.
10. Wang JT, Chen YC, Yang TL, Chang SC. Molecular epidemiology and antimicrobial susceptibility of methicillin-resistant Staphylococcus aureus in Taiwan. Diagn Microbiol
Infect Dis 2002;42:199-203.
11. Song MD, Wachi M, Doi M, Ishino F, Matsuhashi M. Evolution of an inducible penicillin-target protein in methicillin-resistant Staphylococcus aureus by gene fusion.
FEBS lett 1987;221:167-71.
12. Hiramatsu K, Asada K, Suzuki E, Okonogi K, Yokota T. Molecular cloning and nucleotide sequence determination of the regulator region of mecA gene in methicillin-resistant
Staphylococcus aureus (MRSA). FEBS lett 1991;298:133-6.
13. Ryffel C, Tesch W, Birch-Machin I, Reynolds PE, Barberis-Makino L, Kayser FH, Berger-Bächi B. Sequence comparison of mecA genes isolated from methicillin-resistant
Staphylococcus aureus and Staphylococcus epidermidis. Gene 1990;94:137-8.
14. Suzuki E, Hiramatsu K, Yokota T. Survey of methicillin-resistant clinical strains of coagulase-negative staphylococci for mecA gene distribution. Antimicrob Agents
14
15. Suzuki E, Kuwahara-Arai K, Richardson JF, Hiramatsu K. Distribution of mec regulator genes in methicillin-resistant Staphylococcus clinical strains. Antimicrob Agents
Chemother 1993;37:1219-26.
16. Beck WD, Berger-Bächi B, Kayser FH. Additional DNA in and molecular cloning of
mec-specific DNA. J Bacteriol 1986;165:373-8.
17. Dubin DT, Matthews PR, Chikramane SG, Stweart PR. Physical mapping of the mec region of an American methicillin-resistant Staphylococcus aureus strain. Antimicrob
Agents Chemother 1991;35:1661-5.
18. Ito T, Katayama Y, Hiramatsu K. Cloning and nucleotide sequence determination of the entire mec DNA of pre-methicillin-resistant Staphylococcus aureus N315. Antimicrob
Agents Chemother 1999;43:1449-58.
19. Katayama Y, Ito T, Hiramatsu K. A new class of genetic element, Staphylococcus cassette chromosome mec, encodes methicillin resistance in Staphylococcus aureus. Antimicrob
Agents Chemother 2000;44:1549-55.
20. Ito T, Katayama Y, Asada K, Mori N, Tsutsumimoto K, Tiensasitorn C, Hiramatsu K. Structural comparison of three types of staphylococcal cassette chromosome mec
integrated in the chromosome in methicillin-resistant Staphylococcus aureus. Antimicrob
Agents Chemother 2001;45:1323-36.
21. Ma XX, Ito T, Tiensasitorn C, Jamklang M, Chongtrakool P, Boyle-Vavra S, Daum RS, Hiramatsu K. Novel type of staphylococcal cassette chromosome mec identified in community-acquired methicillin-resistant Staphylococcus aureus strains. Antimicrob
Agents Chemother 2002;46:1147-52.
22. National Committee for Clinical Laboratory Standards (NCCLS). Methods for dilution
antimicrobial susceptibility tests for bacteria that grow aerobically, Approved Standard.
Villanova, PA: NCCLS, 2000.
23. Bannerman TL, Hancock GA, Tenover FC, Miller JM. Pulsed-field gel electrophoresis as a replacement for bacteriophage typing of Staphylococcus aureus. J Clin Microbiol 1995;33:551-5.
24. Jorgensen M, Givney R, Pegler M, Vickery A, Funnell G. Typing multidrug-resistant
Staphylococcus aureus: conflicting epidemiological data produced by genotypic and
phenotypic methods clarified by phylogenetic analysis. J Clin Microbiol 1996;34:398-403. 25. Hiramatsu K, Kihara H, Yokota T. Analysis of borderline-resistant strains of
methicillin-resistant Staphylococcus aureus using polymerase chain reaction. Microbiol
Immunol 1992;36:445-53.
26. Matsuhashi M, Song MD, Ishino F, Wachi M, Doi M, Inoue M, Ubukata K, Yamashita N, Konno M. Molecular cloning of the gene of a penicillin binding protein supposed to cause high resistance to beta-lactam antibiotics in Staphylococcus aureus. J Bacteriol
27. Katayama Y, Ito T, Hiramatsu K. Genetic organization of the chromosome region
surrounding mecA in clinical staphylococcal strains: role of IS431-mediated mecI deletion in expression of resistance in mecA-carring, low-level methicillin-resistant Staphylococcus
aureus. Antimicrob Agents Chemother 2001;45:1955-63.
28. Hiramatsu K, Cui L, Kuroda M, Ito T. The emergence and evolution of